Potri.012G017500 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.012G017500.2 pacid=42784139 polypeptide=Potri.012G017500.2.p locus=Potri.012G017500 ID=Potri.012G017500.2.v4.1 annot-version=v4.1
ATGGCAATGGAAGTGACCCAGGTGCTTTTGAATGCCCAATCTATAGATGGGAATGTGCGGAAGCATGCAGAAGAAAGCTTAAAACAGTTTCAGGAGCAAA
ACCTGCCCAGTTTTTTGCTCTCTCTCTCCGGAGAGCTAGCTAATGATGAGAAGCCTGTTGACAGTCGTAAATTAGCAGGTTTAATTCTTAAAAATGCGTT
GGATGCAAAGGAACAGCATAGAAAGCTTGAGCTTGTTCAAAGATGGTTGTCGCTGGACAACAATGCAAAGGGTCAAATTAAGGCATGTTTGTTGAAGACA
CTTGCTTCTCCTGTACCAGATGCACGGTCAACTGCATCTCAAGTTATTGCCAAGATTGCAGGGATTGAGTTGCCTCAGAGACAGTGGCCAGAGCTGATTG
GATCACTTTTATCCAATATTCATCAGTTACCTGCTCATGTCAAGCAAGCTACTTTAGAGACTCTTGGATATTTGTGTGAGGAAGTCTCACCTGATGTTGT
AGATCAAGACCATGTAAACAAGATACTTACAGCAGTAGTACAGGGCATGAACGCAACTGAAGGAAACAATGATGTTAGGCTTGCTGCAACCCGAGCACTG
TACAATGCACTTGGATTTGCTCAGGCGAACTTCAGCAATGATATGGAGCGTGATTATATCATGAGAGTGGTCTGCGAGGCAACTTTATCCCCAGAAATGA
AGATAAGACAAGCTGCTTATGAGTGTCTGGTCTCTATTTCATCAACATACTATGAGAAATTGGCTCCTTATATGCAGGATATCTTTAACATCACGGCGAA
GGCCGTGAGGGAAGATGAAGAACCTGTAGCCCTTCAAGCAATTGAATTCTGGAGTTCAATATGTGATGAGGAGATAGATATCTTGGAAGAATATGGGGGT
GATTTCACTGGGGATTCTGATGTTCCTTGCTTTTACTTTATTAAGCAGGCTCTCCCTGCTTTGGTCCCAATGTTGTTGGAGACTCTCCTAAAGCAAGAGG
AGGATCAGGATCAAGATGAAGGAGCTTGGAATATTGCAATGGCTGGTGGAACATGCCTTGGTTTGGTTGCAAGGACTGTGGGAGATGATATTGTGCAGCT
TGTTATGCAATTCATTGAAGATAACATTACAAAACCAGATTGGAGGCATAGAGAGGCAGCTACTTATGCATTTGGCTCGATCTTGGAAGGTCCTTCCCCT
GAGAAACTAACGCCTCTTGTTAATGTTGCTTTGAACTTCATGCTGACTGCTTTGACAAAGGATCCAAATAACCATGTGAAGGATACAACTGCTTGGACCC
TTGGTCGAATCTTTGAATTCCTCCATGGTTCAACTGTGGATACACCGATAATTACTCAAGCAAATTGCCAACAAATTGTCACGGTACTACTCCAGAGCAT
GAAGGACGTTGCCAATGTAGCAGAGAAGGCATGTGGTGCCCTCTATTTCCTTGCCCAGGGTTATGAAGAGGTGACCCCATCATCACCTCTTACTCCTTAC
TTCCAGGAAATTGTCCAGACACTTCTCTTTGTTACCCACAGAGAGGATGCTGGGGAGTCCCGATTAAGGACTGCTGCATATGAAACGTTGAATGAGGTGG
TGAGATGCTCGACTGATGAAACAGCACCAATGGTGCTGCAACTGGTTCCTGTCATCATGACGGAGCTTCACAATACTCTTGAAGGGCAGAAGCTTTCTTC
TGATGAGAGAGAGAAGCAGGGTGAATTGCAAGGGCTTCTCTGTGGTTGCTTACAGGTCATAATTCAGAAGTTAGGCTCCTCTGAGCCAACTAAGTATGTG
