Potri.011G090200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.011G090200.3 pacid=42782104 polypeptide=Potri.011G090200.3.p locus=Potri.011G090200 ID=Potri.011G090200.3.v4.1 annot-version=v4.1
ATGGTATGCTGTCTTGCTGTTGTGGTGGTTGAATCAGGCTACCTTAGCTTAAAAATGGAGGAACCTTTAAGTGACGAGGCGGCTAAAAAGTCTCCGACTT
CAGACAAACAGAGGCAGCAGTCGTCTGTTAGCTCAATGCAAATCCCTATGAAGGTTTCGAAAACCACAAAACCTACTATTTCAGCCAATTCCCACCTCTT
AACACCAATTGGTTCTATTCGAAAGAGGACTGAACCAAAAAACAGTTCAGATTCAAGTTCAAATGTGACCGCCAAAAACGCATCCTCCTGCAATACCAAG
TCAGTTCCAATCGCCAGAAGAAACAGCACTGGAGGGGTTCCTGAGAAGCAACCTGTCTCTTCAACAAAACGGCAGAATACCAGCGGAAAGACTAATGCTG
TTTCGGACCCAGTGAGGAGGTCCCTTCCAGAGCTTAGAAGGAGTTCCTTGCCTCCTACAAAACCCATGGTTAGAACTGGTAGTGTTTCCGAGACTAGGAA
CTCAGTTCCCATGGACAAGTGTTTGAGAGCATCAACTGGGTCAGGTGTAAGTAGGCTGGAGAAGCCTTCAGTTAAGCCAGCATTGCCAGCCTCTTCTTCT
TCTTCTTCTTCCAGAAGGGTTATTTCCACATCAGTGGACAGTACTGCCAGTAGCATGAGCAGGAAGAAGCTTTCTTCTCCATCTGCTACGTCCCCTTCTA
TTTCTAGTGGTTTGAGAGCCGGGTCCCTATCAACTTCTCGGGATAGGAGTTTCAATTTGACTGGACGGAGAAGAGCAGGTGCTCCTGAAAGCCACGATTC
TCACTTCATTGCTCTTCCACTGGTGGAAACTAAAGCTGGCGATGATGTGAGATTGGATCTTAGAGGACACAAAGTACGTAGTCTCAATGCTAGTGGGTTG
AACTTAGCCCAAAATTTAGAGTTTGTTTATCTTCGAGACAATCTCCTTTCTACATTGGAAGGAATTGAGATACTGAAGCGTGTGAAGGTCCTTGATTTGA
GTTTCAATGAGTTCAAGGGGCCTGGGTTTGAACCTCTTGAGAATTGCCAGGCCCTTCAGCAACTCTATCTTGCTGGGAATCAAATCACATCACTTGTAAA
CCTTCCTCAGCTTCCTAACCTGGAGTTTCTTTCGGTCGCTCAAAACAAGTTAAAGTCTCTGTCAATGGCAGGCCAACCTCGACTACAGGTGTTAGCTGCC
AGCAAAAACAAAATAACAACTCTGAAGGGTTTCCCGCACCTACCATCCCTCGAGCATTTGCGAGTGGAGGAGAATCCTATACTTAAGATGCCTCATTTAG
AAGCCGCCTCGATTTTGCTTGTTGGTCTTACCTTGAAAAAGTTTAATGATAGAGATCTTTCCCGGGAAGAGGTTGCAATTGCAAAGCGTTATCCAGCATG
CACGGCATTGTGTATTAGAGATGGTTGGGAGCTTTGTCGCCCTGAGAATGCAGCAGACTCCACTTTTCACTTTCTGTATGAGCAATGGAAAGAGCATTTC
