Potri.011G123900 [POPLAR]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Flax homologues |
|
|||||||||||||||
| PFAM info |
| |||||||||||||||
|
Representative CDS sequence |
>Potri.011G123900.1 pacid=42780334 polypeptide=Potri.011G123900.1.p locus=Potri.011G123900 ID=Potri.011G123900.1.v4.1 annot-version=v4.1
ATGGCAAACTACCTCGCTCAATTTCAAACTATCAAAAATAGCTGCGATCACATCGTCATTGCAGTTGAAGATGTGAGTGATTTATGGCCGAACATAAAAT
CAGGATTCGATGAGAGAGTTCCAATTAAAAGAGCTAGTTTAAATAACAAGACTCGAAATCCAGTTCTCGTAGAGAATTTTCCTTGCGAATTTATATTAAC
TACTGATTCAAGACTCCGCAGCCGGTTTCCTCAAGAACAGTCTTTGTTTTGGTTTCGCGAACCGTACGCTACTATTGTTCTAGTCACTTGTGAGGATCTT
GATGAGTTTAAAACGATACTTAAGCCGCGGTTGAAGTTAATTGTGCAAAATGACGAGAAGGAATGGTTTATTGTGTTTGTGTCGAGAGCGCATCCGAGTA
ATGATAATGCTGTTAAAATGGCGAAAAAGGTTTATGCTAAGCTTGAAGTTGATTTCAGCTCGAAAAAAAGGGAGAGGTGCTGCAAATATGATATTCATGG
CCCTGAAGCAATTTTTTGGGATGATTTAGAATCGAAAATTATGGAGTGTGTCAGAAACACACTGGATAGGCGTGTACAGTTTTATGAGGATGAGATACGT
AAGCTCACTGAACAGCGGTTCATGCCAGTCTGGAATTTCTGTAATTTCTTTATATTGAAGGAAAGCTTGGCTTTTATGTTTGAGATGGCTCATCTTTATG
AAGATGCTCTACGTGAATATGATGAATTGGAACTCTGTTATTTGGAAACAGTTAATATGCCTGGAAAACAGAGAGAATTTGGAGGAGTAGACCATGGTGA
TGATTGGGCAGCACTGCTTAATCCTGAAAATAAGCCATTGACGCAGATTGTTCAAGATGATTCATTTAGAGAATTTGAATTTAGGCAGTATCTCTTTGCT
TACCAATCAAAGCTTTTATTCAAGCTGAATCGTCCCTTTGAGGTTGCATCAAGGGGTCATTCATTTATAATTGGCTTCTCAAAGGCCCTCACATTGCATG
AGAATATGCTACCTTTTTGTATGCGAGAAGTTTGGGTAATAACTGCTTGCTTGGCTATAATCAATGCTACTGCTCCTAATTACGATGGACTTGTGGCACC
TGACATAGAAAAGGAGTTCTACCGCCTCAAGGGTGATCTTTATTCTTTATGTCGAGTTAAGTTCATGAGGCTGGCCTATTTGATCGGATATGGAGCAGAT
ATAGAAAGAAGTCCTGTCAACAGTGCCTTACTCAGCATGCTACCTTGGCCCAAGCCTCTGGTTTGGCCATCGGTTCCACCTGATGCTTCCCCTGAGGTTC
TTGAGAAGGAAAAGGTGATCCTTCAAGCTACTCCTAAAATCAAACACTTTGGCATCCAGAGGAAGCCACTGCCTCTTGAACCTTCTGTACTTCTGCGAGA
GGCTAATCGGCGAAGGGCTTCCCTTTCTGCTGGGAATGTGTTTGAGATGTTTGATGGCCGCCCAACCTTGATTGATGGATCAGCTTCAGATGCTTCATCA
AGGACCCCCCTATTAAAGAAAATGAATGCGATTTCTATGTCGCGGACTAATTCTTCACCCGGAACTTTTGATGGTTCAGTAGATCGACCTATGAGACTTG
