Pt-TPS7.1 (Potri.011G070900) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pt-TPS7.1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.011G070900.4 pacid=42780894 polypeptide=Potri.011G070900.4.p locus=Potri.011G070900 ID=Potri.011G070900.4.v4.1 annot-version=v4.1
ATGATGTCTAGATCATATACCAATCTATTAGATCTAGCTTCTGGTAACTTTCCGGCAATGGGTCAACCTCGGGAGAGAAAGCGGTTGCCTCGTGTAATGA
CAGTTCCGGGGGTTATTTCTGAGCTTGATGATGATGTGGCTAATAGTGTGACCTCAGATGTACCTTCATCGGTAGTTCAGGACCGGATAATCATTGTAGG
GAATCAGTTGCCGGTAAAAGCAAAGCGCAGACCGGATAATAAAGGATGGAGCTTTAGTTGGGATGAGGATTCGTTGTTATTGCAGTTAAAAGATGGCTTG
CCTGAAGAAATGGAGGTTTTATATGTCGGTTCTTTGCGGGCAGATATTGATTTGAGTGAACAAGAGGATGTCTCACAGATCTTGTTGGACAGGTTCAAGT
GTGTACCAGCATTTCTGCCTCCTGATATTTTGTCCAAGTTTTATCATGGGTTTTGTAAACAGTATTTGTGGCCACTTTTTCATTACATGCTTCCTATTTC
GGGTAATCATGGTGGGCGATTTGATCGGTCTTTGTGGGAGGCATATGTGGCAGCAAATAAGATCTTCTCGCAAAGGGTGATTGAAGTTATAAATCCAGAG
GATGACTATGTTTGGATTCATGACTATCATTTGATGGTGCTGCCTACATTCTTGAGGAGAAGGTTCAATAGATTGAGAATGGGATTCTTTCTTCACAGTC
CATTTCCTTCATCTGAGATATATAGGACTTTGCCTGTTAGGGAAGAGATCCTTAAGGCACTGTTGAATTCGGACCTCATTGGTTTTCACACTTTTGATTA
TGCCCGACATTTTCTATCTTGTTGCAGTCGGATGTTGGGGTTGGAGTATCAGTCAAAGAGGGGTTACATTGGGTTAGAGTATTATGGCAGGACTGTTGGA
ATAAAGATCATGCCAGTTGGGATACATATGGGGCAGATTCAGTCTGTCTTGAAACTTGCGGATAAGGATTGGAGGGTAGAAGAGCTCAAACAACAGTTTG
AAGGAAAGACGGTTTTACTTGGGGTTGATGATATGGACATCTTTAAAGGTGTTAATTTGAAGCTTTTGGCAATGGAGCAGCTGCTGAAACAGCACCCAAA
GTGGCAACGAAGGGCTGTTTTGGTGCAGATAACTAATCCTGCCAGGGGACGAGGGAGAGATCTTGAGGAAGTGCAAGCTGAAATACAGGAAAGCTGCAGG
AGAATTAATGAGACTTTTGGTCGACCTGGCTATGAACCAGTTGTCTTTATTGATCGACCAGTATCTCTCAGTGAAAGATCTGCATATTTCACCATTGCTG
AGTGTGTTGTTGTGGCAGCTGTGAGAGATGGGATGAATCTCACTCCTTATGAGTATATTGTGTGCAGACAGGGAGTTTCTGGGTCAGAGTCTAGCTCAGG
ATCAAGTGGCCCCAAGAAGAGCATGCTGGTTGTATCAGAGTTCATTGGATGTTCCCCTTCACTCAGTGGTGCAATTCGAGTCAATCCATGGAACATTGAA
GCAACCGCAGAAGCAATGAATGAGGCAATTTCAATGGCAGATTCCGAGAAACAATTGCGCCATGAAAAGCACTACAGGTATGTCAGCACGCATGATGTGG
CATATTGGTCAAGGAGTTTCTATCAAGATATGGAGCGGACTTGCAAAGACCATTTCAGAAGGCGCTGCTGGGGAATTGGCTTGAGCTTTGGTTTCAGAGT
