Potri.012G007000 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.012G007000.4 pacid=42783245 polypeptide=Potri.012G007000.4.p locus=Potri.012G007000 ID=Potri.012G007000.4.v4.1 annot-version=v4.1
ATGATGGGTTCTGGGGTTCCTTATGGTTACAATAATGGAGGTGGATCATCATTTTCCTCTTCAAATTTATCACCTTTAGCCCCGCCTTTCACTGTTGATA
GGTCTGTAGCAAAACCCCTTTTGGATTTAACTGAGCCCACTTATCCTGTTAGTTTGAATCCCTCTTTACACAATTGGGCTACCTCTAACTCTCACATTCC
CAATTCAAGACCCGATCTTTTTCCCCTTCCCAACCTGGAATTTAATTCCATACCGTCCCCAAATGTGTTTGGATACTCATCTCCTACTCCCCAGGTGACG
TCTAAGAACCACCCTCTTGTTTTGGCATCGACTGATGCTGTTTTGTATGGTCAGAGTAACCCTAGCCTTGTTGAAGCTGTACCGTATTATCCGTCCTCGT
ATGTCTCGCCTGCAATTGGTAGTGATGGTCACTTGAAAATCCCCCACCAATCTGGTTATGAATTGCTGTCTAATTCTTATGTTGGAACGTCTAATGGATC
ATCTCACGATGATTATACTCAAAGCTCATTGGGTTTGGAGCATGCGACTCAGTGGAGTGGCTTATGGGAGGGAGTTACTGATTGGAATCAGAGCAAAAAA
TTGCAACTTGATGGAGGTTTTTGTGAGAAGGAGAATTTTATTAATCAAGGGTTTTCTGCATTCAAAGATGTGAGCAAATGTGAAGAAACTTCCCTTGGAA
TTGACATGGTTGGAAGGCAAATGCACACCGGGTCTGCAAGCACAGGACAACTGGATTATAAAGCCTTCCTTGTTGAAAAACCCAAGTCCATGCCTACAAC
ACCACCATCATTGATATTCCCTCCAACAGCACCACAAGCCTATCCTCAAGTGTCATCCTCAAATGTAGTCAATTCTCCGAACAATCAAATGCGGCATGTC
ACTTCATATGGAAAGTCCTCAAGGAAGCGTGATGCTAGTTCAAATGATAGGATGCCAATGATGAAGCCTTCACCTGCTGTTGTCATTAGACCCCCGGGAC
AAGATAGATATTCATTTAAAAATATAAATGCTGGTACTGATGGTGATGAGAAGGATTTTGCTGGCAATAACACCTCTTTTGCGCAAGAGCCTAATCCATT
TATTAGCTCCAAAGGCAAAGTTTGCTATGATTCCAGCCAAGTTAATTTTCATCTGAAACAAAATGATGATTCCTTTGCAGAAGTTCCCTCCAAAAACCAT
GAAGAGTTGCTTAGCAACAAAAATATTTCTATTGATTTCTTGGATAAACTTTTCAGAGAAAAAATGGAAAATAGAGTTCCTTGCAAAAATCTAGATTTCT
TCAATTTGGCTATGGATGGTCATGAGGCTGCTGGTTCTGTAGAAATTACTTCTGAGAGTTTAGATCATTACTTCCCTGCTGTGGACTCGCCATGCTGGAA
AGGGGCTCCAGTTTCTCTTCCATCTGCATTCAAAGGTTCTGAGGTTGTGAATCCACAGAATAAGGTAGAGGCATGCAATGGCTTGAATCTGCAAGGCCCT
CAGATTTCCCCTTCCACTACTAATGATGCTGTAAAAGACTGCCCTGAGAAACAAAGTAATATCTCCATGACCTTCAATAATGAGAGTCTGGAGCATCGTC
