Potri.011G164800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.011G164800.1 pacid=42781944 polypeptide=Potri.011G164800.1.p locus=Potri.011G164800 ID=Potri.011G164800.1.v4.1 annot-version=v4.1
ATGCCTGCTCCTCTAAGGAATCTCTTATTTTCTCCCAGTATCTTCTCCTTCACCTTACTTCTATCTATAAACTCTCTTTTCTTTCGTTCTTGTTACTCCA
TTGATGAACAAGGCCAAGCCCTTTTAGCATGGAAGAACAGTTTAAACACCTCCACAGATGTACTCAATTCTTGGAACCCTTTAGACTCAAGTCCATGCAA
ATGGTTTGGGGTCCATTGCAACTCAGATGGCAACATTATAGAGATAAACTTGAAAGCAGTAGATTTGCAAGGCCCCCTGCCTTCAAATTTTCAGCCTCTC
AAATCCTTGAAGAGCCTTATCCTTTCATCAACCAATCTTACTGGTGCAATCCCGGAAGCGTTCGGAGATTATCTTGAGCTTACTCTCATTGATCTTAGTG
ATAATTCTCTCTCAGGTGAAATCCCAGAGGAAATATGCAGACTAAGAAAGTTGGAAACTTTGTCTCTCAATACAAATTTTCTTGAAGGTGCCATTCCTTC
TGATATTGGAAACCTTTCAAGTCTAGTGAATCTGACTCTCTTTGACAATCAACTCAGCGGCGAAATTCCACAGAGCATTGGAGCATTGAGAAGATTACAG
ATTTTTCGAGCAGGTGGGAATAAAAATGTTAAGGGTGAGCTGCCTCAGGAGATTGGAAACTGCACCGAATTGGTTGTTTTAGGCCTCGCCGAAACCAGCA
TTTCTGGGAGTCTTCCTTCATCAATTGGGATGCTGAAAAGGATTCAAACTATAGCCATTTACGCAACACTATTATCAGGTGCTATCCCAGAAGCGATTGG
AGATTGCAGTGAGTTGCAGAACCTGTATCTATATCAGAATTCTATATCCGGTCCAATCCCAAGGCGAATTGGGGAACTCAGCAAGCTTCAGAGTCTGCTC
TTGTGGCAGAACAGCATAGTTGGGGCAATCCCGGATGAGATTGGAAGTTGCACAGAGCTTACAGTCATTGATTTATCCGAGAATCTTCTAGCAGGAAGCA
TACCAAGAAGCTTTGGAAACCTTTTGAAGCTTGAAGAGCTTCAGCTGAGTGTTAATCAGTTATCAGGTACCATACCTGTTGAAATCACAAACTGCACAGC
TCTCACTCATCTTGAAGTAGACAACAATGGAATCTCCGGTGAGATTCCTGCTGGTATCGGCAACCTAAAGAGCTTGACTCTATTCTTTGCCTGGAAGAAC
AATCTCACAGGAAATATCCCAGAGTCTCTTTCAGAGTGTGTCAATCTTCAGGCTCTTGATCTTTCATATAACAGTCTTTTTGGTTCAATACCAAAGCAGG
TTTTCGGATTGCAAAATCTCACCAAGCTACTGATACTTTCCAATGAATTATCGGGCTTTATACCGCCTGATATAGGAAACTGCACCAACTTGTATAGGTT
