External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G54270 446 / 3e-161
LHCB3*1, LHCB3*1, LHCB3*1, LHCB3*1, LHCB3*1, LHCB3*1, LHCB3*1, LHCB3*1, LHCB3*1, LHCB3*1, LHCB3*1,
light-harvesting chlorophyll B-binding protein 3 (.1)
AT2G05070 343 / 1e-120
LHCB2.2
LIGHT-HARVESTING CHLOROPHYLL B-BINDING 2, photosystem II light harvesting complex gene 2.2 (.1)
AT2G05100 343 / 1e-120
LHCB2.3, LHCB2.1
LIGHT-HARVESTING CHLOROPHYLL B-BINDING 2, photosystem II light harvesting complex gene 2.1 (.1)
AT3G27690 343 / 2e-120
LHCB2.3, LHCB2:4
LIGHT-HARVESTING CHLOROPHYLL B-BINDING 2, photosystem II light harvesting complex gene 2.3 (.1)
AT1G29910 337 / 7e-118
AB180, LHCB1.2, CAB3
LIGHT HARVESTING CHLOROPHYLL A/B BINDING PROTEIN 1.2, chlorophyll A/B binding protein 3 (.1)
AT1G29920 337 / 7e-118
AB165, LHCB1.1, CAB2
LIGHT HARVESTING CHLOROPHYLL A/B-BINDING PROTEIN 1.1, chlorophyll A/B-binding protein 2 (.1)
AT1G29930 337 / 7e-118
LHCB1.3, CAB140, CAB1
LIGHT-HARVESTING CHLOROPHYLL A/B-PROTEIN 1.3, CHLOROPHYLL A/B PROTEIN 140, chlorophyll A/B binding protein 1 (.1)
AT2G34430 336 / 1e-117
LHCB1.4, LHB1B1
light-harvesting chlorophyll-protein complex II subunit B1 (.1)
AT2G34420 335 / 2e-117
LHCB1.5, LHB1B2
PHOTOSYSTEM II LIGHT HARVESTING COMPLEX GENE 1.5, photosystem II light harvesting complex gene B1B2 (.1)
AT1G76570 186 / 6e-58
Chlorophyll A-B binding family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G407100
468 / 9e-170
AT5G54270 478 / 2e-173
light-harvesting chlorophyll B-binding protein 3 (.1)
Potri.014G165100
353 / 1e-124
AT2G05100 507 / 0.0
LIGHT-HARVESTING CHLOROPHYLL B-BINDING 2, photosystem II light harvesting complex gene 2.1 (.1)
Potri.002G221400
352 / 6e-124
AT2G05070 499 / 0.0
LIGHT-HARVESTING CHLOROPHYLL B-BINDING 2, photosystem II light harvesting complex gene 2.2 (.1)
Potri.005G239300
347 / 6e-122
AT2G34430 485 / 4e-176
light-harvesting chlorophyll-protein complex II subunit B1 (.1)
Potri.005G239200
345 / 2e-121
AT2G34430 484 / 2e-175
light-harvesting chlorophyll-protein complex II subunit B1 (.1)
Potri.002G189300
345 / 4e-121
AT2G34430 448 / 2e-161
light-harvesting chlorophyll-protein complex II subunit B1 (.1)
Potri.011G079500
344 / 6e-121
AT2G34430 480 / 6e-174
light-harvesting chlorophyll-protein complex II subunit B1 (.1)
Potri.005G258600
203 / 2e-64
AT1G76570 491 / 6e-176
Chlorophyll A-B binding family protein (.1)
Potri.019G063101
189 / 1e-59
AT4G10340 409 / 2e-145
light harvesting complex of photosystem II 5 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037875
453 / 6e-164
AT5G54270 460 / 3e-166
light-harvesting chlorophyll B-binding protein 3 (.1)
Lus10038575
451 / 3e-163
AT5G54270 462 / 9e-167
light-harvesting chlorophyll B-binding protein 3 (.1)
