External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G18590 248 / 3e-80
SOT17, ATSOT17, ATST5C
ARABIDOPSIS SULFOTRANSFERASE 5C, sulfotransferase 17 (.1)
AT1G74100 242 / 4e-78
SOT16, ATSOT16, CORI-7, ATST5A
CORONATINE INDUCED-7, ARABIDOPSIS SULFOTRANSFERASE 5A, sulfotransferase 16 (.1)
AT3G45070 241 / 8e-78
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT3G45080 239 / 7e-77
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT1G13420 239 / 9e-77
SST1, ATST4B
ARABIDOPSIS THALIANA SULFOTRANSFERASE 4B, sulfotransferase 4B (.1)
AT1G74090 231 / 1e-73
SOT18, ATSOT18, ATST5B
ARABIDOPSIS SULFOTRANSFERASE 5B, desulfo-glucosinolate sulfotransferase 18 (.1)
AT5G43690 224 / 4e-71
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT4G26280 223 / 6e-71
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT5G07010 221 / 2e-69
ATST2A
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2A, sulfotransferase 2A (.1)
AT1G28170 219 / 4e-69
SOT7
sulphotransferase 7 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G048200
548 / 0
AT1G18590 273 / 7e-90
ARABIDOPSIS SULFOTRANSFERASE 5C, sulfotransferase 17 (.1)
Potri.011G049100
515 / 0
AT1G18590 266 / 3e-87
ARABIDOPSIS SULFOTRANSFERASE 5C, sulfotransferase 17 (.1)
Potri.011G048712
514 / 0
AT1G18590 265 / 1e-86
ARABIDOPSIS SULFOTRANSFERASE 5C, sulfotransferase 17 (.1)
Potri.011G048300
498 / 1e-178
AT1G18590 244 / 1e-78
ARABIDOPSIS SULFOTRANSFERASE 5C, sulfotransferase 17 (.1)
Potri.011G048100
496 / 5e-178
AT1G74100 251 / 1e-81
CORONATINE INDUCED-7, ARABIDOPSIS SULFOTRANSFERASE 5A, sulfotransferase 16 (.1)
Potri.011G048500
366 / 3e-127
AT1G74100 164 / 2e-48
CORONATINE INDUCED-7, ARABIDOPSIS SULFOTRANSFERASE 5A, sulfotransferase 16 (.1)
Potri.011G048700
337 / 7e-117
AT3G45080 132 / 6e-37
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.012G032700
278 / 2e-92
AT1G74100 295 / 4e-99
CORONATINE INDUCED-7, ARABIDOPSIS SULFOTRANSFERASE 5A, sulfotransferase 16 (.1)
Potri.003G193400
243 / 2e-78
AT5G07010 393 / 6e-137
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2A, sulfotransferase 2A (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017878
266 / 3e-87
AT1G18590 289 / 3e-96
ARABIDOPSIS SULFOTRANSFERASE 5C, sulfotransferase 17 (.1)
Lus10026147
223 / 2e-70
AT5G07010 425 / 2e-149
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2A, sulfotransferase 2A (.1)
Lus10008673
223 / 2e-70
AT5G07010 429 / 3e-151
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2A, sulfotransferase 2A (.1)
Lus10033720
196 / 8e-60
AT1G74100 229 / 6e-73
CORONATINE INDUCED-7, ARABIDOPSIS SULFOTRANSFERASE 5A, sulfotransferase 16 (.1)
