Potri.011G163600 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.011G163600.4 pacid=42781380 polypeptide=Potri.011G163600.4.p locus=Potri.011G163600 ID=Potri.011G163600.4.v4.1 annot-version=v4.1
ATGCGTGGTTGTGATTTTTATGAGAACTTTTGGAGGTTGCTTATGGAAATTGGAGTGAAGAAGACAATTTTTTCACCTAAGGCTATCATAAACCAGAAGT
TTGGTAGCAAGGCATGTTATAAGGTAGAGGAGGTACAGGAGGAGTCAACTCAGAATGGGTTCCCAGGATTGGCAATTCCTCAGAAAGCTCCTCTTTTTCG
GTGCCAGCTAGAGCTTCCAGAATTTACTGTTGTTTCAGATATATGTAGAAAGAAGAAGGATGCCGAGCAATCTGCAGCAGATTTGGCTTTAAAAAGGCTA
GGCCACCATTCCGTTGCTGAAAATCCATCTGATAAGGATCCTTGTGATGCTTTGATTGATCAAATCAAATATCTATTTTCAGACGAGTTCTCCCTACCTC
TTCATCCATTGAGGGGTCACCTTAGAGCAGCTTTGCTGAGAAGAGGTGATCTTTATGGTCTGGTACCTGCTTCTGTCATTACTACCTGTGATACAAAGAC
GAGTAATCTGTGTAAATTACTCAACCCTGAGGTGGAATTAAAACCTTTTTTGGCTTTGTCATTAATTATGAGGACTATACCAAGATTATCAGGCTGTGTT
ACATCCAAGGGGCAGCTTTCAATTCAGAAGCAAAATCCATATCCCACTGAGATCATAGAGTCATCAGATATCCAGCAATCTGATTCCCCAGAAAGCATAT
TGGTTAAAGCAATACAAATTCCAGCTTCACTGGATAAGACTGTTCAGCCAGTGACTCTTAATATTTCATCCGCCGGATATTACCTGGATGTTATTGCAGA
ACAACTTGGTGTAACAGATGCCAGCAAGGTTTTGCTCTCAAGGACTATTGGTAAAGCTTCCTCAGAGACCAGATTGTATTTTGCTGCTTCTGAGTCACTT
GTGATGGATCTGTTGTCGGACCTTGCAAATGTGAAAGATTTTCATGTGGAAGGACCTCCAAATGCAAGGGCAAGTTACTTCTGTGGTCAAGGAATATACG
GTGATGCAATCATGGCTTCCATTGGATACACATGGAGGTCTAAAGAACTTTTCCATGAACACGTGTCCCTCCAATCATATTACAGGATGCTTATAAGCAA
GATACCTAGTGGAAATTACAAGTTGTCTAGAGAAGCAATACTTGCAGCAGAGTTGCCCTCAGTATTCACTACTAAAGCAAATTGGAGAGGTTCCTTCCCA
AGGGAAATCCTATTCGCATTCTGCCATCAGCATAGGCTATCTGAGCCTATCTTCTCAACTACAAGTGTTCCTTTAAAGGCATCATGCGAGTTACTGAGAT
CACAGAAGAAGTTGAAGGTTACTGAGGTAGCTGGACTTGCGACTGAATATGCAAATGGAGGAGGTTTAAATGCGGGCGATGGAGAATCAGTAGGACTGGA
AAGCAACTTTAGATGTGAAGTAAAAGTATTTTCCAAGGGTCGGGATTTGATCATAGAGTGTTCCCCTAAGGAGATTTATAGGAAGCAGACTGATGCTACC
CACAGTGCTTCTCTGAAAGTTCTCTCATGGCTGAATGCATATTTTAAGGACCTTGGCATGCCTTTGGAGAAGCTAAATTGCTCTGCTGATGCCCTTGACA
TTAGTTTTTCTCTTGAAAACTTTCATAAAGAGTTTGCTTTATCCCAATCCATTCATAATGTTCAGCAAAGTGGGACTCAAGGATCCAAATTACCAGAATC
AAAAAGTACTGATATGCAATATACTTTGTCAGGACAAGATGTTTGCTTACCTAACATTGAAGGTTCAGATTCTGGTGTTTTCCCCTCTAATGGATCTTTG
TTGTGTATAAGTTACTCTGTATCCTTAGTGACAGAAGGTGGACATACAAAAGAGCTTATTGAAAGTAAAGATGAATTTGAGTTTGAGATGGGGGCTGGAG
CAGTGATTTCTGCTCTTGAAGCAGTTGTTACCCAAATGTCAGCTGGTCAATGTGCTCATTTCAACATGAATCTGCCCCCTCAAGAGTTCATTCTAGCTGC