Lus10001741
350 / 5e-123
AT3G27690 481 / 1e-174
LIGHT-HARVESTING CHLOROPHYLL B-BINDING 2, photosystem II light harvesting complex gene 2.3 (.1)
Lus10001169
350 / 5e-123
AT3G27690 484 / 1e-175
LIGHT-HARVESTING CHLOROPHYLL B-BINDING 2, photosystem II light harvesting complex gene 2.3 (.1)
Lus10008191
345 / 4e-121
AT1G29910 452 / 5e-163
LIGHT HARVESTING CHLOROPHYLL A/B BINDING PROTEIN 1.2, chlorophyll A/B binding protein 3 (.1)
Lus10027966
343 / 2e-120
AT1G29910 446 / 2e-160
LIGHT HARVESTING CHLOROPHYLL A/B BINDING PROTEIN 1.2, chlorophyll A/B binding protein 3 (.1)
Lus10042219
342 / 5e-120
AT1G29930 444 / 1e-159
LIGHT-HARVESTING CHLOROPHYLL A/B-PROTEIN 1.3, CHLOROPHYLL A/B PROTEIN 140, chlorophyll A/B binding protein 1 (.1)
Lus10008594
341 / 1e-119
AT1G29930 445 / 4e-160
LIGHT-HARVESTING CHLOROPHYLL A/B-PROTEIN 1.3, CHLOROPHYLL A/B PROTEIN 140, chlorophyll A/B binding protein 1 (.1)
Lus10007362
293 / 3e-101
AT1G29930 370 / 5e-131
LIGHT-HARVESTING CHLOROPHYLL A/B-PROTEIN 1.3, CHLOROPHYLL A/B PROTEIN 140, chlorophyll A/B binding protein 1 (.1)
Lus10038490
252 / 4e-85
AT1G29910 337 / 2e-118
LIGHT HARVESTING CHLOROPHYLL A/B BINDING PROTEIN 1.2, chlorophyll A/B binding protein 3 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00504
Chloroa_b-bind
Chlorophyll A-B binding protein
Representative CDS sequence
>Potri.011G126700.1 pacid=42782073 polypeptide=Potri.011G126700.1.p locus=Potri.011G126700 ID=Potri.011G126700.1.v4.1 annot-version=v4.1
ATGGGAAATGGCAAATACATCATGGGTAATGAGTTGTGGTATGGACCTGACAGAGTGAAGTACTTGGGACCTTTTTCAGCTCAGACTCCTTCATACCTGA
CTGGAGAATTCCCAGGTGACTATGGATGGGACACTGCTGGTTTATCAGCTGATCCTGAGGCCTTTGCTAAGAACAGAGCTCTTGAGGTCATCCATGGCAG
ATGGGCTATGCTAGGAGCACTAGGTTGCATCACCCCAGAAGTCCTTGAGAAATGGGTTAAGGTAGACTTCAAGGAACCAGTTTGGTTCAAAGCTGGAGCT
CAAATCTTCTCAGAAGGTGGCTTGGACTACTTGGGCAACCCCAACCTTGTCCATGCCCAAAGCATCCTTGCTGTCCTTGGATTCCAAGTTGTCCTCATGG
GCCTTGTTGAGGGATTCCGCATCAATGGTCTCCCTGGAGTAGGAGAGGGTAATGACCTGTACCCTGGTGGCCAATACTTCGACCCTCTTGGCCTGGCCGA
TGATCCAACTACATTTGCCGAGCTCAAGGTGAAGGAAATCAAGAATGGCCGGCTAGCTATGTTCTCCATGTTTGGATTCTTTGTTCAGGCTATTGTTACT
GGAAAAGGTCCCCTGGAGAACCTCTTGGACCACCTTGACAACCCAGTTGCCAACAACGCCTGGGTCTACGCCACCAAGTTTGTGCCTGGGTCATAA
AA sequence
>Potri.011G126700.1 pacid=42782073 polypeptide=Potri.011G126700.1.p locus=Potri.011G126700 ID=Potri.011G126700.1.v4.1 annot-version=v4.1
MGNGKYIMGNELWYGPDRVKYLGPFSAQTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCITPEVLEKWVKVDFKEPVWFKAGA
QIFSEGGLDYLGNPNLVHAQSILAVLGFQVVLMGLVEGFRINGLPGVGEGNDLYPGGQYFDPLGLADDPTTFAELKVKEIKNGRLAMFSMFGFFVQAIVT
GKGPLENLLDHLDNPVANNAWVYATKFVPGS
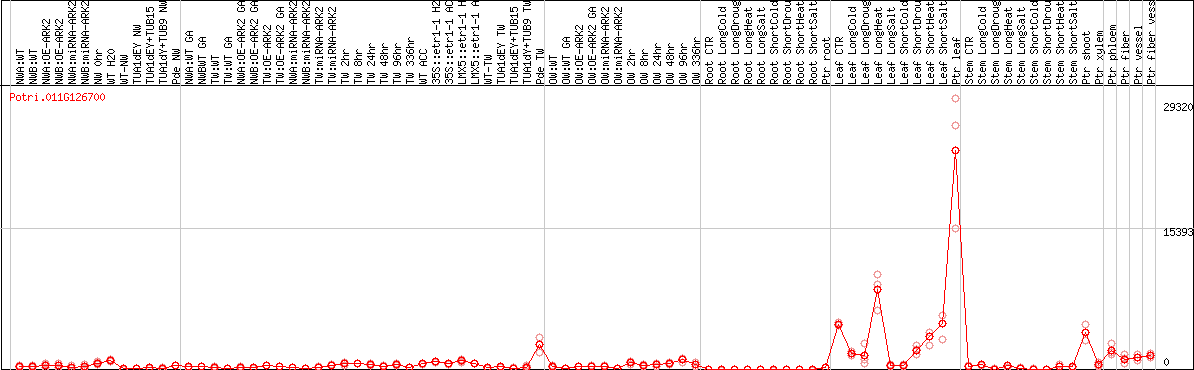
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G126700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.