CAGCCTCTTCTTTTAAGAGGCCTTTAGTTGCTAATGTGTTGTTTAGAGAAGGAATTGATGATGCTGTGAAATATGGACCCTGCCAAAGGAAATCAAGCTA
TTGCAATGAAGCTCAAATTTCTGATGTCATTGATGAACCTAGGAAAGAAAGTATACTGCCTGATTTTAAACCTGTCCACACTAAGCAGAAAAGTCTTGAA
GAAGGTGAATGGCCATCTAAGAAAAATTCTGATGTTGCAGGTGTTAGAAGGAAAATCAACGACAACCCTGATGATTGTTCATCTCATGTGCCATATCATG
CTATAGAACATGTCTTATGTTCACCACCTTCCTCAGAACATGCTCCTGCTCAGCACACTCAATCACAGGTTGGGGAATCATCGTCAAAGATGCATGCTCG
AACACTTGTTGATACAATGCACAACCTCTCAGAATTGCTTCTATTTTATTCTTCAAATGACACTTGTGAATTGAAGGATGAAGATTTTGATGTTCTTAAC
GATGTGATCAACAATCTTGATATTTTTATATCCAAGAATTCAGAAAGGAAGAATTCAACACAAGAATCATTGATTCCTCGGCGAGCCACTTCTCAATCTC
CTGGAAAGTTATCTGAGCTTTACAAGGGCCAACTTGAGTTCCAGCATTTTGAGGACGAGAAAGAGTGTAAAATTGTGTCTGATGAGAGAAAGGAGAAACT
GTCAAATTTCGTTTCTATGAGGGGTGCTACAGATACAGTCAAAGACGATAACGTGACCCAGGCTATAAAGAAGGTTCTTGCCCAGAATTTTCCGATCAAA
GAAGAATCAGAGTCTCAAATTCTTCTGTATAAGAATCTATGGCTTGAAGCAGAAGCTTCACTATGTGTGGTGAATTGTATGGATCGCTTTAATCGATTGA
AGATAGAGATAGAGAAAGGCAGCTCCCAAAAAGTAAATGAATTTTCTTCTGCAGCCCCAGTTGTACCTGAAAATTCTATGATCATGGAGAATCTTTTGGG
GCCTAAGGTTTCATCTGACATATTGCCTGCTGAAGATGAGGGTAGCCCTGTTCACAATGTTCCAGATTCATCTATCTTGAGCAGAAACAGCCATTCAGAT
GATGTCATGGCCAGATTTCATATCATAAAAAGCCGGGTTGATGACTCTAATTCTCTGAATACTTCAGCCATGGATTTGTCAAGTCCTAAGGTTTCTCCTG
ATCTGAACAAGGTTGACAAATTTGCACATGACACAAAAGATAGCTCTAAGTCCCATATATCTTTCCAGGATTCTATAAGGGGAGCAAGCAGTCATGCAGA
TAATGTTATGGACAGATTTCATATCTTAAAATGCCGGGTTGAAAACTCTAGTTCTGTGAATACAGCAACTGGGGGGATATTGGCAAGTTCTATGGTTTCT
CCTGATCAGAACCAGGTTGACAAGTTGGCACATGATACAAAAGATAGCATCATGTCCTATACCATCCAGGATTCCCCTATGTCAGGAAGGAGCAGCCATG
CAGATGATGTTATGACCAGATTTTGTATTTTAAATGGCCGGGATGACAACTCTAATTCTGTGACTATATCAGCTGTAGAGAAATTGTCAAGTTCCAAGGT
TTCCTCTGATCTGAACAAGGTCAGCAAATTGACAGATGATACAAAAGATAGCATAAAGGCAGATGTAACCACCCAGGATTCTTCCATGTCAAGTGCATCA
AGCCAGGCAGAAGATGTTGAAGCATCTGTTATTTTGAAACACAGGGATGGCAATTCAAGTTCCTTGGATATGGAAGAGCACCAGCGAGTGAGTATTGATA