CAGAGATTTATGTTGCTGCCGAACATGCTCTGAAGCATACAATTTCTGATGCAGATCTTTGGAAAGCTTTGTCATCTGTAGAGGAGTTTGAGCAAAAGTA
TCTCGAGCTAACCAAAGGTGCTGCTGACAATTACCATCATTCCTGGTGGAAAAGGCATGGAGTTGTTCTCGATGGTGAAATAGCAGCTGTTTGCTTTGGG
CATGGAAATTTTGATCTAGCTGCAAAGTCTTATGAGAAAGTTTGTGCCCTTTATGCTGGCGAAGGATGGCAGGAATTATTGGCTGATGTCCTTCCCAATT
TAGCTGAGTGCCAGAAGATGCTAAATGATCAAGCTGGCTATTTAGCATCTTGCGTGAGATTGCTGTCTTTAGATAAAGGTTTATTCTCAACAAAGGAGCG
CCAAGCATTTCAGGCCGAGGTTCTTCGTCTTGCACACAGTGAGATGAAGGACCCTGTGCCCTTGGATGTTTCATCACTGATTACGTTTTCTGGAAATCCT
GGACCTCCCTTGGAGTTATGTGATGGGGATCCTGGTATTCTCTCTGTAACTGTTTGGAGTGGCTTTCCTGATGATATAACTCTTGATTCACTTAACCTCA
CTTTGACTGCTACATTTAATGCTGACGAGGGTGCTAAGGCATTAAGGAGCTCTACTGCCACCATACTGAAGCCTGGTAGGAATACCATTACCCTTGCTTT
ACCTCCTCAAAAACCAGGTTCCTATGTCTTGGGGGTTCTTACTGGGCAGATTGGTCAGTTGAGATTTAGATCTCATAGTTTTTCAAAGGTTGGCCCAGCG
GACAGTGATGATTTCATGAGTTATGAAAAACCCACTAGACCTATACTCAAGGTTTTCAAACCAAGACCACTAGTTGATCTTGCTGCAGCTATTTCATCTG
CTTTACTGATAAATGAAACTCAGTGGGTTGGGGTTATAGTACGGCCTATTGACTACTCTCTCAAAGGTGCTGTCTTGTATATTGATACTGGTCCAGGTCT
GAATATTGAAGAGTCGCATGTCATTGAGATGGAGACCCGTGTCAATATATCACAGAGCAGTGCTGAAATGACAAACTCTAATGGCACACAAAAGGATTGT
TCTTCAGCTTCTAAAAAAGAATTTCAGCAGTTAAAACTTCAAGATGGTAGAATAGAGTTTCCTGCTTGGGCAAGTGATGTAAATTCTGTTCTCTGGATTC
CTGTTCGTGCAATCAGTGACAGGCTTCCTAGAGGGTCATCCTCAGTCACCCCTCAGAAACAAAGCAATCTAGATGGCATGAGGACAATTGCTTTGAAACT
TGAATTTGGAGTATCCCACAACCAGATATTTGAAAGAACTGTAGCAGTACATTTCACTGATCCTTTCCATGTGAGCACACGTGTTGCAGACAAGTGTAAT
GATGGTACTCTGCTGTTGCAGGTGATACTGCATTCCCAAGTGAAGGCTACGTTGACCATATATGATGCTTGGCTTGAGCTTCAAGATGGATTTATTCATA
CTGGACAAGGAACTGGAAGACCAACTTCAAGCTTCTTCCCGCTCATGATTTCTCCAACCTCTAGAGCAGGAATCATGTTCAGTATTCGCTTGGGAAAAGT
AATTGACAAAGATGAAGTGGAAGCACTGCAGACAGAGAGCATACTAAATATCCGATATGGAATTTATGGTGAGCGAACAAACGGAGCACATCCTCCTGTT
TCCGTGGATGGCATTGAACCTGAGGATGCCAGGCAGGACTTGCTATTCAAGAGTGCTATTGTACTGCAGAGACCAGTGCTTGACCCATGCCTGGCTGTTG
GTTTTCTTCCTCTTCCTTCAACTGGCCTCAGAGTTGGACAACTCATCACCATGCAGTGGAGAGTTGAAAGATTGAAGGGCCTGGAGGACAATGGAATTTC
