External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G44680 96 / 3e-25
HDA09, HDA9
histone deacetylase 9 (.1)
AT4G38130 50 / 4e-09
ATHDA19, ATHD1, RPD3A, HDA19, HD1
ARABIDOPSIS HISTONE DEACETYLASE 19, ARABIDOPSIS HISTONE DEACETYLASE 1, histone deacetylase 1 (.1.2)
AT5G35600 50 / 6e-09
HDA7
histone deacetylase7 (.1)
AT5G63110 49 / 1e-08
RPD3B, CAT1, AXE1, ATHDA6, SIL1, RTS1, HDA6
RNA-MEDIATED TRANSCRIPTIONAL SILENCING 1, histone deacetylase 6 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G460000
117 / 2e-33
AT3G44680 739 / 0.0
histone deacetylase 9 (.1)
Potri.012G083800
58 / 1e-11
AT5G63110 716 / 0.0
RNA-MEDIATED TRANSCRIPTIONAL SILENCING 1, histone deacetylase 6 (.1)
Potri.015G082500
57 / 3e-11
AT5G63110 728 / 0.0
RNA-MEDIATED TRANSCRIPTIONAL SILENCING 1, histone deacetylase 6 (.1)
Potri.004G209800
50 / 8e-09
AT4G38130 815 / 0.0
ARABIDOPSIS HISTONE DEACETYLASE 19, ARABIDOPSIS HISTONE DEACETYLASE 1, histone deacetylase 1 (.1.2)
Potri.009G170700
50 / 9e-09
AT4G38130 811 / 0.0
ARABIDOPSIS HISTONE DEACETYLASE 19, ARABIDOPSIS HISTONE DEACETYLASE 1, histone deacetylase 1 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020570
106 / 3e-29
AT3G44680 806 / 0.0
histone deacetylase 9 (.1)
Lus10006274
106 / 5e-29
AT3G44680 756 / 0.0
histone deacetylase 9 (.1)
Lus10019486
55 / 1e-10
AT5G63110 727 / 0.0
RNA-MEDIATED TRANSCRIPTIONAL SILENCING 1, histone deacetylase 6 (.1)
Lus10001356
51 / 4e-09
AT4G38130 809 / 0.0
ARABIDOPSIS HISTONE DEACETYLASE 19, ARABIDOPSIS HISTONE DEACETYLASE 1, histone deacetylase 1 (.1.2)
Lus10001605
50 / 1e-08
AT4G38130 833 / 0.0
ARABIDOPSIS HISTONE DEACETYLASE 19, ARABIDOPSIS HISTONE DEACETYLASE 1, histone deacetylase 1 (.1.2)
Lus10043337
39 / 5e-05
AT5G63110 629 / 0.0
RNA-MEDIATED TRANSCRIPTIONAL SILENCING 1, histone deacetylase 6 (.1)
PFAM info
Representative CDS sequence
>Potri.011G156250.1 pacid=42781109 polypeptide=Potri.011G156250.1.p locus=Potri.011G156250 ID=Potri.011G156250.1.v4.1 annot-version=v4.1
ATGACACACCATCTTGTTCTCTCTTACGAGCTTCACAAGAAGATGGAAGTTTTTAGACCCCACAAGGCATATCCGGTTGAGCTTGCTCAGTTTCATTCAG
AGGATTATGTTGAATTTTTGCATCGGATTACGCCAGATACTCAGCACTTGTTTGCGGGGGAAATGGCTAGATGTGTGGATTTTTCGTTGTGA
AA sequence
>Potri.011G156250.1 pacid=42781109 polypeptide=Potri.011G156250.1.p locus=Potri.011G156250 ID=Potri.011G156250.1.v4.1 annot-version=v4.1
MTHHLVLSYELHKKMEVFRPHKAYPVELAQFHSEDYVEFLHRITPDTQHLFAGEMARCVDFSL
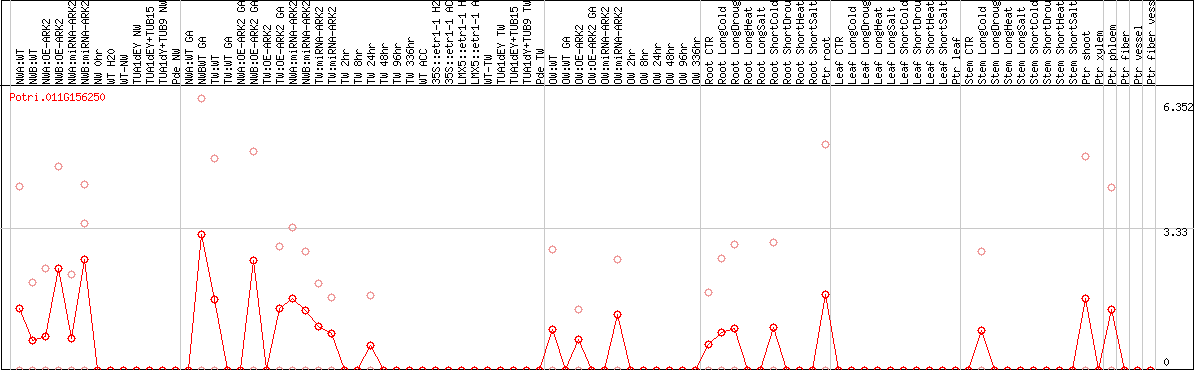
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G156250 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.