Pt-TOP1.2 (Potri.011G092700) [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pt-TOP1.2 | ||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.011G092700.7 pacid=42781767 polypeptide=Potri.011G092700.7.p locus=Potri.011G092700 ID=Potri.011G092700.7.v4.1 annot-version=v4.1
ATGCATGCCAGCACAGACCCTAGTTGGACTCTAATGGTTGTTCAGGCACCTCCAAAGCCAATTCTCAATAATTTCAATGACGACGACGACGACGACGGGC
CCATAGTTTTCAAGAGAGGTGGTAACTCAACCTCTAAGCAAAATCAACCAAACCTTGAAGCAAAGAAACCATCTTCATCTTCTCAGAATTCCAATGGTCA
AAGCTTGAATGCCCACAAAGGCAAATCACCCATTTCATCTTCCAATGCATCCCCTGTGAAATCTCCTATAGGGAGCCCTAAAGCATCAGCTTCATCTGCC
AAGGCATCCCCAATGAAGCCACCAGTGACATATTCTAGAGTGCCAAGTTCTTCAATTGCCGTTAAGGAGGAGACCAAACCGATTGTGGATGATGATGATT
CTGAGGATGATAAGCCCTTGAGTTCTAGATTTAAGGGGAGCACTAACAATGCCAACAAAGGGGTTACTGTACCTGCTCGTGTCAAGGATGAACATTCAGA
TGATGATGAAGTCCCTTTATCCTCGAGGTTTGCAGCAAAGTCAAATGCAGGGTCATCAAGCTGTAAGACTATCAGTTCTAGTGAGAAGATATTGTTGGTT
TCAAAAATTCAGCTAAATGGTTCTACTGCAAAGGACAAGCAACAAAAATCTTCTGCTGTGCCAACTAAGAGACCTATGGATAATAAGGTTCCTTCAAACC
AGTCTTTTGCTAAAAAACCAAAACTTTCAGATGCGTCTACAACGAAGCAAGTTACTGTCAAGGCTGAGCTGAAGGCAGATGATGATGACCACCTTACAAT
TTCCCAGAGAATGAAGAAAGCAAGCTCCTCATCTAACAAATCATTGTCTGTGAAGCAAAATGTAACAAAAGTTGTTTCTTTTTCATCCAAGAAGACGACC
AAGAATAACAAGAAACAAATGCAAAATTCAAAATATTCAAAGTCAACAAAAGTACAACCTGGCTCCAGTGATGGGCAGAAAAAATGGAAAACCTTGGTTC
ACAACGGTGTCATCTTCCCACCTCCATACAAGCCTCATGGGGTTAAGATGCTTTACAAGGGGAAACCAGTTGATTTGACTCCTGAACAGGAGGAGGTTGC
AACCATGTTTGCGGTGATGAAAGATACAGAATATGTTCAAAAACCACAATTCCTTCAAAACTTTTGGAATGATTGGCGTACATTACTTGGGAAGAATCAT
GTAATACAGAAGCTAGAGGCCTGTGATTTCAACCCAATATATGAATGGCATCAGAAAGAAAAGGAGAAGAAGAAACAAATGAGTACTGAAGAGAAGAAGG
CTGTGAGAGAAGAGAAATTGAAGCAAGAGGAGATGTATATGTGGGCGATTGTTAATGGGGTTAAAGAGAAGGTTGGGAATTTCAGAGTTGAACCACCTGG
TCTATTTCGAGGACGGGGAGAGCATCCAAAGATGGGAAAGCTGAAAAGCCGTATTCGTCCCAGTGACATTACTATAAATATTGGAAAGGATGCCCCAATC
CCAAAATGTCCTATTCCTGGTGAAAGGTGGAAAGAAGTGAAGCATGATAACACTGTTACATGGTTGGCATTCTGGAATGATCCAATCAATCCAAAGGAAT
TCAAATATGTATTCTTGGCTGCCAGCAGTTCCTTGAAGGGGCAAAGTGACAAGGAGAAATATGAAAAAGCAAGGAAGTTGAAGGACTATATACACAACAT
CAGAACAGCTTACACAAAGGATTTTGCAAGTAAAGATATTACAAAGCGGCAAATAGCAGTGGCAACCTATCTTATTGATAAACTAGCTCTTAGGGCGGGT