ATGGGTACATGGATCTGATAAGGCTTGCTCGAATGAATAAAGATGGAACGAAAGATAGAACTTTGGATGTAAATATGGAACCTCTCATCCCAAATTTTCG
AGCTGACTCTACTGAAGACAAACCTACTGTGAAGGAGTTTCGTTTATTCATAAATGATGATGTTGAGACTCAATCTCGCCTCACCGATAGGTTCGGAGAC
CAATCCCATGCTGGCTGGTATGATAGCTGTTCCTCAGATTGGGAACATGTCTTGAAGGAAGAGCTTGCAGGGCAGGGTTATTAA
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.012G007000.4 pacid=42783245 polypeptide=Potri.012G007000.4.p locus=Potri.012G007000 ID=Potri.012G007000.4.v4.1 annot-version=v4.1
MMGSGVPYGYNNGGGSSFSSSNLSPLAPPFTVDRSVAKPLLDLTEPTYPVSLNPSLHNWATSNSHIPNSRPDLFPLPNLEFNSIPSPNVFGYSSPTPQVT
SKNHPLVLASTDAVLYGQSNPSLVEAVPYYPSSYVSPAIGSDGHLKIPHQSGYELLSNSYVGTSNGSSHDDYTQSSLGLEHATQWSGLWEGVTDWNQSKK
LQLDGGFCEKENFINQGFSAFKDVSKCEETSLGIDMVGRQMHTGSASTGQLDYKAFLVEKPKSMPTTPPSLIFPPTAPQAYPQVSSSNVVNSPNNQMRHV
TSYGKSSRKRDASSNDRMPMMKPSPAVVIRPPGQDRYSFKNINAGTDGDEKDFAGNNTSFAQEPNPFISSKGKVCYDSSQVNFHLKQNDDSFAEVPSKNH
EELLSNKNISIDFLDKLFREKMENRVPCKNLDFFNLAMDGHEAAGSVEITSESLDHYFPAVDSPCWKGAPVSLPSAFKGSEVVNPQNKVEACNGLNLQGP
QISPSTTNDAVKDCPEKQSNISMTFNNESLEHRPASSFKRPLVANVLFREGIDDAVKYGPCQRKSSYCNEAQISDVIDEPRKESILPDFKPVHTKQKSLE
EGEWPSKKNSDVAGVRRKINDNPDDCSSHVPYHAIEHVLCSPPSSEHAPAQHTQSQVGESSSKMHARTLVDTMHNLSELLLFYSSNDTCELKDEDFDVLN
DVINNLDIFISKNSERKNSTQESLIPRRATSQSPGKLSELYKGQLEFQHFEDEKECKIVSDERKEKLSNFVSMRGATDTVKDDNVTQAIKKVLAQNFPIK
EESESQILLYKNLWLEAEASLCVVNCMDRFNRLKIEIEKGSSQKVNEFSSAAPVVPENSMIMENLLGPKVSSDILPAEDEGSPVHNVPDSSILSRNSHSD
DVMARFHIIKSRVDDSNSLNTSAMDLSSPKVSPDLNKVDKFAHDTKDSSKSHISFQDSIRGASSHADNVMDRFHILKCRVENSSSVNTATGGILASSMVS
PDQNQVDKLAHDTKDSIMSYTIQDSPMSGRSSHADDVMTRFCILNGRDDNSNSVTISAVEKLSSSKVSSDLNKVSKLTDDTKDSIKADVTTQDSSMSSAS
SQAEDVEASVILKHRDGNSSSLDMEEHQRVSIDNGYMDLIRLARMNKDGTKDRTLDVNMEPLIPNFRADSTEDKPTVKEFRLFINDDVETQSRLTDRFGD
QSHAGWYDSCSSDWEHVLKEELAGQGY
|
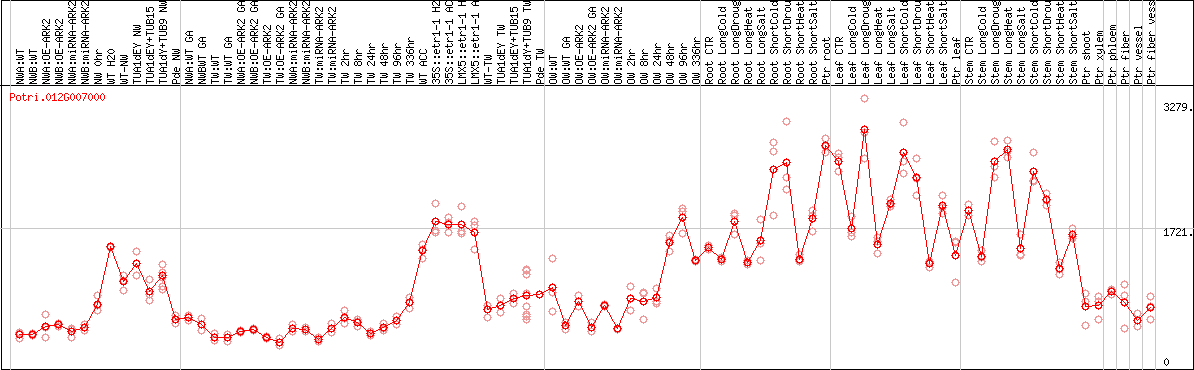
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G007000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
