Potri.011G052500 [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||
|
Representative CDS sequence |
>Potri.011G052500.3 pacid=42781294 polypeptide=Potri.011G052500.3.p locus=Potri.011G052500 ID=Potri.011G052500.3.v4.1 annot-version=v4.1
ATGGAGCCGTCTCGCGCTCCTAAATCATCTCCATTTCTCGAAAACTCTACTTCCCGCAGATTCCCAACCACTCCCTTCCTCATTTCCCTCGCCTTTCTTC
TCCTTACAACCTCGGCAACCTCTGCACCAACAATAAATTCCTTTAATTTCCTTGAATATTACGCCGAACACTGCAACAACGTAGTCCCTGAATCACCAAT
CACAGGTACTCTCATAAACAATGCTTCTTTCTTTGAAGACAAAATCAAAATCCTTAATTTTGATGTCGCATACTTTACTGGTGGCTCACAAATCATACCC
AAAAAAAGGGACTCTGATTCGGCTCCGAGTGTCCTCTCTTTTAAGCCAAAAAAATTCGATCTTCAGCAAACTGTGAACCCTTATGTGGTTTCCCTTCGAG
GGTCTCTTAAATTTCGCTTTCCTGCTCGGTTTGATTGGAGTAATGTGACACGGGATCGCAGGAATTCGAAGAGAATCCGGTACCGCCCGCCGAGAACTCC
AGTCAGGTCGAGGTATCTACTGTTTGAGCTATATGGGTTCTGGTCAATGAATACAGGGAAGCTGTGTATGGTGGGATCTGGTTCAGGTAACAGTGGTCTA
AGTTCTTTAAATGCTGCTTTTAAGGCTAATTATCCTGTGGGTATTAGTGACTTTAGTGGCTTGATTAATGGGGTTTTGGAAAGTTTGGATTTTCAGGATA
GTTATTTTGAGCAGGTTTCGATTTTGGGCATTCCTCATTTTGGTGAATACAAGTATACATTGGTAGATAAAGAGAATGTGGATGTGGGTTTTAGTGGAAC
TTATGATAGTGTCGGGGGGAGAGAGAATTTGCCTATCGAATCGGTCGATAGAAGTATGTGTTTAAATGAAATGTATAGGCATGCTAGGATATTGGAATTG
GAGTATGGGAGTGATTGTAGTGGTGATAATGGTGGTAAATGTAATCCTCTTAGTGGGAGTTCTGGGGTTTTGCCGAAGATTATGACAATTCAGGGAATTA
GATGTGACCATGAGCGTGGGCGTGAAGCTCGGGTTTTGATTGGGTTTTCAGATTCTGCCGTTGTTAATGTTTATGGACCTTATGGCTCTGAAAGAGTTTT
TGATCCTTATACAACACTAATTGGCGAGGGGGTCTGGGATGAGAAGAGGAATAGACTATTTGTGGTTGCATGTCGTGTCTTGAATTTTAATGATTCATCA
GCTAATGCTACTGTTGGTGATTGCTCGATTCAATTGACCTTGCGGTTCCCCAGAACCTTGACAATTAGAGATCAGAGTGTTGTAGTGGGGCAAATTTATA
GCAACAAAACTGTGAATGACACGAGCTACTTTCCTGGAATTGGGTTTCATGGTTCTGAGTTTCGGACAAGGCGACTTCGAGGTCTGGCTTATGAGTATAC
TATGCTTGACAAGGTGCATAAATCTTGTGCAGAAAAGAAGTCCATGAAAGGTAAAGGAAAGACATACCCACATGGTTATTCATCTGACATGCGGTTTGAT
ATGTTAGTGAGAAATGGAAAAGGACACGTAGCACAGGGCTTTTCAACCCCATTGTTTGTGGGTTATCAGCTGTTTGAGCCATATCCAATGACCAACAACT
ACAGTGGTCATCTAAATATTAGCTACAAGATGTTGTTCACGGGTATGCTTCCATCAAATGACTCTGGCACAATTTCAGCTGAAGGAACGTACGATGATGA
GAATGGGGTTCTATGCATGATAGGTTGTCGGCATTTGATATCACGTATGGGAAATTCAATGAAAAATGATTCAACTGACTGTGAGATTCTTGTCAATGTC
CAGTTTTCTCCACTGAATGGAAAGGGTCACGGTAACATCAAAGGAACCATTGAAAGCGTAAGGAAAAACTCAGACCCTCTTCACTTTGAGAAGCTTGAGA