TGTGGCGCTTGATCCTAATTTCAAAAAATTAAATATAGATCAAATTGAATCTGCATATATAAAGTCCAAAAACAGAGCTATACTCTTGGACTATGATGGA
ACTGTAATGCCTCAAACCACCATCAATAAGACCCCAAACCAAGAGGTCATTTCAATCATAAATACTCTTTGTAGTGATGTCAAGAACACCGTCTTTGTTG
TCAGTGGAAGGGGAAGGGATAGTTTAGGGAAGTGGTTTGCTCATTGCAAGAAACTTGGAATTGCTGCTGAGCATGGGTACTTCATGAGGTGGTCTGTGGA
TGAGGACTGGGAGAACTGCGGGCAGAGTAGTGATTTTGGATGGACTCAAATAGCTGAACCTGTTATGAATCTGTACACAGAAGCCACTGATGGATCTTCC
ATCGAGACGAAAGAGAGCGCTTTGGTTTGGCACCACAGAGATGCAGATCCAGGTTTTGGAGCTGCTCAGGCCAAGGAATTGTTGGACCATCTTGAGAGCG
TGCTTGCAAATGAGCCTGTTGCAGTCAAAAGTGGTCAATGCATTGTGGAAGTTAAGCCCCAGGGAATTAGTAAAGGTTCTGTGGCAGAAAAGATCTTTAC
ATCAATGGCTGAAAGTGGAAGACAGGCTGATTTTGTATTGTGTATTGGGGATGACAGATCAGATGAGGATATGTTTGAAAGCATTGATAATGCAATAGCG
AACGGTATCCTAACCTCAAGCAAATCTGTCTTCGCTTGCACTGTTGGTCAGAAACCAAGCAAAGCCAAGTATTATTTGGATGACACAACTGATGTCATAA
ACATGCTCGAAGCTCTTGCAGAAGCTTCCGACCCTTCACCCTCTGCAGGAAGCTCCCCCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.011G070900.4 pacid=42780894 polypeptide=Potri.011G070900.4.p locus=Potri.011G070900 ID=Potri.011G070900.4.v4.1 annot-version=v4.1
MMSRSYTNLLDLASGNFPAMGQPRERKRLPRVMTVPGVISELDDDVANSVTSDVPSSVVQDRIIIVGNQLPVKAKRRPDNKGWSFSWDEDSLLLQLKDGL
PEEMEVLYVGSLRADIDLSEQEDVSQILLDRFKCVPAFLPPDILSKFYHGFCKQYLWPLFHYMLPISGNHGGRFDRSLWEAYVAANKIFSQRVIEVINPE
DDYVWIHDYHLMVLPTFLRRRFNRLRMGFFLHSPFPSSEIYRTLPVREEILKALLNSDLIGFHTFDYARHFLSCCSRMLGLEYQSKRGYIGLEYYGRTVG
IKIMPVGIHMGQIQSVLKLADKDWRVEELKQQFEGKTVLLGVDDMDIFKGVNLKLLAMEQLLKQHPKWQRRAVLVQITNPARGRGRDLEEVQAEIQESCR
RINETFGRPGYEPVVFIDRPVSLSERSAYFTIAECVVVAAVRDGMNLTPYEYIVCRQGVSGSESSSGSSGPKKSMLVVSEFIGCSPSLSGAIRVNPWNIE
ATAEAMNEAISMADSEKQLRHEKHYRYVSTHDVAYWSRSFYQDMERTCKDHFRRRCWGIGLSFGFRVVALDPNFKKLNIDQIESAYIKSKNRAILLDYDG
TVMPQTTINKTPNQEVISIINTLCSDVKNTVFVVSGRGRDSLGKWFAHCKKLGIAAEHGYFMRWSVDEDWENCGQSSDFGWTQIAEPVMNLYTEATDGSS
IETKESALVWHHRDADPGFGAAQAKELLDHLESVLANEPVAVKSGQCIVEVKPQGISKGSVAEKIFTSMAESGRQADFVLCIGDDRSDEDMFESIDNAIA
NGILTSSKSVFACTVGQKPSKAKYYLDDTTDVINMLEALAEASDPSPSAGSSP
|
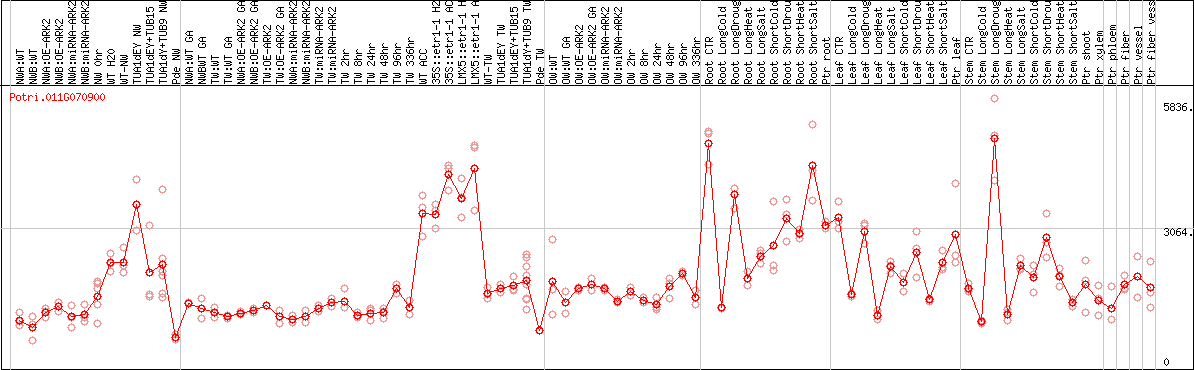
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G070900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
