Potri.011G102700 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.011G102700.1 pacid=42781854 polypeptide=Potri.011G102700.1.p locus=Potri.011G102700 ID=Potri.011G102700.1.v4.1 annot-version=v4.1
ATGTTTTGTTCAGCCTTTTGCTTCCGATCCTTTGTTTTTCTTTTATCTTTAATCTCAGTCACCTGCTCCGACTACACTAATGAGACTGACTTATTAGCCT
TAATCCAATTCAAGAACAAAATAGTGGATGACCCTCTTGGGATCATGAGCTCGTGGAATAGTACTATCCATTTCTGCCAGTGGCATGGTGTCTCTTGTGG
TCGTAGGCACCAAAGGGTCAGAGTGTTAGCCCTACAGTCCCTCAAACTTTCAGGTACCATATCACCCCATATTGGCAATCTGAGCTTTTTAAGGGAACTA
CACCTTCAAAACAATAGTTTCTTTCATGAAATCCCTCCACAGGTTGGCCGTTTGCGTAGTTTGCAGATATTTTCTCTACACAATAATTCCATCTCTGGTC
AAATTCCTCCTAGCATATCTGATTGCTCTAACCTTATTTCAATTAAAATTGAATTCAACAATCTGACAGGAGAAATTCCTATGGAGCTAGGCTCTTTGTT
GAAGCTCAAAAACCTTACTCTGGAAGTCAATGGTCTTACAGGGACTATCCCTCCTTCTTTAGGGAATCTATCCTCCCTTGAGATCCTTCGGCTAGAAAAA
AATAAAATTCTGTTTGGGAATGTTCCTAGTACTCTAGGAAAATTGAAGAATCTTAGGATTTTAAATCTGATGGATAACAGGCTTTCAGGTGTCATTCCAC
CCTCAATCTTCAATCTTTCTTCATTGACAGCTTTGGATATAGGATTCAACCTTTTCCATGGCAATCTTCCATCTGATATAGGCATATCTCTCCCAAATCT
TGAATTCTTTTCAATCGCCTCAAACCAGTTCACTGGTTCCATTCCTGTTTCCATATCAAATGCTTCAAATATTGAATTGCTTCAAGTAAGTTTAAATAAT
CTAACAGGAGAAGTTCCTACATTAGAAAAGCTGCATAGGCTCAACTTTTTCACCCTTTTCTCCAACCATTTAGGAAGTGGACAAGCTAATGACTTAAGCT
TTCTCTCCTCATTGACCAATGCCACCACTTTAGAGTATTTGTCTATTAAACGAAATAACTTTGGCGGGGAGTTGCCGAAACAAATCAGCAATCTCTCAAC
CATGCTCGGGGTGATCAGTCTACCGGAAAATAACATACTTGGAAGCATCCCAGCTGGAATAGAGAAGCTGGTGAACTTGAAAGTTTTTGATGTTGGCAAC
AACAAAATCTCAGGTATTATTCCTTCTAGCATTGGAGAGCTTCAAAATCTTGAGGGATTGGTTCTTGACTACAACAATTTATCAGGGCGTATTCCATCCT
CTGTAGGGAATCTGACCAAATTAATGGCTCTCTATTTAGGGGATAACAGTCTTGAAGGTAGCATCCCCTCAAGTCTAGGGAACTGCAAGAAATTGTTAGT
GTTGACTCTGTGTGGAAACAATCTTAGTGGTGACATACCCCCAGGACTTTTTGGGATCTTCTCATTACTGTATATCTGTTTTTCTAAAAACCATTTTTCG
GGTTCCCTTCCCATTGAAATAGGAAAGTTAATAAATCTAGAATTCTTAGATGTATCTGGAAACATGTTATCAGGTGAGATTCCTAGCAGTCTTGGGGGTT
GTATAAGTCTTGAAGACTTGTATATGAATTCCAACTTCTTTCATGGGTCCATTCCTTCAGCATTGAGTTCACTGAGAGGAGTTCTACAGTTCAACTTTTC
TCATAACAACTTGTCAGGGAAAATTCCAGAGTTTTTTCAGGGCTTTAACTCATTAGAGATGTTGGATCTCTCTTATAACAACTTCGAGGGCATGATACCA
GATGAAGGAATTTTCAAGAACTCAACTGCAGTTTCAGTCATAGGAAATAGCCAGCTATGTGGTGGTAACACCGAGTTGGGGCTGCCTAGATGCAAGGTCC
ACCAACCAAAGAGGCTGAAACTCAAATTGAAGATAGCAATCTTTGCCATTACAGTGCTTCTAGCATTGGCCCTTGTTGTCACTTGTTTATTTCTTTGTTC
ATCAAGAAGGAAAAGAAGAGAAATTAAATTGAGCTCTATGCGGAATGAGCTATTGGAGGTGTCTTATCAAATTCTCTTAAAAGCTACCAATGGATTCTCT