AGTTGATGATCCTGGAAGGATTCACTCATTGTTATCCTCAGAAGCTTGCTGGTTGGAATACCATGTAACGTTGTTGCGTGTGACAAAACCACCAGAAGAA
AGAATGGAGCAGGCTCTTTTCAGCCCTCCGCTCTCAAAGCAACGGGTGGAATATGCTGTGCAGCATATCAAAAAATCTTGTGCTGCTACTTTGGTTGACT
TTGGATGCGGCTCTGGAAGTTTATTGGATTCTCTGTTAGATTACCCAACTTCACTAGAAAAGATTGTTGGTGTTGACATATCACAGAAGAGTCTTGGCCG
TGCTGCAAAGGTACTGCATGCAAAACTGAGCTCAAAGTCAGATGCAGGCATCAAATCTGCTATTCTTTATGATGGTTCCATCACAGAATTTGAACCTCAA
TTGTGTGGATTTGATATTGGCACTTGCTTAGAGGTAATAGAGCATATGGAGGAGGACCAGGCCTGTCGTTTTGGTGATATAGCATTGAGTTATTTTCGTC
CAAAGGTTCTTATAGTTTCCACCCCAAATTACGAATACAACGTGATTCTACAAAGATCTAGTCCTGTAACTCAAGAAGAATACCCAGACGAGAAAAGCCA
GTCAGAGTCATGTAAATTTCGCAACCATGACCACAAGTTTGAGTGGACAAGAGAGCAGTTCAACCACTGGGCATCTGAACTAGCGAAGAAGCACAATTAC
AGCGTCGAGTTCAGTGGGGTTGGTGGTTCTGGTGATGTGGAACCAGGTTTTGCTTCTCAGATAGCCGTCTTTAAACAGGAGAGCCTGCTTGATGAAGATG
ATCTTCTGACAAAACAAAATTCATCTCAGCATTGCAAAGTTGTATGGAATTGGAACTGCGCTGACAGATCGGTAGTCGGTACCCTCTCTAACAAGTAA
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.011G163600.4 pacid=42781380 polypeptide=Potri.011G163600.4.p locus=Potri.011G163600 ID=Potri.011G163600.4.v4.1 annot-version=v4.1
MRGCDFYENFWRLLMEIGVKKTIFSPKAIINQKFGSKACYKVEEVQEESTQNGFPGLAIPQKAPLFRCQLELPEFTVVSDICRKKKDAEQSAADLALKRL
GHHSVAENPSDKDPCDALIDQIKYLFSDEFSLPLHPLRGHLRAALLRRGDLYGLVPASVITTCDTKTSNLCKLLNPEVELKPFLALSLIMRTIPRLSGCV
TSKGQLSIQKQNPYPTEIIESSDIQQSDSPESILVKAIQIPASLDKTVQPVTLNISSAGYYLDVIAEQLGVTDASKVLLSRTIGKASSETRLYFAASESL
VMDLLSDLANVKDFHVEGPPNARASYFCGQGIYGDAIMASIGYTWRSKELFHEHVSLQSYYRMLISKIPSGNYKLSREAILAAELPSVFTTKANWRGSFP
REILFAFCHQHRLSEPIFSTTSVPLKASCELLRSQKKLKVTEVAGLATEYANGGGLNAGDGESVGLESNFRCEVKVFSKGRDLIIECSPKEIYRKQTDAT
HSASLKVLSWLNAYFKDLGMPLEKLNCSADALDISFSLENFHKEFALSQSIHNVQQSGTQGSKLPESKSTDMQYTLSGQDVCLPNIEGSDSGVFPSNGSL
LCISYSVSLVTEGGHTKELIESKDEFEFEMGAGAVISALEAVVTQMSAGQCAHFNMNLPPQEFILAAVDDPGRIHSLLSSEACWLEYHVTLLRVTKPPEE
RMEQALFSPPLSKQRVEYAVQHIKKSCAATLVDFGCGSGSLLDSLLDYPTSLEKIVGVDISQKSLGRAAKVLHAKLSSKSDAGIKSAILYDGSITEFEPQ
LCGFDIGTCLEVIEHMEEDQACRFGDIALSYFRPKVLIVSTPNYEYNVILQRSSPVTQEEYPDEKSQSESCKFRNHDHKFEWTREQFNHWASELAKKHNY
SVEFSGVGGSGDVEPGFASQIAVFKQESLLDEDDLLTKQNSSQHCKVVWNWNCADRSVVGTLSNK
|
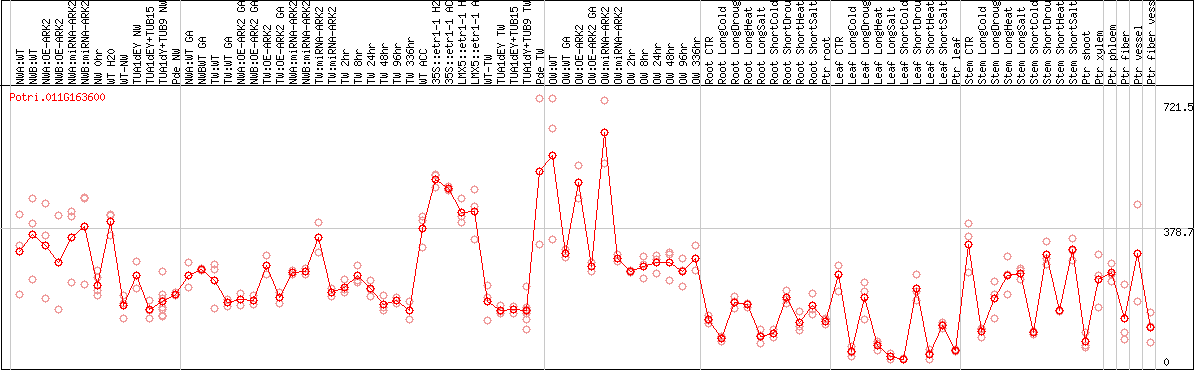
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G163600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
