Potri.011G082300 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.011G082300.2 pacid=42781685 polypeptide=Potri.011G082300.2.p locus=Potri.011G082300 ID=Potri.011G082300.2.v4.1 annot-version=v4.1
ATGCTATATAATTTCGTTCCTTACTCCAAATTCCCAAAACTTGTAGCCTATATCCCTTTCTTTCACTTTGTTCATACTGCCAAAAAGCATTACCGCACCA
CCCCATTTCCCTATATTCATATGTTTCTTTCACTTTTTTCTTCAACAAATCATGGTCCTCATTCTTATTTTGATTCTGATTCTTCAACTACTAATTCCAC
TTCCCATCTTGTTGAAACTGAGAATTCTTATGATGCTAATGATGGGTTTGGAAAAATGGGTTTTAATTTTAGCAATGCTCACTGTCAAGTTTATTCTGAT
GGTTTTAGTGATGAGGTTGAAGAATCTGATGAAGAACGCGGCCAAGGTAACAGTTCATGTACTGATAGGAATCAAGGCACCAAAATGGATGAGGATTGGA
GGAGAATTGAGATACACGAAGATGAATTTCGACATCCTTTGGTCAGAGAAATTTGTAGGTTGATTGAATGCAGGTCGGCTTGGAACCATAAGCTTGAAGG
GAAAATGAGGCACTTGCTAAGAGGCCTCAAGCCTCGACTTGTTTGTGCTGTTTTGCTGTCTCAATCTGATGAAAGGGTTGCTTTGGATTTCTTCTTTTGG
TCCGACAGGCAATGGCGGTATCGGCATGATCCAATTGTGTACTGTGTGATGCTTGATGTTCTTAGCAAAACTAAACTGTGTCAAGGTGCTAGGAGGGTTC
TTCGGCTCATGGTACGTAGGGGAATTCAGCGTACGCCTCAAGATTTTTGTTGTGTGATGGTGTCTTATAGTCGGGCTGGTAAGTTGAGGAATGCAATGCA
GGTCTTGACCATGATGCAGAAAGCAGGTATTGAACCTAATTTGTTGGTTTGTAACACTGCTATTCATGTTTTGGTGATGGCAAATATGTTGGAAAAGGCT
TTGAGATTTCTAGAGAGAATGCAACTTCTTGGAATCATGCCAAATGTTGTCACTTATAACTGTTTGATCAAGGGATATTGTGATCTTCATAGGGTTGAAG
ATGCCATGGAGCTAATTTCTGAAATGCCATTGAAAGGATGCTCCCCAGATAAAGTTAGTTACTATACTGTAATGGGCTTCCTTTGCAAAAACAGAAGGAT
CAGAGAAGTGATGGACGTAATAGAGAAAATGGAGGATACTAAGTTGTTAGCGGATCAGGTTACTTATAATACATTAATACATATGCTTTGTAAGCATCAG
CATGCAGATGAGGCACTTCAGTTTTTAAGAGAAGCACAAAAAAGGGGTTTTCAGGTTGATAAGGTTGGGTATAGTGCGATAGTTGATTCATATTGTAAGG
AGGGAAGGATGGACCAGGCGAAAGAAATTGTGAATGAAATGTTCACAAGGGGCTGCATCCCTGATGTTGTGACTTACACAGCAATTATTAATGGATTTTC
TCAGGCTGGGGAGGTTGGTCAAGCTAGGAAGATGCTTCAGCAAATGTACAAGCATGGCTGCAAACCAAACACTGTATCATATACAGCCTTCTTAAAAGGG
CTTTGTCAAAAGGGGAACTCATCAGAGGCAAGGGAGATGATGAAGGCGAGTGAGGAACAGTGGTGGACCCCAAATGCTATTACCTATAGTGTTGTGATGC
ATGGGTTTCGGAGGGAAGGAAAGCTGTCTGATGCTTGTGATGTGGTGAGGGAGATGATTGGCAAGGGCTTTTTCCCAACTCCAGTTGAAATTAACTTACT
