External link
Symbol
UBC.8
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G64230 295 / 1e-104
UBC28
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
AT5G41700 293 / 4e-104
ATUBC8, UBC8
ARABIDOPSIS THALIANA UBIQUITIN CONJUGATING ENZYME 8, ubiquitin conjugating enzyme 8 (.1.2.3.4.5)
AT4G27960 291 / 6e-103
UBC9
ubiquitin conjugating enzyme 9 (.1.2)
AT5G53300 290 / 1e-102
UBC10
ubiquitin-conjugating enzyme 10 (.1.2.3.4)
AT3G08690 290 / 2e-102
ATUBC11, UBC11
ubiquitin-conjugating enzyme 11 (.1.2)
AT5G56150 288 / 6e-102
UBC30
ubiquitin-conjugating enzyme 30 (.1.2)
AT2G16740 270 / 6e-95
UBC29
ubiquitin-conjugating enzyme 29 (.1)
AT3G08700 251 / 2e-87
UBC12
ubiquitin-conjugating enzyme 12 (.1)
AT3G13550 167 / 8e-54
EMB144, COP10, CIN4, FUS9
FUSCA 9, EMBRYO DEFECTIVE 144, CONSTITUTIVE PHOTOMORPHOGENIC 10, CYTOKININ-INSENSITIVE 4, Ubiquitin-conjugating enzyme family protein (.1.2)
AT1G78870 151 / 9e-48
UBC35 ,UBC13A
UBIQUITIN CONJUGATING ENZYME 13A, ubiquitin-conjugating enzyme 35 (.1.2.3)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039323
293 / 5e-104
AT1G64230 303 / 8e-108
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
Lus10027570
293 / 5e-104
AT1G64230 303 / 8e-108
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
Lus10014187
292 / 2e-103
AT1G64230 303 / 7e-108
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
Lus10022726
292 / 2e-103
AT1G64230 303 / 7e-108
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
Lus10038827
292 / 2e-103
AT1G64230 301 / 3e-107
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
Lus10014942
292 / 2e-103
AT1G64230 301 / 3e-107
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
Lus10032352
291 / 3e-103
AT5G53300 302 / 2e-107
ubiquitin-conjugating enzyme 10 (.1.2.3.4)
Lus10021385
291 / 5e-103
AT1G64230 302 / 1e-107
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
Lus10033937
290 / 2e-102
AT5G53300 300 / 2e-106
ubiquitin-conjugating enzyme 10 (.1.2.3.4)
Lus10009422
285 / 1e-100
AT3G08690 285 / 2e-100
ubiquitin-conjugating enzyme 11 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0208
UBC
PF00179
UQ_con
Ubiquitin-conjugating enzyme
Representative CDS sequence
>Potri.011G168200.7 pacid=42781371 polypeptide=Potri.011G168200.7.p locus=Potri.011G168200 ID=Potri.011G168200.7.v4.1 annot-version=v4.1
ATGGCTTCCAAGAGGATTAACAAGGAGTTGAAAGACCTGCAGAAGGACCCTCCTGCTTCATGCAGTGCTGGTCCTGTTGCTGATGATATGTTCCACTGGC
AAGCAACAATTATGGGCCCAGCTGACAGCCCATTTACTGGTGGTGTATTCCTTGTGTCCATTCACTTTCCTCCAGATTACCCATTCAAGCCACCCAAGGT
CTCCTTCCGTACAAAGGTTTTCCACCCTAACATCAACAGCAATGGAAGCATTTGTCTTGACATTCTCAAAGAGCAGTGGAGCCCTGCCCTTACAATATCC
AAGGTTCTTCTGTCTATCTGCTCACTGCTGACTGATCCAAACCCGGATGATCCCCTGGTGCCTGAAATTGCTCACATGTACAAGACAGACAGAGCCAAGT
ATGAAACTACTGCACGGTCCTGGACCCAGAAATATGCCATGGGCTAG
AA sequence
>Potri.011G168200.7 pacid=42781371 polypeptide=Potri.011G168200.7.p locus=Potri.011G168200 ID=Potri.011G168200.7.v4.1 annot-version=v4.1
MASKRINKELKDLQKDPPASCSAGPVADDMFHWQATIMGPADSPFTGGVFLVSIHFPPDYPFKPPKVSFRTKVFHPNINSNGSICLDILKEQWSPALTIS
KVLLSICSLLTDPNPDDPLVPEIAHMYKTDRAKYETTARSWTQKYAMG
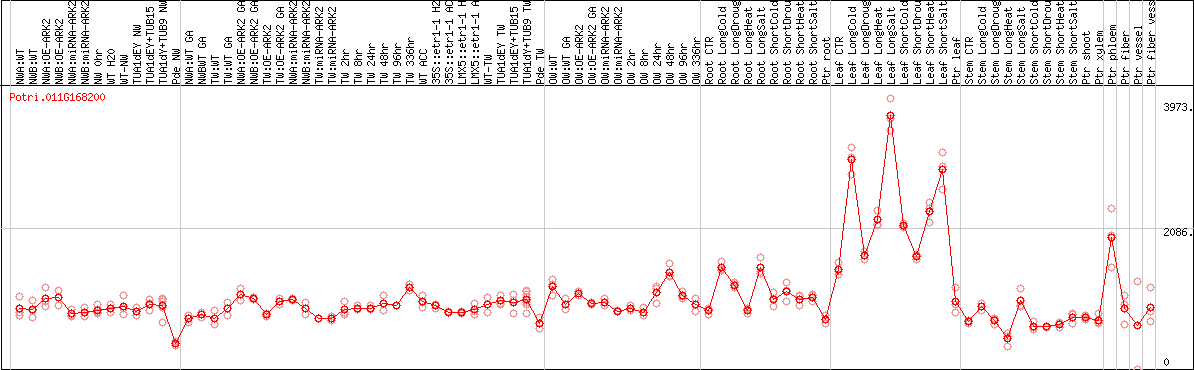
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.011G168200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.