TTCATGCAATATGTAGATCAGATTATGGGGCTCTTCCTGAGGGTCTTTGCTTGTAGAAGTGCAACTGTGCATGAGGAGGCCATGTTAGCCATAGGAGCCC
TTGCCTATGCAACTGGTCCTGATTTTGCAAAATACATGCCAGAGTTTTACAAGTACCTAGAAATGGGCCTTCAGAATTTTGAGGAGTACCAAGTCTGTGC
TGTGACTGTCGGTGTGGTGGGTGATATCTGCAGGGCATTGGAAGATAAAATTTTGCCTTACTGTGATGGTATTATGACTCAGCTTCTCAAGGATTTATCA
AGCAACCAGCTGCATAGATCTGTGAAGCCTCCTATTTTCTCTAGCTTTGGCGACATTGCCCTGGCAATAGGAGAGAATTTTGAGAAGTACTTGATGTATG
CCATGCCAATGCTGCAAAGTGCCGCAGAGTTGTCTGCCCACACATCTGTTGCTGACGATGAAATGACTGAATACACAAATTCTTTGAGAAATGGAATTTT
GGAGGCATACTCTGGAATTCTTCAAGGGTTTAAGAATTCTCCTAAAACTCAGCTCTTGATTCCTTATGCACCTCACATCCTGCAGTTCTTGGATAGCATG
TACATGGAGAAAGACATGGATGATGTCGTTATGAAAACTGCAATTGGAGTCCTGGGAGATTTAGCAGATACACTGGGAAGTAATGCTGGCTCTTTGATAC
AGCAATCTCTTTCTAGCAAAGACTTTTTAAATGAATGTTTGTCTTCTGATGACCATATGATTAAGGAATCTGCTGAATGGGCCAAGTTGGCCATCAGTCG
TGCCATTTCTGTTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.012G017500.2 pacid=42784139 polypeptide=Potri.012G017500.2.p locus=Potri.012G017500 ID=Potri.012G017500.2.v4.1 annot-version=v4.1
MAMEVTQVLLNAQSIDGNVRKHAEESLKQFQEQNLPSFLLSLSGELANDEKPVDSRKLAGLILKNALDAKEQHRKLELVQRWLSLDNNAKGQIKACLLKT
LASPVPDARSTASQVIAKIAGIELPQRQWPELIGSLLSNIHQLPAHVKQATLETLGYLCEEVSPDVVDQDHVNKILTAVVQGMNATEGNNDVRLAATRAL
YNALGFAQANFSNDMERDYIMRVVCEATLSPEMKIRQAAYECLVSISSTYYEKLAPYMQDIFNITAKAVREDEEPVALQAIEFWSSICDEEIDILEEYGG
DFTGDSDVPCFYFIKQALPALVPMLLETLLKQEEDQDQDEGAWNIAMAGGTCLGLVARTVGDDIVQLVMQFIEDNITKPDWRHREAATYAFGSILEGPSP
EKLTPLVNVALNFMLTALTKDPNNHVKDTTAWTLGRIFEFLHGSTVDTPIITQANCQQIVTVLLQSMKDVANVAEKACGALYFLAQGYEEVTPSSPLTPY
FQEIVQTLLFVTHREDAGESRLRTAAYETLNEVVRCSTDETAPMVLQLVPVIMTELHNTLEGQKLSSDEREKQGELQGLLCGCLQVIIQKLGSSEPTKYV
FMQYVDQIMGLFLRVFACRSATVHEEAMLAIGALAYATGPDFAKYMPEFYKYLEMGLQNFEEYQVCAVTVGVVGDICRALEDKILPYCDGIMTQLLKDLS
SNQLHRSVKPPIFSSFGDIALAIGENFEKYLMYAMPMLQSAAELSAHTSVADDEMTEYTNSLRNGILEAYSGILQGFKNSPKTQLLIPYAPHILQFLDSM
YMEKDMDDVVMKTAIGVLGDLADTLGSNAGSLIQQSLSSKDFLNECLSSDDHMIKESAEWAKLAISRAISV
|
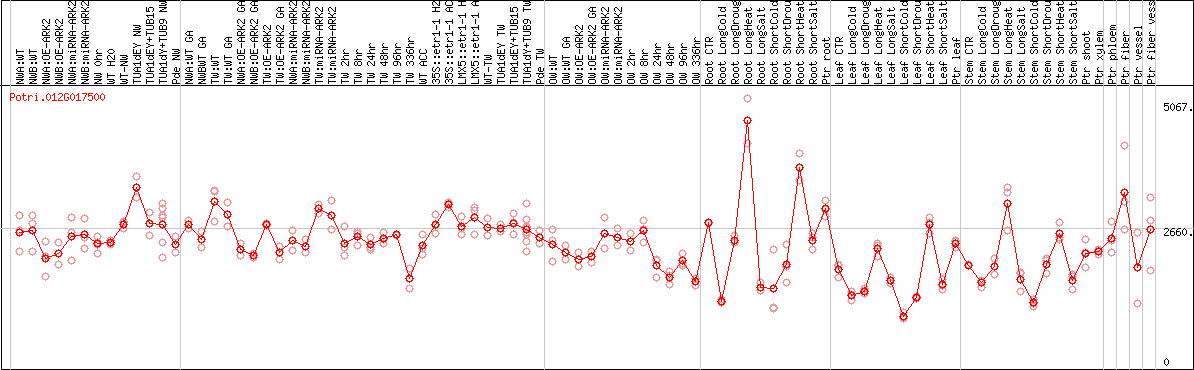
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G017500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