AATGAGAAGGATGATGATGAAGCTGATACTGTTGGTTGCTGCACATTAAAAGTAGCAAATGTTGAGTGCATCCCTCCGAATAAGTTGAAGTTTGACTTCC
TTGGTAAAGATTCAATTCGATATGAAAACACAGTCGAGGTTGAGCTTCCTGTGTACAAGGCAATTAGACAGTTTCAAGTTGCAAAAAAGCCAACTGATGA
TCTTTTTGACTCTCTGGATACAAGTAAACTAAATGCTCACCTAAAAGAACTCATGCCTGGCCTTACAGCAAAAGTTTTTCGTACATACAATGCATCTATC
ACATTGGATGAAAAGTTATATGAGGAGACAGAAGATGGTGACGTTGCTGAAAAATTTGTCATTTATAACCAAGCAAACAAACAGGTTGCAATCATTTGTA
ATCATCAACGTACTATTTCAAAATCTCATGATGCACAAATGTCAAGATTGACCGAGAAACTTGAGGAACTTAAGGGCACACTGAAGGAGTTCAAAACTGA
TCTGGACAGGGCCAAAAAGGGGAAGCCCCCATTGAAGGATGCAGATGGAAAGCAAAAGAGGAACTTGACTCCTGAAGCCATAGAGAAGAAGATAGCAACA
ACCAATCAAAAGATTGAGAAAATGGAACTGTCCATGAAGACCAAAGAGGATCTGAAAACGGTAGCATTGGGCACGTCCAAAATTAATTACCTTGATCCTA
GGATTACAGTTGCATGGTGCAAGCGGCATGAAGTTCCTATTGAGAAGATATTCAACAAGTCTCTCCTAGCGAAGTTTACTTGGGCAATGGACGTTGACCC
CCAGTTCAGATTCTGA
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.011G092700.7 pacid=42781767 polypeptide=Potri.011G092700.7.p locus=Potri.011G092700 ID=Potri.011G092700.7.v4.1 annot-version=v4.1
MHASTDPSWTLMVVQAPPKPILNNFNDDDDDDGPIVFKRGGNSTSKQNQPNLEAKKPSSSSQNSNGQSLNAHKGKSPISSSNASPVKSPIGSPKASASSA
KASPMKPPVTYSRVPSSSIAVKEETKPIVDDDDSEDDKPLSSRFKGSTNNANKGVTVPARVKDEHSDDDEVPLSSRFAAKSNAGSSSCKTISSSEKILLV
SKIQLNGSTAKDKQQKSSAVPTKRPMDNKVPSNQSFAKKPKLSDASTTKQVTVKAELKADDDDHLTISQRMKKASSSSNKSLSVKQNVTKVVSFSSKKTT
KNNKKQMQNSKYSKSTKVQPGSSDGQKKWKTLVHNGVIFPPPYKPHGVKMLYKGKPVDLTPEQEEVATMFAVMKDTEYVQKPQFLQNFWNDWRTLLGKNH
VIQKLEACDFNPIYEWHQKEKEKKKQMSTEEKKAVREEKLKQEEMYMWAIVNGVKEKVGNFRVEPPGLFRGRGEHPKMGKLKSRIRPSDITINIGKDAPI
PKCPIPGERWKEVKHDNTVTWLAFWNDPINPKEFKYVFLAASSSLKGQSDKEKYEKARKLKDYIHNIRTAYTKDFASKDITKRQIAVATYLIDKLALRAG
NEKDDDEADTVGCCTLKVANVECIPPNKLKFDFLGKDSIRYENTVEVELPVYKAIRQFQVAKKPTDDLFDSLDTSKLNAHLKELMPGLTAKVFRTYNASI
TLDEKLYEETEDGDVAEKFVIYNQANKQVAIICNHQRTISKSHDAQMSRLTEKLEELKGTLKEFKTDLDRAKKGKPPLKDADGKQKRNLTPEAIEKKIAT
TNQKIEKMELSMKTKEDLKTVALGTSKINYLDPRITVAWCKRHEVPIEKIFNKSLLAKFTWAMDVDPQFRF
|
DESeq2's median of ratios [POPLAR]
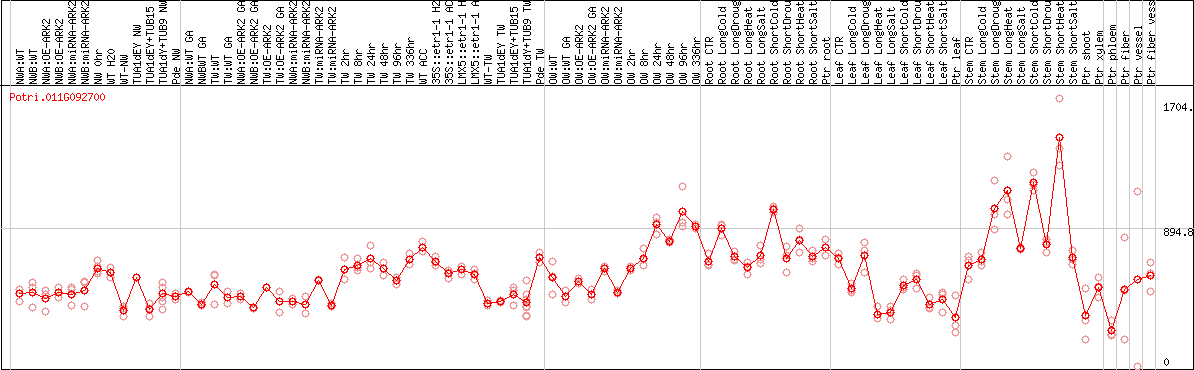
Coexpressed genes
Potri.011G092700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