GAGGCTGAATGGCAACAGGTTAGGAGGTACTATTCCATCGGAGATCGAAAAGTTGAAGAGTTTGAATTTCATTGATTTGAGCAACAATCTTCTTGTTGGA
AGAATCCCTTCATCAGTATCTGGATGTGAAAATCTTGAGTTTCTTGATCTCCATTCAAATGGGATCACTGGTTCTGTACCTGATACTCTGCCTAAAAGCC
TACAGTATGTGGATGTTTCAGACAATAGGCTTACAGGTTCACTGGCTCATAGCATTGGGTCATTGATTGAATTAACCAAGCTTAATTTGGCGAAGAATCA
ACTTACAGGAGGAATTCCAGCAGAGATATTGTCATGCAGTAAGCTACAATTGCTTAACCTTGGAGACAATGGTTTCTCTGGAGAAATTCCAAAGGAACTT
GGTCAAATTCCATCACTGGAAATCTCTCTCAACCTAAGCTGCAACCAGTTTTCTGGCAAGATCCCATCGCAATTCTCTGATCTCAGCAAGTTGGGAGCAC
TTGACATCTCTCACAACAAGCTTGAAGGAAGCTTGGATGTTCTTGCAAATCTCCAAAACCTCGTTTCCCTTAATGTCTCCTTCAATGACTTCTCTGGTGA
ATTGCCTAACACCCCATTTTTTCGAAAGCTTCCGATAAGTGACCTTGCTTCGAATCAAGGACTCTACATTTCTGGAGGTGTAGCAACACCAGCAGACCAC
TTGGGACCTGGAGCACACACTAGATCAGCAATGAGACTACTGATGTCTGTGCTTCTCAGTGCTGGTGTAGTACTGATACTACTAACAATCTACATGCTAG
TTCGTGCTCGTGTTGATAATCATGGGCTGATGAAAGATGATACATGGGAAATGAATTTGTATCAGAAGCTTGAATTCTCCGTCAACGACATTGTTAAAAA
TCTAACATCATCTAATGTCATAGGCACTGGCAGCTCTGGAGTTGTGTACAGGGTGACCCTTCCAAATTGGGAGATGATAGCTGTAAAGAAGATGTGGTCA
CCAGAAGAGTCTGGAGCATTCAATTCTGAAATTCGGACACTCGGTTCAATCAGACACCGAAATATTGTCCGGCTTCTAGGCTGGTGTTCAAACAAGAATT
TGAAATTGCTTTTCTATGATTACCTCCCAAATGGAAGCTTGAGCTCCCTCCTTCATGGTGCTGGCAAGGGAGGAGCAGAGTGGGAAGCTAGATATGATGT
TTTGTTAGGTGTGGCACACGCACTTGCATACCTGCACCATGATTGTGTGCCACCTATTCTACATGGGGATGTTAAAGCCATGAATGTGCTTTTAGGACCA
GGCTACGAACCTTACCTAGCTGACTTTGGCCTAGCAAGAGTGGTAAATAACAAATCTGATGATGATTTATGCAAGCCAAGTCCAAGACCTCAACTTGCTG
GTTCTTATGGATATATGGCTCCAGAGCATGCATCGATGCAGCGTATTACCGAGAAAAGTGATGTGTACAGTTTTGGGGTGGTGCTTTTAGAGGTTTTAAC
AGGAAGGCATCCATTAGACCCAACTTTGCCTGATGGTGCACACTTAGTTCAATGGGTTAGGGAACACTTAGCAAGCAAGAAAGATCCAGTTGACATCCTT
GATTCAAAGCTTAGAGGAAGAGCTGACCCTACAATGCACGAAATGCTCCAAACATTAGCAGTTTCATTCCTGTGCATCAGTACTCGAGCTGATGATCGTC