ACTTCAGTCTCTCTGCCGGATCGGCAGAGTGGATGAGGCAAAAAAGTTTATGGAGGAGTGTCTTAACATGGGGTGTGCTGTCAATGCTGTAAACTTTACT
ACTGTAATTCACAGATTTTGTCAGCAGGATGACATAGAAGCAGCTTTATCTTTGCTTGATGACATGTATCTAAGCAACAAACACCCTGATGCAGTCACTT
ATACGACGATAATTGATGCATTAGGAAAGAAAGGTAGAATAGAAGAGGCCACTGAGCTTACGTTGAAGATGCTGAAGAAGGGTATAGATCCTACACCAGT
AACATATAGAACAGTGATTCATCGATATGGTCAAATAGGTAGGGTGGAAGATTTATTAAATTTATTAGATAAAATGCTTACAAGGCAAGAGTGTAGAACT
GCATTTAATCAAGTTATTGAGAAGCTATGTACTTTTGGAAACCTAGAGGCGGCTGATAAACTCTTAGGTAAGGTTTTGAGAACAGCCTCAAGAATTGATG
CTAACACATGCCATGTGCTAATGGAGAGTTATTTGAGAAAAGGGATCCCTCTGTCAGCATATAAAGTGGCTTGTCGTATGTTTAGTCGAAGTCTGATTCC
TGATTTAAAGTTGTGTGAGAAGGTGTGCAAGAAACTAATGCAGGAAGGAAAGTCAGAGGAGGCAGACAATCTGTTCTTACGGTTTGTTGAGCGTGGGAAT
ATTTCATCGCATTGTCTACGGCATTCTCAAACCGAGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.011G082300.2 pacid=42781685 polypeptide=Potri.011G082300.2.p locus=Potri.011G082300 ID=Potri.011G082300.2.v4.1 annot-version=v4.1
MLYNFVPYSKFPKLVAYIPFFHFVHTAKKHYRTTPFPYIHMFLSLFSSTNHGPHSYFDSDSSTTNSTSHLVETENSYDANDGFGKMGFNFSNAHCQVYSD
GFSDEVEESDEERGQGNSSCTDRNQGTKMDEDWRRIEIHEDEFRHPLVREICRLIECRSAWNHKLEGKMRHLLRGLKPRLVCAVLLSQSDERVALDFFFW
SDRQWRYRHDPIVYCVMLDVLSKTKLCQGARRVLRLMVRRGIQRTPQDFCCVMVSYSRAGKLRNAMQVLTMMQKAGIEPNLLVCNTAIHVLVMANMLEKA
LRFLERMQLLGIMPNVVTYNCLIKGYCDLHRVEDAMELISEMPLKGCSPDKVSYYTVMGFLCKNRRIREVMDVIEKMEDTKLLADQVTYNTLIHMLCKHQ
HADEALQFLREAQKRGFQVDKVGYSAIVDSYCKEGRMDQAKEIVNEMFTRGCIPDVVTYTAIINGFSQAGEVGQARKMLQQMYKHGCKPNTVSYTAFLKG
LCQKGNSSEAREMMKASEEQWWTPNAITYSVVMHGFRREGKLSDACDVVREMIGKGFFPTPVEINLLLQSLCRIGRVDEAKKFMEECLNMGCAVNAVNFT
TVIHRFCQQDDIEAALSLLDDMYLSNKHPDAVTYTTIIDALGKKGRIEEATELTLKMLKKGIDPTPVTYRTVIHRYGQIGRVEDLLNLLDKMLTRQECRT
AFNQVIEKLCTFGNLEAADKLLGKVLRTASRIDANTCHVLMESYLRKGIPLSAYKVACRMFSRSLIPDLKLCEKVCKKLMQEGKSEEADNLFLRFVERGN
ISSHCLRHSQTE
|
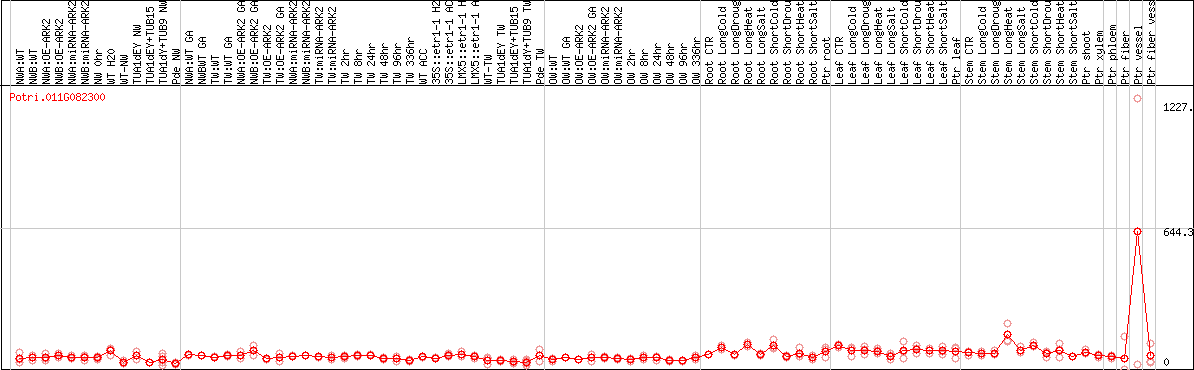
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G082300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
