External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G23590 254 / 5e-85
ATMES8
methyl esterase 8 (.1)
AT2G23620 253 / 7e-85
ATMES1
ARABIDOPSIS THALIANA METHYL ESTERASE 1, methyl esterase 1 (.1)
AT2G23600 249 / 4e-83
ATMES2, ACL, ATME8
ARABIDOPSIS THALIANA METHYL ESTERASE 2, ARABIDOPSIS METHYL ESTERASE 8, acetone-cyanohydrin lyase (.1)
AT2G23610 239 / 3e-79
ATMES3
ARABIDOPSIS THALIANA METHYL ESTERASE 3, methyl esterase 3 (.1)
AT4G37150 217 / 1e-70
ATMES9
ARABIDOPSIS THALIANA METHYL ESTERASE 9, methyl esterase 9 (.1)
AT5G10300 215 / 8e-70
AtHNL, HNL, ATMES5
HYDROXYNITRILE LYASE, ARABIDOPSIS THALIANA METHYL ESTERASE 5, methyl esterase 5 (.1)
AT2G23560 213 / 7e-69
ATMES7
ARABIDOPSIS THALIANA METHYL ESTERASE 7, methyl esterase 7 (.1)
AT2G23580 205 / 6e-66
ATMES4, ABE4
ARABIDOPSIS THALIANA METHYL ESTERASE 4, ALPHA/BETA FOLD HYDROLASE/ESTERASE 4, methyl esterase 4 (.1.2)
AT2G23550 201 / 2e-64
ATMES6, ABE1
ALPHA/BETA FOLD HYDROLASE/ESTERASE 1, methyl esterase 6 (.1.2)
AT3G50440 185 / 1e-57
ATMES10
ARABIDOPSIS THALIANA METHYL ESTERASE 10, methyl esterase 10 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038592
222 / 3e-72
AT2G23620 306 / 4e-105
ARABIDOPSIS THALIANA METHYL ESTERASE 1, methyl esterase 1 (.1)
Lus10022467
215 / 1e-69
AT3G50440 238 / 2e-78
ARABIDOPSIS THALIANA METHYL ESTERASE 10, methyl esterase 10 (.1)
Lus10015532
208 / 4e-67
AT2G23600 240 / 2e-79
ARABIDOPSIS THALIANA METHYL ESTERASE 2, ARABIDOPSIS METHYL ESTERASE 8, acetone-cyanohydrin lyase (.1)
Lus10023326
183 / 2e-57
AT3G50440 263 / 3e-88
ARABIDOPSIS THALIANA METHYL ESTERASE 10, methyl esterase 10 (.1)
Lus10005402
176 / 2e-54
AT2G23580 204 / 3e-65
ARABIDOPSIS THALIANA METHYL ESTERASE 4, ALPHA/BETA FOLD HYDROLASE/ESTERASE 4, methyl esterase 4 (.1.2)
Lus10009205
173 / 1e-53
AT2G23580 194 / 3e-61
ARABIDOPSIS THALIANA METHYL ESTERASE 4, ALPHA/BETA FOLD HYDROLASE/ESTERASE 4, methyl esterase 4 (.1.2)
Lus10009203
171 / 5e-52
AT2G23620 205 / 6e-65
ARABIDOPSIS THALIANA METHYL ESTERASE 1, methyl esterase 1 (.1)