TGAACACAATGGTGAGGTGCTGTATGAAGTCAGTGCAAATTCTGAAAATTGGATGCTTGCTGGGAGAAAGCGCGGACATGTTACTCTCTCAACAATCCAA
GGTTCAAGAATAGTCATCTCAGTATTATGTGTGCCGTTGGTAGCTGGTTATGTCCGCCCTCCTCAACTTGGGCTACCAGATGTTGATGAGTCAAATATAA
GCTGCAATCCTCCTGGGCCCCATCTGGTTTGTGTCATGCCTCCAGCTCTCAGCTCCTCCTTCTGCATTCCGCCTTGA
|
|||||||||||||||
|
AA sequence
|
>Potri.011G123900.1 pacid=42780334 polypeptide=Potri.011G123900.1.p locus=Potri.011G123900 ID=Potri.011G123900.1.v4.1 annot-version=v4.1
MANYLAQFQTIKNSCDHIVIAVEDVSDLWPNIKSGFDERVPIKRASLNNKTRNPVLVENFPCEFILTTDSRLRSRFPQEQSLFWFREPYATIVLVTCEDL
DEFKTILKPRLKLIVQNDEKEWFIVFVSRAHPSNDNAVKMAKKVYAKLEVDFSSKKRERCCKYDIHGPEAIFWDDLESKIMECVRNTLDRRVQFYEDEIR
KLTEQRFMPVWNFCNFFILKESLAFMFEMAHLYEDALREYDELELCYLETVNMPGKQREFGGVDHGDDWAALLNPENKPLTQIVQDDSFREFEFRQYLFA
YQSKLLFKLNRPFEVASRGHSFIIGFSKALTLHENMLPFCMREVWVITACLAIINATAPNYDGLVAPDIEKEFYRLKGDLYSLCRVKFMRLAYLIGYGAD
IERSPVNSALLSMLPWPKPLVWPSVPPDASPEVLEKEKVILQATPKIKHFGIQRKPLPLEPSVLLREANRRRASLSAGNVFEMFDGRPTLIDGSASDASS
RTPLLKKMNAISMSRTNSSPGTFDGSVDRPMRLAEIYVAAEHALKHTISDADLWKALSSVEEFEQKYLELTKGAADNYHHSWWKRHGVVLDGEIAAVCFG
HGNFDLAAKSYEKVCALYAGEGWQELLADVLPNLAECQKMLNDQAGYLASCVRLLSLDKGLFSTKERQAFQAEVLRLAHSEMKDPVPLDVSSLITFSGNP
GPPLELCDGDPGILSVTVWSGFPDDITLDSLNLTLTATFNADEGAKALRSSTATILKPGRNTITLALPPQKPGSYVLGVLTGQIGQLRFRSHSFSKVGPA
DSDDFMSYEKPTRPILKVFKPRPLVDLAAAISSALLINETQWVGVIVRPIDYSLKGAVLYIDTGPGLNIEESHVIEMETRVNISQSSAEMTNSNGTQKDC
SSASKKEFQQLKLQDGRIEFPAWASDVNSVLWIPVRAISDRLPRGSSSVTPQKQSNLDGMRTIALKLEFGVSHNQIFERTVAVHFTDPFHVSTRVADKCN
DGTLLLQVILHSQVKATLTIYDAWLELQDGFIHTGQGTGRPTSSFFPLMISPTSRAGIMFSIRLGKVIDKDEVEALQTESILNIRYGIYGERTNGAHPPV
SVDGIEPEDARQDLLFKSAIVLQRPVLDPCLAVGFLPLPSTGLRVGQLITMQWRVERLKGLEDNGISEHNGEVLYEVSANSENWMLAGRKRGHVTLSTIQ
GSRIVISVLCVPLVAGYVRPPQLGLPDVDESNISCNPPGPHLVCVMPPALSSSFCIPP
|
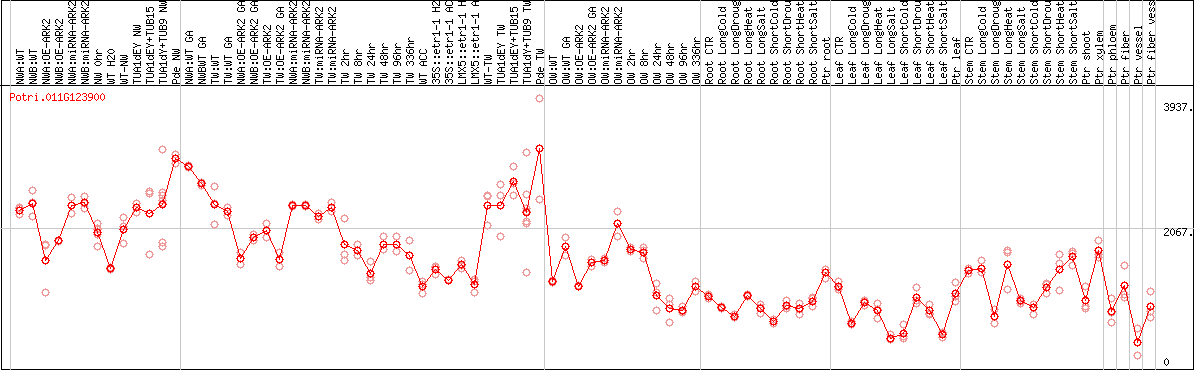
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G123900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