TATCTTCAAACTCGATTTATAGGCACCAAGCTGCGGAATCCATTTGGAGAATGGATATGGAAATAACCATGGTTTTGATTTCTAGTACTCTTGCTTGTAT
CCTTGTGGGTTTGCAGCTCTATCATGTGAAAAGGCATCCTGATGTGCTAACCTTCATTTCTTTTATGATGCTTCTTGTTCTTACTCTGGGTCATATGATA
CCTCTATTGTTAAATTTCGAAGCCTTGTTTTTGTCAAACCGTAACCAACAAAATGTTTTTCTTGAGAGTGGTGGATGGCTTGAAGTCAATGAAGTAGCAG
TAAGGGTGGTAAAGATGGTAGCCTTTCTTTTGATATTCCGGCTTCTCCAACTCACTTGGTCTGCTAGGCCGAGCGATGGAAGCAACAAAAACGTGTGGAT
TTCTGAGAAAAGAGTTCTTTATTTGTCTTTACCGATGTACATTGTTGGTGGACTCATTGCATGGTATGTGCATCACTGGAAAAACACTTCTAGGAGCCCA
CACCTGCTACAAGGTCATAAGGTTTACCAACAACATTATCCCTGGACAGATCTTAAATCATATGCTGGTTTGGTTCTGGATGGTTTCTTACTCCCACAGA
TAATGTTTAACTTATTCTTGAACTCTAGTGAAAAAGCTCTTGCCCCTTCGTTTTATGCTGGGACAACTGTTATTCGTTTGCTGCCTCATGCATATGACCT
TTACAGGGCTCACAGTTCCACATGGTACCTTGATTTGTCATACTTGTATGCAAATCACACATATGATTTTTATTCTACTGCATGGGACATCATAATCCCT
TTGTGTGGCCTGTTGTTTGCTATCCTCATTTACTTACAGCAGCAATTTGGTGGTCGTTGCTTTCTTCCAAAGAGATTTAGAGGGGGTCCTGCATATGAAA
AGGTTCCCATAGTTAGCAACGAGGAACTGCAAGAGATAACAACTCATTGA
|
||||||||||||||||||||
|
AA sequence
|
>Potri.011G052500.3 pacid=42781294 polypeptide=Potri.011G052500.3.p locus=Potri.011G052500 ID=Potri.011G052500.3.v4.1 annot-version=v4.1
MEPSRAPKSSPFLENSTSRRFPTTPFLISLAFLLLTTSATSAPTINSFNFLEYYAEHCNNVVPESPITGTLINNASFFEDKIKILNFDVAYFTGGSQIIP
KKRDSDSAPSVLSFKPKKFDLQQTVNPYVVSLRGSLKFRFPARFDWSNVTRDRRNSKRIRYRPPRTPVRSRYLLFELYGFWSMNTGKLCMVGSGSGNSGL
SSLNAAFKANYPVGISDFSGLINGVLESLDFQDSYFEQVSILGIPHFGEYKYTLVDKENVDVGFSGTYDSVGGRENLPIESVDRSMCLNEMYRHARILEL
EYGSDCSGDNGGKCNPLSGSSGVLPKIMTIQGIRCDHERGREARVLIGFSDSAVVNVYGPYGSERVFDPYTTLIGEGVWDEKRNRLFVVACRVLNFNDSS
ANATVGDCSIQLTLRFPRTLTIRDQSVVVGQIYSNKTVNDTSYFPGIGFHGSEFRTRRLRGLAYEYTMLDKVHKSCAEKKSMKGKGKTYPHGYSSDMRFD
MLVRNGKGHVAQGFSTPLFVGYQLFEPYPMTNNYSGHLNISYKMLFTGMLPSNDSGTISAEGTYDDENGVLCMIGCRHLISRMGNSMKNDSTDCEILVNV
QFSPLNGKGHGNIKGTIESVRKNSDPLHFEKLEISSNSIYRHQAAESIWRMDMEITMVLISSTLACILVGLQLYHVKRHPDVLTFISFMMLLVLTLGHMI
PLLLNFEALFLSNRNQQNVFLESGGWLEVNEVAVRVVKMVAFLLIFRLLQLTWSARPSDGSNKNVWISEKRVLYLSLPMYIVGGLIAWYVHHWKNTSRSP
HLLQGHKVYQQHYPWTDLKSYAGLVLDGFLLPQIMFNLFLNSSEKALAPSFYAGTTVIRLLPHAYDLYRAHSSTWYLDLSYLYANHTYDFYSTAWDIIIP
LCGLLFAILIYLQQQFGGRCFLPKRFRGGPAYEKVPIVSNEELQEITTH
|
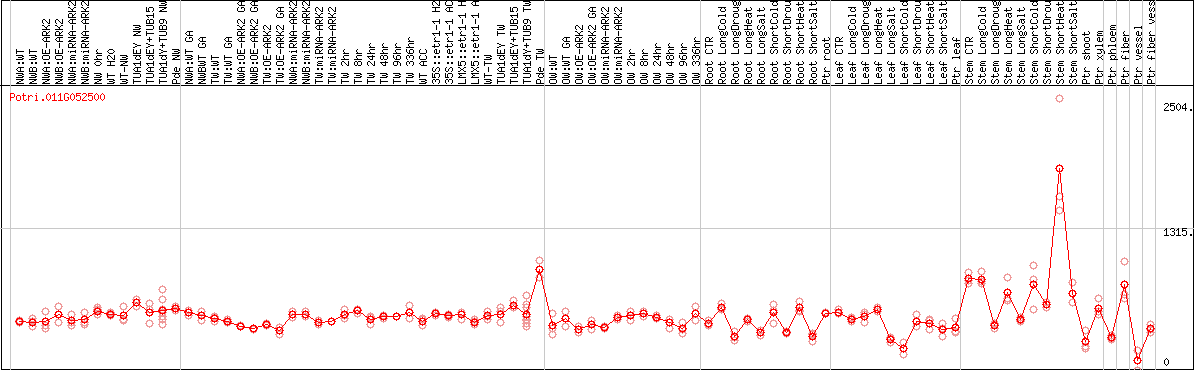
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G052500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