Lus10005400
162 / 4e-49
AT2G23610 206 / 8e-66
ARABIDOPSIS THALIANA METHYL ESTERASE 3, methyl esterase 3 (.1)
Lus10005401
164 / 7e-49
AT2G23600 184 / 5e-56
ARABIDOPSIS THALIANA METHYL ESTERASE 2, ARABIDOPSIS METHYL ESTERASE 8, acetone-cyanohydrin lyase (.1)
Lus10029011
149 / 6e-44
AT3G10870 293 / 1e-99
ARABIDOPSIS THALIANA METHYL ESTERASE 17, methyl esterase 17 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0028
AB_hydrolase
PF12697
Abhydrolase_6
Alpha/beta hydrolase family
Representative CDS sequence
>Potri.011G082400.1 pacid=42781978 polypeptide=Potri.011G082400.1.p locus=Potri.011G082400 ID=Potri.011G082400.1.v4.1 annot-version=v4.1
ATGGGAGACATCAACAACCAAGCAGCCCATCTTGTTTTAATACACGGTTCTTCCGCAGGAGCTTGGGTGTGGTACAAGGTGAAGCCAATGCTAGAGGCAG
CCGGCCACAGCATCACAGCTCTTGACATGAGTGCATCAGGGGTGAACACAAAAACACTCGAAGAAGTTCGAACTTTTGACCAGTATAACGAGCCCTTGAT
TGAGTTCATGGCCAACTTACCTGAAAATGAAAAGGTTGTATTGGTGGGCCATAGTTTGGGTGGCTTGAACCTGGCCTTCGCCATGGAGAAGTTCCCAGAG
AAGATTTCTCTTGCTGTTTTTGTTACTGCAATCTTGCCTGATACTCAGCACCAGCCATCGTATATGTTAGAGAAGTTTATTGAAAGCATTTCCGGTGCAG
ACGAAGAGCAAGACACTGCAGTGGTTTCATCCACGCCGTTTCAGCTCACCCCAATTGAGGATCTTACTCTCCAAGCGCTTTTAAACAGACCAGGATCAAT
GTTCGTTGAAAGTCTGTCCAAAGCAAACAAGTTCACTGAAGACAGATATGGATCCGTTCCCCGAGTTTATATTGTCTGTACTGAGGATATATTGCTTTCT
CCGTCGTTGCAACGCTACATGATTGAACAAAATGAGGTTAAGGAAGTGATGGAGATTCCTGCAGATCATATGGCAGTCTTTTCTAAGCCTAAGGAACTCA
GCCAGTGCATACTGGAGTTGGCACAGAAGCACGCTTAG
AA sequence
>Potri.011G082400.1 pacid=42781978 polypeptide=Potri.011G082400.1.p locus=Potri.011G082400 ID=Potri.011G082400.1.v4.1 annot-version=v4.1
MGDINNQAAHLVLIHGSSAGAWVWYKVKPMLEAAGHSITALDMSASGVNTKTLEEVRTFDQYNEPLIEFMANLPENEKVVLVGHSLGGLNLAFAMEKFPE
KISLAVFVTAILPDTQHQPSYMLEKFIESISGADEEQDTAVVSSTPFQLTPIEDLTLQALLNRPGSMFVESLSKANKFTEDRYGSVPRVYIVCTEDILLS
PSLQRYMIEQNEVKEVMEIPADHMAVFSKPKELSQCILELAQKHA
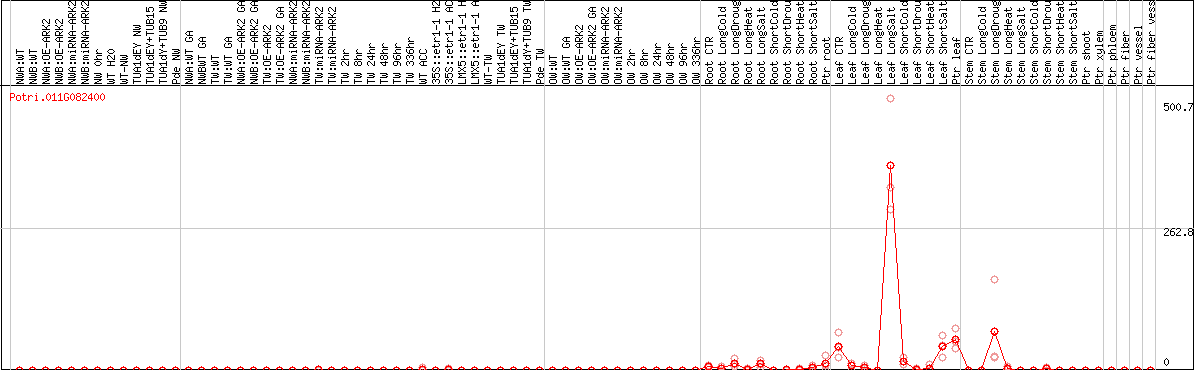
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G082400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.