External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G33820 460 / 8e-165
ATMBAC1
Mitochondrial substrate carrier family protein (.1)
AT5G46800 172 / 8e-52
BOU
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
AT1G79900 123 / 4e-33
ATMBAC2, BAC2
RABIDOPSIS MITOCHONDRIAL BASIC AMINO ACID CARRIER 2, Mitochondrial substrate carrier family protein (.1)
AT5G66380 103 / 1e-25
ATFOLT1
folate transporter 1 (.1)
AT5G01340 89 / 4e-20
AtmSFC1
mitochondrial succinate-fumarate carrier 1, Mitochondrial substrate carrier family protein (.1)
AT4G11440 85 / 3e-18
Mitochondrial substrate carrier family protein (.1)
AT1G34065 84 / 4e-18
SAMC2
S-adenosylmethionine carrier 2 (.1)
AT1G14140 80 / 5e-17
Mitochondrial substrate carrier family protein (.1)
AT5G58970 79 / 8e-17
ATUCP2
uncoupling protein 2 (.1.2)
AT1G25380 79 / 1e-16
ATNDT2
NAD+ transporter 2, ARABIDOPSIS THALIANA NAD+ TRANSPORTER 2, NAD+ transporter 2 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G046801
575 / 0
AT2G33820 453 / 4e-162
Mitochondrial substrate carrier family protein (.1)
Potri.001G142600
166 / 2e-49
AT5G46800 434 / 1e-154
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
Potri.003G091500
160 / 3e-47
AT5G46800 443 / 4e-158
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
Potri.001G182000
126 / 6e-34
AT1G79900 424 / 1e-150
RABIDOPSIS MITOCHONDRIAL BASIC AMINO ACID CARRIER 2, Mitochondrial substrate carrier family protein (.1)
Potri.003G053900
120 / 6e-32
AT1G79900 419 / 2e-148
RABIDOPSIS MITOCHONDRIAL BASIC AMINO ACID CARRIER 2, Mitochondrial substrate carrier family protein (.1)
Potri.007G019000
110 / 4e-28
AT5G66380 463 / 8e-166
folate transporter 1 (.1)
Potri.012G066000
84 / 3e-18
AT1G74240 446 / 6e-158
Mitochondrial substrate carrier family protein (.1)
Potri.016G119500
83 / 5e-18
AT5G01340 542 / 0.0
mitochondrial succinate-fumarate carrier 1, Mitochondrial substrate carrier family protein (.1)
Potri.008G100100
79 / 2e-16
AT5G15640 338 / 5e-116
Mitochondrial substrate carrier family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015420
465 / 5e-167
AT2G33820 415 / 7e-147
Mitochondrial substrate carrier family protein (.1)
Lus10013998
268 / 3e-90
AT2G33820 215 / 3e-69
Mitochondrial substrate carrier family protein (.1)
Lus10004565
172 / 7e-52
AT5G46800 481 / 2e-173
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
Lus10000906
170 / 5e-51
AT5G46800 482 / 1e-173
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
Lus10011451
125 / 2e-33
AT1G79900 410 / 4e-145
