External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G21200 325 / 2e-110
ATGA2OX8
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
AT1G50960 310 / 7e-105
ATGA2OX7
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 7, gibberellin 2-oxidase 7 (.1)
AT5G05600 171 / 3e-50
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G19010 157 / 4e-45
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.3)
AT3G11180 157 / 1e-44
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
AT1G62380 154 / 2e-44
ATACO2, ACO2
ACC oxidase 2 (.1)
AT5G58660 153 / 1e-43
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G10490 152 / 4e-43
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT2G38240 152 / 4e-43
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G05010 151 / 4e-43
ACO4, EAT1, EFE
ethylene forming enzyme, ethylene-forming enzyme (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G134000
572 / 0
AT4G21200 340 / 1e-116
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.001G418200
488 / 7e-175
AT4G21200 369 / 8e-128
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.004G022800
369 / 1e-127
AT4G21200 441 / 7e-156
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.011G026700
357 / 3e-123
AT4G21200 444 / 1e-157
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.010G096800
315 / 1e-106
AT4G21200 362 / 8e-125
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.008G145300
295 / 9e-99
AT4G21200 354 / 1e-121
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.001G278200
193 / 4e-59
AT5G58660 312 / 2e-105
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G451900
168 / 2e-49
AT4G10490 498 / 3e-178
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.016G117100
164 / 2e-47
AT5G05600 491 / 4e-175
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018372
329 / 4e-112
AT4G21200 419 / 2e-147
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10007640
325 / 1e-110
AT4G21200 429 / 1e-151
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10006759
312 / 2e-105
AT4G21200 419 / 2e-147
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10029081
293 / 7e-98
AT4G21200 365 / 3e-126
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10030567
292 / 2e-97
AT4G21200 370 / 2e-128
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10012912
291 / 2e-97
AT4G21200 368 / 2e-127
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10013069
291 / 2e-96
AT4G21200 366 / 5e-126
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10021002
166 / 2e-48
AT3G19000 389 / 3e-135
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10035334
156 / 8e-45
AT1G05010 495 / 6e-178
ethylene forming enzyme, ethylene-forming enzyme (.1)
Lus10029992
156 / 8e-45
AT1G05010 496 / 2e-178
ethylene forming enzyme, ethylene-forming enzyme (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF03171
2OG-FeII_Oxy
2OG-Fe(II) oxygenase superfamily
CL0029
Cupin
PF14226
DIOX_N
non-haem dioxygenase in morphine synthesis N-terminal
Representative CDS sequence
>Potri.011G134100.1 pacid=42780364 polypeptide=Potri.011G134100.1.p locus=Potri.011G134100 ID=Potri.011G134100.1.v4.1 annot-version=v4.1
ATGGATCCTCCATTTGAAGAGAAATACAAATCCCTCTTAGCCAATGCTACTCTATTATCAAAAGACAAGGATGATGTCATAATGAGTGATTACGAGGAGT
ATGAGCTGCCTATCATAGATCTTCACCGTTTGACTCTCTCCTTCTCGGAGAGAGAGCAATGTATAAAAGAGATAAGGCAGGCTGCAAGTGAATGGGGCTT
CTTTCAAGTTGTGAATCATGGGATTCCACAGGAGATTTTGGAGCGCATTCAACTGGAGCAAAGGAAGCTTTTCCATCATCCATTCAGCAAAAAGGCTGAA
GAAAACATTTTGAATTTGTCTGAAAATAATGGTTACAGATGGGGTAACCATACAGCCACTTGTTTGAGGCAGATATCATGGTCAGAAGCCTTCCACATAC
CTCTCACAGATATTTCAAAAATTGGTGGTGAATACAAGAGTCTCAGAGAGAGCATTGAAGCTTACGCAGCAAGTGCTGAGAAATTGGCTAAAGAGATGAC
AGAGATTCTAGCTAAGAATCTGGATATTTCCTCCACTTACTTTCAAGAGAATTTTCTGCCGGAAACTAATTATCTTCGAATGAACAGGTATCCTCCATGT
CCATTCTATTCTGAGGTGTTTGGCATACTGCCTCATACCGATAGCTGTTTCGTCAATGTATTAATTCAAGACCAGATTGGAGGATTGCAACTTCGGGTAA
ATGGAGAATGGATTAGCGTTAAACCTCATCCAGAAGCTCTACTAATTAATCTTGGTGACTTATTTCAGGCATTGAGCAATGATGTTTACAAAAGCATTAG
GCATCGGGTAGTACTTGCCTCCAAACAAGTTGAAAGGCTTTCTTTGGCATATTTATATTGCCCCAGAAATGATGCTGTCATACAGAGTGGAATGAAGCCA
TCAATTTATAGGAAGTTCACATTCGAAGAGTTGATGAAGCAAAATTCAAGGGATATTGAAGAAACTGGCCGCAAACTCGGGATCTCAAGGTTTCTCATGT
GA
AA sequence
>Potri.011G134100.1 pacid=42780364 polypeptide=Potri.011G134100.1.p locus=Potri.011G134100 ID=Potri.011G134100.1.v4.1 annot-version=v4.1
MDPPFEEKYKSLLANATLLSKDKDDVIMSDYEEYELPIIDLHRLTLSFSEREQCIKEIRQAASEWGFFQVVNHGIPQEILERIQLEQRKLFHHPFSKKAE
ENILNLSENNGYRWGNHTATCLRQISWSEAFHIPLTDISKIGGEYKSLRESIEAYAASAEKLAKEMTEILAKNLDISSTYFQENFLPETNYLRMNRYPPC
PFYSEVFGILPHTDSCFVNVLIQDQIGGLQLRVNGEWISVKPHPEALLINLGDLFQALSNDVYKSIRHRVVLASKQVERLSLAYLYCPRNDAVIQSGMKP
SIYRKFTFEELMKQNSRDIEETGRKLGISRFLM
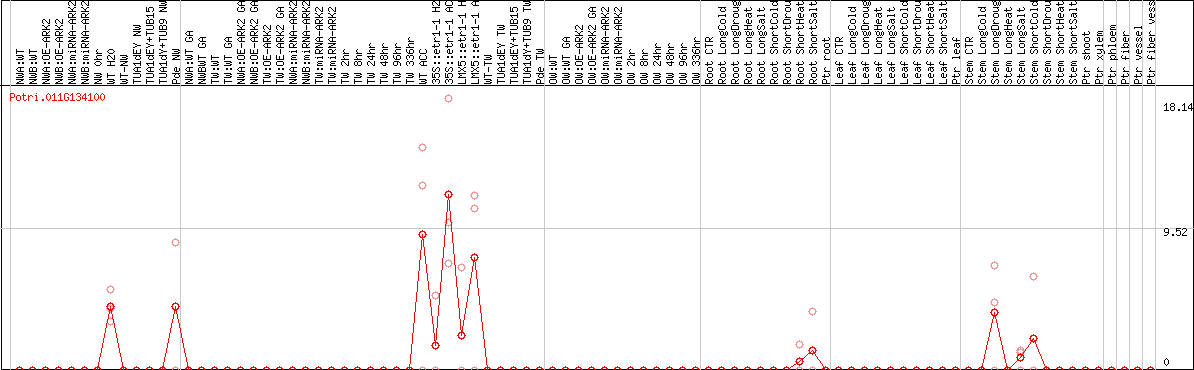
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G134100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.