CCTCCTGGTTATCTGCTCAAAGATGCGTTGGTAGATCAACCTTTCGAAGGAGATGCTTGCCACTGCCACTTTGTTTTTGTTCAAGACAACAATCTAAGTG
CCGCTCCACAGTTAGTCTTAAAGTATCAATGGTTTGTGGGAGAAAGAGCACTTTCAAGCTTTGCTGCCATTCCTGATGCCACTGGAGAGGTATACTGGCC
AAAGCATGAAGACATCGGTAAATTTCTGAAAGTGGAATGTACTTCCGTAATGGGAGAAATAGAGTATCCTCCTATTTTTGCCCTATCATCATGTGTTTCA
CCCGGAAATGGAATTCCAAAGGTTGTGAACCTTGAAGTGCAAGGAGAGTTGGTGGAAGGAAATGTTATCAAAGGCTATGCTGGGATTGCATGGTGTGGTG
GAACTCCAGGGAAAGGTGTTGCTAGTTGGCTGAGGAGAAGATGGAATAGTAGCCCAGTGGTTATTGCAGGGGCAGAAGATGAGGAGTATTGTTTAACTTT
AGATGACATTGATTCCAGTTTGGTTTTTATGTACACACCAGTCACAGAAGAAGGCGCCAAAGGGGAACCTCAATATAAATATACTGATTTTGTTAAAGCA
GCACCTCCATCGGTCAGCAATGTTCGAATTATTGGAGATATTGTAGAAGGAAATATCATTAAAGGAGTTGGGGATTACTTTGGGGGGAAAGAGGGCCCCA
GCAAGTTTGAATGGTTACGGGAGAATAAGAATACTGGAGATTTTGTATCCATATCAACTGGTACATCTGAGTATGCCTTGACCAATGAAGATGTTGGACG
ATGCTTAGCTTTTGTATATTCACCCATTAATTTTGAAGGACAGGAAGGGAAATCTGTTTCAATATTTTCTCACCCAGTGAAGCAAGCACCTCCTAAAGTA
AAGAATATAAAGATAATTGGTCATTTACGGGAAAACAGTAAGGTCACTGTAACTGCTACCGTTACTGGAGGAACTGGAGGAACTGAAGGTTCTAGCAGAG
TTCAGTGGTTTAAAACAAGTTCTTCAACATTGGATGGTGAGAATAGTCTTGATGCATTGATCACAGCAAAGATTGCAAAGGCATTACGAATACCTTTAGG
AGCAGTTGGTTATTACATTGTTGCGAAATATACTCCTATGACTCCAGATGGTGAATCTGGTGAACCAGCATATGCTATATCTGAAAAAGCAGTTGAGACT
CTTCCCCCAAGCCTAAATTTCTTATCAATTTCGGGCGACTACACTGAAGGTGGAATATTGACAGCATCATACGGTTATGTAGGTGGTCATGAAGGGAAAA
GTGAATATAATTGGTTTCTTCATGAGTTTGAAAGAGACAATGGAACCTTGATACTTGAGGGTTCTGGAGTTCTTCGGTATTGTGTTACTAGAGATGCAAT
TGGTAAATTCATATCTTTTCAGTGCATCCCAGTCCGTGATGATGGAATAGCAGGTGAGCCAAGAACTTGCATGGGGGTGGAACGCATTCGACCAGGAAGT
CCAAGATTGCTTTCTCTGCAAATAGTTGGAAATGCTATTGAAGGAACTTCACTAAGCGTTGACAAAAAATACTGGGGAGGTGAAGAGGGGAATTCAGTTT
TCTGCTGGTTTCGGAGTAGTTCAGATGGTGCTCAAATTGAAATTCAAGGAGCCAATACTTCGTCATACATGCTTTCAGTTGATGACATTGGCTCCTTTGT
