External link
Symbol
ARRO.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G14130 367 / 5e-128
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G14120 319 / 4e-109
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT2G38240 120 / 2e-31
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G21200 115 / 9e-30
ATGA2OX8
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
AT1G50960 112 / 1e-28
ATGA2OX7
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 7, gibberellin 2-oxidase 7 (.1)
AT1G52800 111 / 2e-28
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G03070 111 / 2e-28
AOP1.1, AOP1
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G80330 111 / 3e-28
ATGA3OX4
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 4, gibberellin 3-oxidase 4 (.1)
AT1G52820 109 / 7e-28
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT5G24530 110 / 8e-28
DMR6
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G455400
548 / 0
AT1G14130 376 / 1e-131
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.008G145300
130 / 2e-35
AT4G21200 354 / 1e-121
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.009G025900
125 / 3e-33
AT4G25300 410 / 3e-143
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Potri.018G033400
122 / 2e-32
AT1G52820 269 / 4e-89
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G176200
122 / 2e-32
AT1G52820 496 / 8e-179
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.010G201000
122 / 6e-32
AT3G21420 290 / 5e-96
LATERAL BRANCHING OXIDOREDUCTASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.016G117100
122 / 6e-32
AT5G05600 491 / 4e-175
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.010G096800
121 / 6e-32
AT4G21200 362 / 8e-125
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.006G101100
118 / 1e-30
AT5G05600 462 / 1e-163
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031054
368 / 1e-128
AT1G14130 352 / 3e-122
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10016659
124 / 4e-33
AT1G52820 445 / 1e-158
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10005037
119 / 9e-31
AT3G11180 471 / 3e-166
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10030567
118 / 1e-30
AT4G21200 370 / 2e-128
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10012912
117 / 2e-30
AT4G21200 368 / 2e-127
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10008097
115 / 5e-30
AT1G52800 221 / 2e-70
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10029081
112 / 2e-28
AT4G21200 365 / 3e-126
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10011980
111 / 3e-28
AT1G17020 382 / 1e-132
senescence-related gene 1 (.1)
Lus10013069
112 / 4e-28
AT4G21200 366 / 5e-126
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10015573
110 / 5e-28
AT5G24530 510 / 0.0
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF03171
2OG-FeII_Oxy
2OG-Fe(II) oxygenase superfamily
CL0029
Cupin
PF14226
DIOX_N
non-haem dioxygenase in morphine synthesis N-terminal
Representative CDS sequence
>Potri.011G146600.1 pacid=42780342 polypeptide=Potri.011G146600.1.p locus=Potri.011G146600 ID=Potri.011G146600.1.v4.1 annot-version=v4.1
ATGGGTAAGAGTGGTGTCCCAATAATTGATCTGCATGAGTTTCCAGGACAGTATGAGAAACTGAGGAGAGCATGTGTGGAGTGGGGCTGTTTCAGGATTG
TGAATCACAATATCTCTTTGGCCCTCATGGCAGATATGAAGAGGGTTGTGAGGTCATTGCTAGACCTGCCCTTTGATGTCAAAAAGCGCAACATAGATGT
CATAGCTGGGAGTGGCTACATGGCACCAAGCAAAGTCAACCCTCTTTATGAGGCTTTAGGACTTCATGATATGAGGTCTTCTCAAGCCGTGGAAACTTTC
TGCTCCCAAATGGATGCCTCTCCTTACCAAAGAGAGGTTATCGAGATGTATGCTAAGGCAATTCATGGGGCAGCCATGGATATAGCACGAAAGTTGGCTG
AAAGTATGGGGTTGAATGGTGATTTGTTCGAGTCATGGATTTCCCAGTTCAGGATCAACAAGTACAGCTTCACCCCCGAAACCATAGGCTCGTCTGGAGT
CCAAATACATACAGATTCTGGGTTCCTGACTATCCTTCAAGATGATGAAAATGTTGGTGGACTTGAAGTGATGGACCCATCTGGTGTGTTTGCTGCAGTT
GATCCTTCACCAGGCACCCTCCTGGTCAATCTTGGAGACGTTGCTACGGCATGGAGCAATGGAGGGCTTCGTAACGTGCAGCATCGAGTGCAATGCAAGG
AAGCCACAATTCGGATCTCAATCGCTTCATTCCTGGTCGGTCCCAAGGACGAAGTGGAAGCTCCGCCGGAACTTGTGGACTCTGAACACCCACGTCTCTT
TGTCCCTTTCAAATATGAAGATTATAGAAAGCTCAGACTTACCACAAAATTGGAGGCTGGTGAAGCCCTTGAACTTAAACGCATCCTGTCCTAA
AA sequence
>Potri.011G146600.1 pacid=42780342 polypeptide=Potri.011G146600.1.p locus=Potri.011G146600 ID=Potri.011G146600.1.v4.1 annot-version=v4.1
MGKSGVPIIDLHEFPGQYEKLRRACVEWGCFRIVNHNISLALMADMKRVVRSLLDLPFDVKKRNIDVIAGSGYMAPSKVNPLYEALGLHDMRSSQAVETF
CSQMDASPYQREVIEMYAKAIHGAAMDIARKLAESMGLNGDLFESWISQFRINKYSFTPETIGSSGVQIHTDSGFLTILQDDENVGGLEVMDPSGVFAAV
DPSPGTLLVNLGDVATAWSNGGLRNVQHRVQCKEATIRISIASFLVGPKDEVEAPPELVDSEHPRLFVPFKYEDYRKLRLTTKLEAGEALELKRILS
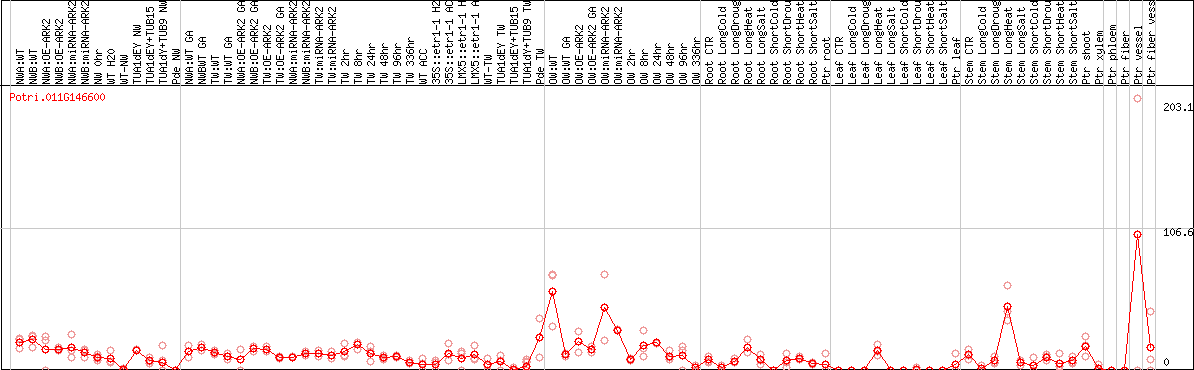
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G146600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.