External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G34460 371 / 5e-130
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G18890 111 / 3e-27
AtTic62
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT4G31530 65 / 1e-11
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT2G37660 57 / 4e-09
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT5G18660 55 / 2e-08
PCB2, DVR
PALE-GREEN AND CHLOROPHYLL B REDUCED 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT4G18810 53 / 2e-07
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT3G46780 50 / 1e-06
PTAC16
plastid transcriptionally active 16 (.1)
AT1G09480 45 / 4e-05
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G33590 44 / 5e-05
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT1G32100 44 / 6e-05
ATPRR1
pinoresinol reductase 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G150300
110 / 7e-27
AT3G18890 481 / 1e-164
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.T124404
99 / 8e-23
AT3G18890 541 / 0.0
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.009G111323
99 / 8e-23
AT3G18890 541 / 0.0
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.008G204900
67 / 2e-12
AT5G18660 610 / 0.0
PALE-GREEN AND CHLOROPHYLL B REDUCED 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.009G037000
60 / 6e-10
AT3G46780 503 / 2e-175
plastid transcriptionally active 16 (.1)
Potri.001G244800
59 / 2e-09
AT3G46780 513 / 7e-179
plastid transcriptionally active 16 (.1)
Potri.006G086800
52 / 2e-07
AT2G37660 450 / 2e-160
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.001G253900
51 / 5e-07
AT4G31530 399 / 1e-139
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.004G061200
47 / 7e-06
AT4G18810 812 / 0.0
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10040884
394 / 2e-138
AT2G34460 401 / 7e-142
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10004936
387 / 8e-136
AT2G34460 399 / 4e-141
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10042346
107 / 9e-26
AT3G18890 483 / 2e-164
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10026321
104 / 1e-24
AT3G18890 479 / 6e-163
translocon at the inner envelope membrane of chloroplasts 62, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016502
59 / 1e-09
AT3G46780 533 / 0.0
plastid transcriptionally active 16 (.1)
Lus10040768
58 / 4e-09
AT3G46780 540 / 0.0
plastid transcriptionally active 16 (.1)
Lus10037912
50 / 1e-06
AT5G18660 577 / 0.0
PALE-GREEN AND CHLOROPHYLL B REDUCED 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10019734
49 / 3e-06
AT1G09480 286 / 4e-96
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10022632
48 / 4e-06
AT4G35250 640 / 0.0
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10007308
49 / 5e-06
AT4G18810 850 / 0.0
NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF13460
NAD_binding_10
NAD(P)H-binding
Representative CDS sequence
>Potri.011G079600.1 pacid=42781630 polypeptide=Potri.011G079600.1.p locus=Potri.011G079600 ID=Potri.011G079600.1.v4.1 annot-version=v4.1
ATGGCCACTGCTTTGGTCATAAAAACTCGTCTTCTCTCTGCACCACGCCTCCATCACCATCACCACTGTTACCCTCTTCTCTTCTCTACCACATCCCCCA
AACCTCCCAAGTCTCAATCCCTCAAATCCATCAAGATGGAAAGTGGAAATGAAATTACGGAAGAAGCTAAAGAGAATGTGAATCAGAAGAAGAAGAAAAT
ATTTGTGGCTGGAGCCACTGGAAGTACTGGGAAAAGGATTGTTGAACAGCTTCTAGCCAAGGGTTTTGAGGTCAAGGCTGGTGTTAGGGATTTAGACAAG
GCCAAAACTATTCTTTCTGAACACAACCCATCTCTTCAAATTGTGACAGCCGATGTAACTAAGGGGTCTGACAAGCTAGTACAGGCCATAGGGGATGATT
CCGAGGCTGTAATATGTGCCACTGGGTTTCGGCCTGGCTGGAACTTGTTTGCTCCATGGAAGGTTGATAATTTAGGCACTGTGAACCTTGTTGAAGCTTG
CCGTAAACTTGGTGTGAAAAGATTCATTCTAATAAGCTCAATTTTAGTCAATGGAGCTGCAATGGGACAGATACTCAATCCAGCTTACATTTTTCTCAAT
GTTTTTGGGCTAACCCTAGTGGCGAAGCTTCAGGCAGAGAATTACATCAGGAAGTCTGGCATAAACTACACTATAGTTAGACCAGCTGGTTTGAAAAATG
AACCACCCTCAGGAAATCTTGTAATTGAGCCAGAGGATACTCTTTATGAAGGTATCATATCAAGGGATGTGGTTGCGGAAGTTGCTGTTGAGGCACTAGG
TCTTCCGGAATCATCTTACAAAGTCGTGGAAATAGTCTCTCGTGCTGACGCTCCGAAGCGTACATACGAGGATCTTTTTGGTTCTATCAAGCAAAAGTGA
AA sequence
>Potri.011G079600.1 pacid=42781630 polypeptide=Potri.011G079600.1.p locus=Potri.011G079600 ID=Potri.011G079600.1.v4.1 annot-version=v4.1
MATALVIKTRLLSAPRLHHHHHCYPLLFSTTSPKPPKSQSLKSIKMESGNEITEEAKENVNQKKKKIFVAGATGSTGKRIVEQLLAKGFEVKAGVRDLDK
AKTILSEHNPSLQIVTADVTKGSDKLVQAIGDDSEAVICATGFRPGWNLFAPWKVDNLGTVNLVEACRKLGVKRFILISSILVNGAAMGQILNPAYIFLN
VFGLTLVAKLQAENYIRKSGINYTIVRPAGLKNEPPSGNLVIEPEDTLYEGIISRDVVAEVAVEALGLPESSYKVVEIVSRADAPKRTYEDLFGSIKQK
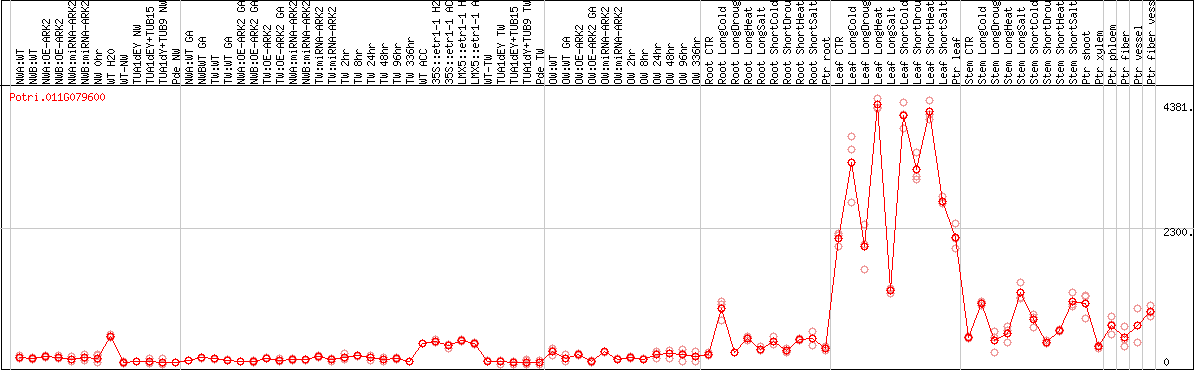
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G079600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.