Lus10033721
193 / 4e-56
AT1G18590 223 / 6e-67
ARABIDOPSIS SULFOTRANSFERASE 5C, sulfotransferase 17 (.1)
Lus10031623
184 / 1e-55
AT2G03760 201 / 3e-62
ARABIDOPSIS THALIANA SULFOTRANSFERASE 1, sulphotransferase 12 (.1)
Lus10033718
182 / 7e-55
AT1G74100 209 / 3e-65
CORONATINE INDUCED-7, ARABIDOPSIS SULFOTRANSFERASE 5A, sulfotransferase 16 (.1)
Lus10041611
182 / 2e-54
AT3G45070 239 / 9e-77
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10003070
179 / 1e-53
AT3G45070 219 / 2e-69
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10018047
174 / 1e-51
AT1G74100 208 / 7e-65
CORONATINE INDUCED-7, ARABIDOPSIS SULFOTRANSFERASE 5A, sulfotransferase 16 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00685
Sulfotransfer_1
Sulfotransferase domain
Representative CDS sequence
>Potri.011G048400.1 pacid=42782343 polypeptide=Potri.011G048400.1.p locus=Potri.011G048400 ID=Potri.011G048400.1.v4.1 annot-version=v4.1
ATGGAATCCTCTTTGCCTTCCACCCAAATCTTGGATTCTGGTGAAAGTGCTGAGAGGAAAAATGAAGCTAAAAATTACAATGAAGTCATGTCCACCCTTC
CAAAAGTAAAAGGCTTGAATGGGTGTGATTATTACCTGTACCAATGCTTTTGGTACGATCCCTCCTTCTTGGAAGGAATCATGTCTGTTCAAGAGCGCTT
CAATCCTCAATCCACTGATATATTTCTCGCCAGTTTTCCAAAAACTGGCACAACTTGGCTTAAGGCCCTCACTTTTGCTATTTGTACACGATCCCGTTTA
AGTGGTTCAACAACTAGTTCTTTACTTTCCAAGATGCCACATGACTGTGTTCCTTTTATGGAATATGACCTTGCTCAGAACCCAAGTAATCGGGACTTGG
CAATTCCTCTGGTATCTACTCATGTTCCTTACACTTGTTTACCTAAATCTATCATTTCTTCTTGTTGCAAGATTATTTACATTTGCAGGGACGCAAAGGA
TGTTTTTGTTTCTTTGTGGTACTTTCTTGCCAGGCTGCAAATGTCGGAAAATGTTGAGCCTCTTCTTCTGGAAGAGGCTTTTGAGTTGTTTTGCAATGGA
ATTGCAAATTTTGGACCATATTGGGACCATGTTTTAGGGTATTGGAGAGCAAGCTTAGAGTTCCCTGAGAAGATATTGTTCTTGACATACGAGGAAATGA
AGAAAGACACCGCTGCTCATATTAAGAAAGTAGCTGAGTTCATGGATTGTTCTTTCACCTTAGAGGAAGAGGGGGAGGGGGAGGTGCAAAAGATAATAAG
CATGTGTAGTTTTGAGGAGTTGAGCACCTTAGAGGTGAATAAACATGGGCAGCGCCGTCTAGACACATCAATTACTATTCAAAATAGTATATACTTCAGG
AAAGGTGAGATAGGCGACTGGGCAAATCACTTGACACCTGAAATGGGAGCACGTCTAGATGACATAATGGAGCGGAAGCTCAAGGGTTCAGGCTTGAAGC
TGCCCAGGTGA
AA sequence
>Potri.011G048400.1 pacid=42782343 polypeptide=Potri.011G048400.1.p locus=Potri.011G048400 ID=Potri.011G048400.1.v4.1 annot-version=v4.1
MESSLPSTQILDSGESAERKNEAKNYNEVMSTLPKVKGLNGCDYYLYQCFWYDPSFLEGIMSVQERFNPQSTDIFLASFPKTGTTWLKALTFAICTRSRL
SGSTTSSLLSKMPHDCVPFMEYDLAQNPSNRDLAIPLVSTHVPYTCLPKSIISSCCKIIYICRDAKDVFVSLWYFLARLQMSENVEPLLLEEAFELFCNG
IANFGPYWDHVLGYWRASLEFPEKILFLTYEEMKKDTAAHIKKVAEFMDCSFTLEEEGEGEVQKIISMCSFEELSTLEVNKHGQRRLDTSITIQNSIYFR
KGEIGDWANHLTPEMGARLDDIMERKLKGSGLKLPR
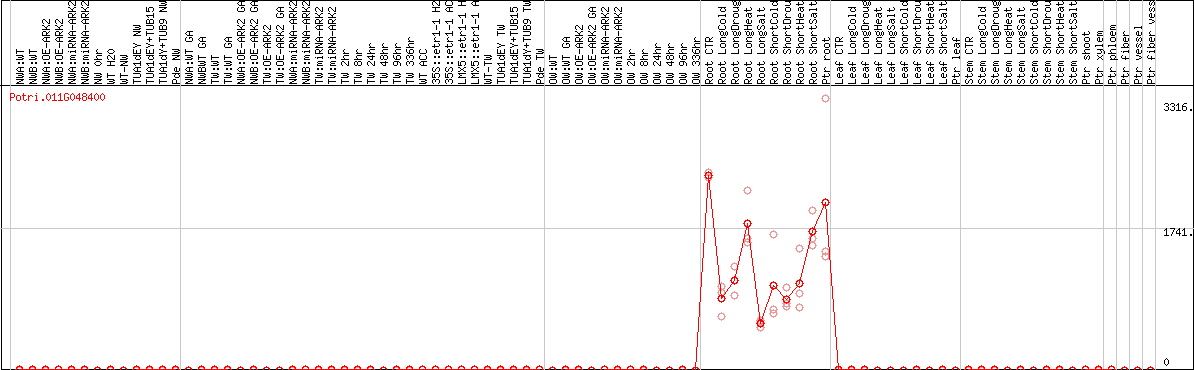
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G048400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.