TCAGCGAATTTGGTTGGTATAGGTAGCTTTGGGTCTGTGTATAAAGGAATGCTTGACCAAAATGGAATGGTCATTGCTGTCAAAGTGCTTAATCTCATGC
GTCAAGGAGCTTCTAGGAGTTTCATAGCTGAATGTGAAGCCCTGAGAAACATCAGACATCGGAATCTTGTCAAGGTATTAACTGCATGCTCAAGTATTGA
TTACCATGGTAATGATTTCAAGGCTATAGTTTATGAGTTCATGGCTAATGGGAGTCTAGAGGATTGGCTGCATCCAACTGGTACAGGAGGTGGCACAGCG
CTGACTCTAAATCTTCTCCAGAGACTTAACATTGCCATCGATGTTGCGTGTGCACTAGAGTACCTTCATCATCATTGTGAAATGCCAATAGCTCACTGTG
ATCTTAAACCAAGCAATGTTCTTCTCGATGATGAACTCACTGGACATGTAGGTGACTTTGGGCTAGCAAAATTTCTTTCTGGAGCAAGCCTTGACAATCC
AACAAATGAATCAACTTCAATTGGAGTAAGAGGAACTATAGGTTACGCTCCTCCAGAGTATGGCGTAGGAGGTGAAGTGTCTGCATATGGTGATACATAT
AGCTATGGCATACTCTTGTTGGAGATGTTCACAGGAAAGAGGCCCACTGATGAAATGTTTAGAGAAGGCTCAAATCTTCATAACTTTGTCAAGAGAGCTG
TGCCTGAACAAGTGAAACAGATAACAGATCCAACTCTTCTTCAGGAAGAACCAACTGGGGACGACGACAAACGTGAAATTAGCAGCATGAGAAACAGCAG
ACCTCTCGAGTGCTTGAACTCAATATTACGAATTGGAATCTCATGTTCTGTTGAATTTCCAAGAGAGCGCATGAAGATTAGCGACGCCGTTGCTCAACTC
CATTCTGTCCGAAATGAACTTCAATCAACTGGTGGTAAGCAAGGATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.011G102700.1 pacid=42781854 polypeptide=Potri.011G102700.1.p locus=Potri.011G102700 ID=Potri.011G102700.1.v4.1 annot-version=v4.1
MFCSAFCFRSFVFLLSLISVTCSDYTNETDLLALIQFKNKIVDDPLGIMSSWNSTIHFCQWHGVSCGRRHQRVRVLALQSLKLSGTISPHIGNLSFLREL
HLQNNSFFHEIPPQVGRLRSLQIFSLHNNSISGQIPPSISDCSNLISIKIEFNNLTGEIPMELGSLLKLKNLTLEVNGLTGTIPPSLGNLSSLEILRLEK
NKILFGNVPSTLGKLKNLRILNLMDNRLSGVIPPSIFNLSSLTALDIGFNLFHGNLPSDIGISLPNLEFFSIASNQFTGSIPVSISNASNIELLQVSLNN
LTGEVPTLEKLHRLNFFTLFSNHLGSGQANDLSFLSSLTNATTLEYLSIKRNNFGGELPKQISNLSTMLGVISLPENNILGSIPAGIEKLVNLKVFDVGN
NKISGIIPSSIGELQNLEGLVLDYNNLSGRIPSSVGNLTKLMALYLGDNSLEGSIPSSLGNCKKLLVLTLCGNNLSGDIPPGLFGIFSLLYICFSKNHFS
GSLPIEIGKLINLEFLDVSGNMLSGEIPSSLGGCISLEDLYMNSNFFHGSIPSALSSLRGVLQFNFSHNNLSGKIPEFFQGFNSLEMLDLSYNNFEGMIP
DEGIFKNSTAVSVIGNSQLCGGNTELGLPRCKVHQPKRLKLKLKIAIFAITVLLALALVVTCLFLCSSRRKRREIKLSSMRNELLEVSYQILLKATNGFS
SANLVGIGSFGSVYKGMLDQNGMVIAVKVLNLMRQGASRSFIAECEALRNIRHRNLVKVLTACSSIDYHGNDFKAIVYEFMANGSLEDWLHPTGTGGGTA
LTLNLLQRLNIAIDVACALEYLHHHCEMPIAHCDLKPSNVLLDDELTGHVGDFGLAKFLSGASLDNPTNESTSIGVRGTIGYAPPEYGVGGEVSAYGDTY
SYGILLLEMFTGKRPTDEMFREGSNLHNFVKRAVPEQVKQITDPTLLQEEPTGDDDKREISSMRNSRPLECLNSILRIGISCSVEFPRERMKISDAVAQL
HSVRNELQSTGGKQG
|
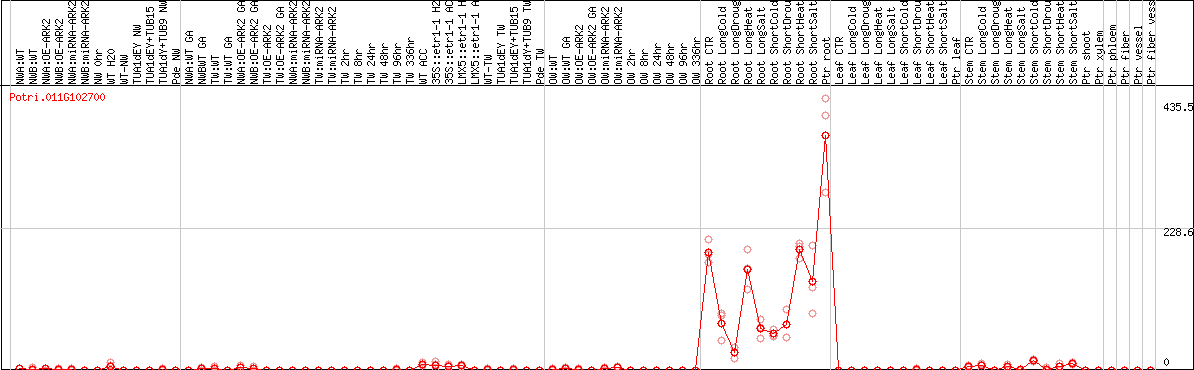
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G102700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