CAATGATGAAGGACGTTGTTGCAATGCTGAAGGAGATTAGGCACGTTGAGACCGTAAGGCCAGAACCTGACCTGTCAAAGGGGGTTAATTTGACAGCAGT
TCGCTCATCACCTCCTGCTAAGATTGTAGTTTCACAAGGATCTTCTAATTGTTCCTTTGCCTTCTCAGATTATTCAATCTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.011G164800.1 pacid=42781944 polypeptide=Potri.011G164800.1.p locus=Potri.011G164800 ID=Potri.011G164800.1.v4.1 annot-version=v4.1
MPAPLRNLLFSPSIFSFTLLLSINSLFFRSCYSIDEQGQALLAWKNSLNTSTDVLNSWNPLDSSPCKWFGVHCNSDGNIIEINLKAVDLQGPLPSNFQPL
KSLKSLILSSTNLTGAIPEAFGDYLELTLIDLSDNSLSGEIPEEICRLRKLETLSLNTNFLEGAIPSDIGNLSSLVNLTLFDNQLSGEIPQSIGALRRLQ
IFRAGGNKNVKGELPQEIGNCTELVVLGLAETSISGSLPSSIGMLKRIQTIAIYATLLSGAIPEAIGDCSELQNLYLYQNSISGPIPRRIGELSKLQSLL
LWQNSIVGAIPDEIGSCTELTVIDLSENLLAGSIPRSFGNLLKLEELQLSVNQLSGTIPVEITNCTALTHLEVDNNGISGEIPAGIGNLKSLTLFFAWKN
NLTGNIPESLSECVNLQALDLSYNSLFGSIPKQVFGLQNLTKLLILSNELSGFIPPDIGNCTNLYRLRLNGNRLGGTIPSEIEKLKSLNFIDLSNNLLVG
RIPSSVSGCENLEFLDLHSNGITGSVPDTLPKSLQYVDVSDNRLTGSLAHSIGSLIELTKLNLAKNQLTGGIPAEILSCSKLQLLNLGDNGFSGEIPKEL
GQIPSLEISLNLSCNQFSGKIPSQFSDLSKLGALDISHNKLEGSLDVLANLQNLVSLNVSFNDFSGELPNTPFFRKLPISDLASNQGLYISGGVATPADH
LGPGAHTRSAMRLLMSVLLSAGVVLILLTIYMLVRARVDNHGLMKDDTWEMNLYQKLEFSVNDIVKNLTSSNVIGTGSSGVVYRVTLPNWEMIAVKKMWS
PEESGAFNSEIRTLGSIRHRNIVRLLGWCSNKNLKLLFYDYLPNGSLSSLLHGAGKGGAEWEARYDVLLGVAHALAYLHHDCVPPILHGDVKAMNVLLGP
GYEPYLADFGLARVVNNKSDDDLCKPSPRPQLAGSYGYMAPEHASMQRITEKSDVYSFGVVLLEVLTGRHPLDPTLPDGAHLVQWVREHLASKKDPVDIL
DSKLRGRADPTMHEMLQTLAVSFLCISTRADDRPMMKDVVAMLKEIRHVETVRPEPDLSKGVNLTAVRSSPPAKIVVSQGSSNCSFAFSDYSI
|
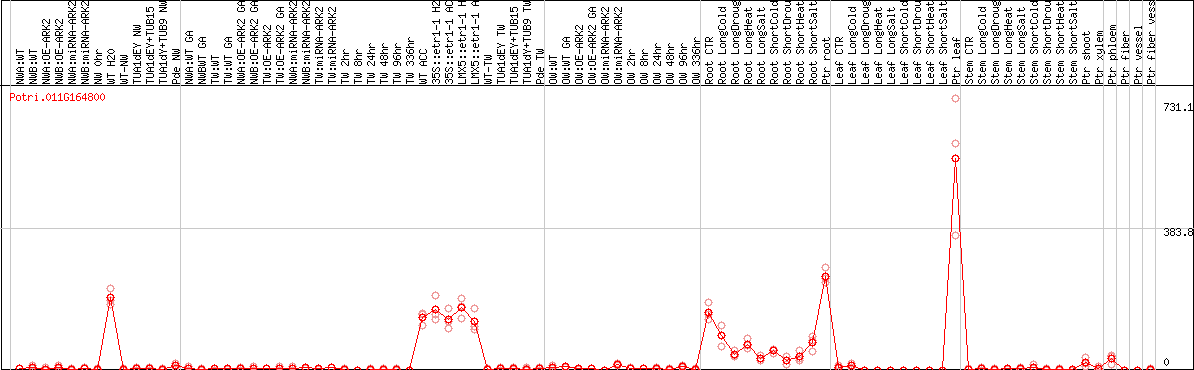
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G164800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