RABIDOPSIS MITOCHONDRIAL BASIC AMINO ACID CARRIER 2, Mitochondrial substrate carrier family protein (.1)
Lus10037544
125 / 2e-33
AT1G79900 408 / 2e-144
RABIDOPSIS MITOCHONDRIAL BASIC AMINO ACID CARRIER 2, Mitochondrial substrate carrier family protein (.1)
Lus10027817
86 / 4e-19
AT5G01340 526 / 0.0
mitochondrial succinate-fumarate carrier 1, Mitochondrial substrate carrier family protein (.1)
Lus10026395
86 / 9e-19
AT2G30160 483 / 5e-173
Mitochondrial substrate carrier family protein (.1)
Lus10029770
84 / 4e-18
AT5G42130 478 / 5e-169
MitoFerrinLike1, Mitochondrial substrate carrier family protein (.1)
Lus10042258
83 / 6e-18
AT2G30160 484 / 2e-173
Mitochondrial substrate carrier family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00153
Mito_carr
Mitochondrial carrier protein
Representative CDS sequence
>Potri.011G056000.1 pacid=42781761 polypeptide=Potri.011G056000.1.p locus=Potri.011G056000 ID=Potri.011G056000.1.v4.1 annot-version=v4.1
ATGGGAGAGACATTCGCTTACAAGGAATACTTAGCCGGTTTACTTGCTGGTGTTGCTACTGTTATCACTGGTCACCCTTTTGACACTGTCAAGGTGATGC
TACAGAAACATAATACAGAAGCACATGGGATTAAGTACAAGAATGGTTTGCATTGCACGGCTAGGATACTACAGACTGAAGGAGTTAAAGGACTGTATAG
AGGGGCAACATCATCTTTTGTTGGAGTGGCTTTTGAGAGTTCCCTTCTTTTTGGCATTTATTCCCAAACAAAGCAGTCATTGCAGGGAGGAGGTCAAAGT
GATGTGCCCCGACCCCAAGTAATAATTCCATCAGCAGCATATGGCGGAGCTATTATTAGTTTTGTATTATGCCCATCAGAGCTAGTAAAATGTAGGATGC
AAATTCAAGGCACTGACTCCTTGGTTCCAAAGTTCAGTAGATACAGTAGCCCACTTGATTGTGCCCTCCAAACAATGAAAAATGAAGGGGTTACAGGCAT
TTTTCGTGGAGGTTTTACAACATTGTTGAGAGAATCTATTGGAAGTGCTGTCTTCTTTAGTGTTTATGAGTATGTCCGTTATTACATGCATTTACAATTG
AAACCCACTTTATCTGACCACAGCAATTTAACTGACATGGGGATCGGAATTGTGACCGGTGGCCTTAGTGGCGTAGCTTTTTGGTCTGCTGTTTTGCCCT
TGGATGTGGCGAAAACTATCATCCAGACAGCTCCAGATAAAAGCTCCACAAGAAATCCATTTGCAGTTCTGAATTCTATCTACAGCAGGGCTGGACTTAA
AGGATGCTATACAGGTTTCGGTCCGACAATAGTGAGAGCATTTCCTGCTAATGCCGCTGCAATAGTCACCTGGGAGCTAGCCATGAAAATGCTAGGGGAT
AAATCGTGA
AA sequence
>Potri.011G056000.1 pacid=42781761 polypeptide=Potri.011G056000.1.p locus=Potri.011G056000 ID=Potri.011G056000.1.v4.1 annot-version=v4.1
MGETFAYKEYLAGLLAGVATVITGHPFDTVKVMLQKHNTEAHGIKYKNGLHCTARILQTEGVKGLYRGATSSFVGVAFESSLLFGIYSQTKQSLQGGGQS
DVPRPQVIIPSAAYGGAIISFVLCPSELVKCRMQIQGTDSLVPKFSRYSSPLDCALQTMKNEGVTGIFRGGFTTLLRESIGSAVFFSVYEYVRYYMHLQL
KPTLSDHSNLTDMGIGIVTGGLSGVAFWSAVLPLDVAKTIIQTAPDKSSTRNPFAVLNSIYSRAGLKGCYTGFGPTIVRAFPANAAAIVTWELAMKMLGD
KS
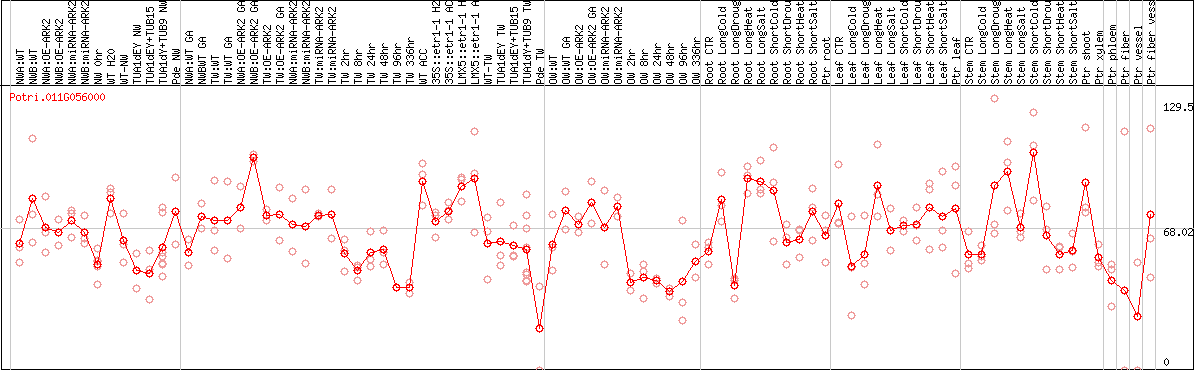
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G056000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.