CTCGGTTTCATGTGAGCCTGTGAGAAGTGATTGGGCTTGTGGTCCTACCATTTTTTCTGAACAGATTGGACCTATTATACCAGGTCCTCCAACTTGTCAG
TCATTGGAGTTTCTGGGATCGATGATGGAAGGACAACGTTTGAGCTTTGTTGCATCATATAGTGGAGGAGAGAGAGGGAATTGCTTCCATGAATGGTTTA
GAGTGAAAAGTGGTGGTATCAGGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.011G090200.3 pacid=42782104 polypeptide=Potri.011G090200.3.p locus=Potri.011G090200 ID=Potri.011G090200.3.v4.1 annot-version=v4.1
MVCCLAVVVVESGYLSLKMEEPLSDEAAKKSPTSDKQRQQSSVSSMQIPMKVSKTTKPTISANSHLLTPIGSIRKRTEPKNSSDSSSNVTAKNASSCNTK
SVPIARRNSTGGVPEKQPVSSTKRQNTSGKTNAVSDPVRRSLPELRRSSLPPTKPMVRTGSVSETRNSVPMDKCLRASTGSGVSRLEKPSVKPALPASSS
SSSSRRVISTSVDSTASSMSRKKLSSPSATSPSISSGLRAGSLSTSRDRSFNLTGRRRAGAPESHDSHFIALPLVETKAGDDVRLDLRGHKVRSLNASGL
NLAQNLEFVYLRDNLLSTLEGIEILKRVKVLDLSFNEFKGPGFEPLENCQALQQLYLAGNQITSLVNLPQLPNLEFLSVAQNKLKSLSMAGQPRLQVLAA
SKNKITTLKGFPHLPSLEHLRVEENPILKMPHLEAASILLVGLTLKKFNDRDLSREEVAIAKRYPACTALCIRDGWELCRPENAADSTFHFLYEQWKEHF
PPGYLLKDALVDQPFEGDACHCHFVFVQDNNLSAAPQLVLKYQWFVGERALSSFAAIPDATGEVYWPKHEDIGKFLKVECTSVMGEIEYPPIFALSSCVS
PGNGIPKVVNLEVQGELVEGNVIKGYAGIAWCGGTPGKGVASWLRRRWNSSPVVIAGAEDEEYCLTLDDIDSSLVFMYTPVTEEGAKGEPQYKYTDFVKA
APPSVSNVRIIGDIVEGNIIKGVGDYFGGKEGPSKFEWLRENKNTGDFVSISTGTSEYALTNEDVGRCLAFVYSPINFEGQEGKSVSIFSHPVKQAPPKV
KNIKIIGHLRENSKVTVTATVTGGTGGTEGSSRVQWFKTSSSTLDGENSLDALITAKIAKALRIPLGAVGYYIVAKYTPMTPDGESGEPAYAISEKAVET
LPPSLNFLSISGDYTEGGILTASYGYVGGHEGKSEYNWFLHEFERDNGTLILEGSGVLRYCVTRDAIGKFISFQCIPVRDDGIAGEPRTCMGVERIRPGS
PRLLSLQIVGNAIEGTSLSVDKKYWGGEEGNSVFCWFRSSSDGAQIEIQGANTSSYMLSVDDIGSFVSVSCEPVRSDWACGPTIFSEQIGPIIPGPPTCQ
SLEFLGSMMEGQRLSFVASYSGGERGNCFHEWFRVKSGGIR
|
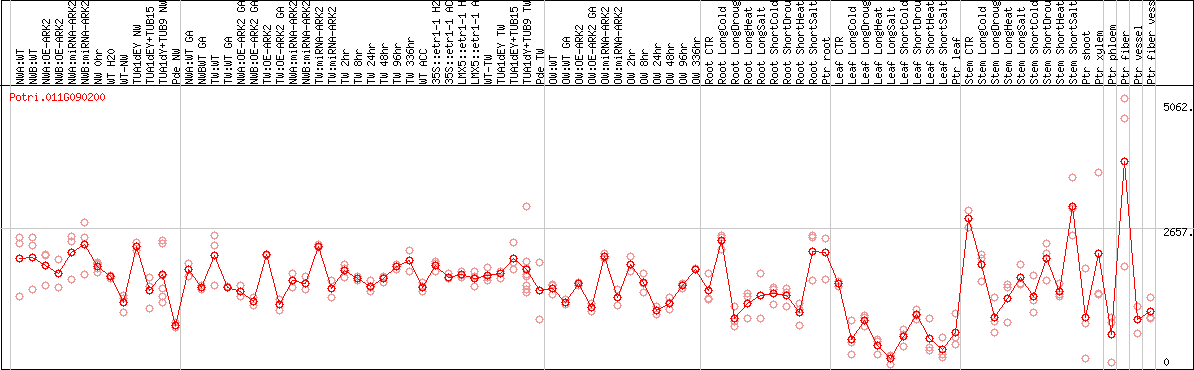
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G090200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
