Potri.012G022700 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.012G022700.2 pacid=42782616 polypeptide=Potri.012G022700.2.p locus=Potri.012G022700 ID=Potri.012G022700.2.v4.1 annot-version=v4.1
ATGAGCTACGCTGCCTACAAGATGATGCACTGGCCAACCACCATAGACACGTGCGTCTCCGGTTTCGTAACCCATTCTCGCTCCGAGTCAGCCCATCTTC
CTCAACTCCACACTGACGATCTTGACTCTGACTGGCCTTCACGCCGCCGCCACGGTGGAGGAATTGGCCCTACTCCGAACCTAATTGTTGCTTCCGGTAA
TGTTCTTGAACTCTATGTTGTCAGGGTTCAAGAAGAGGGGGCGAGGAGTTCGGGTGAATTGAAGCGTGGTGGAGTTATGGATGGTGTTGCTGGAGCTTCT
CTTGAGCTTGTTTGCCATTACAGGTTGCATGGTAATGTTGAGTCAATGGGGGTGTTGTCTGTTGAAGGTGGTGATGATTCTAGGAGAAGGGATTCTATTA
TATTGGCATTTAAAGATGCAAAGATTTCTGTTTTGGAGTTTGATGATTCAATTCATGGACTTCGCACAAGCTCTATGCATTGTTTTGAAGGTCCAGACTG
GCGTCATTTGAAAAGAGGTAGAGAATCATTTGCAAGAGGTCCATTGGTAAAGGTTGATCCTCAAGGAAGATGCGGAGGTGTTCTTGTTTATGATTTGCAG
ATGATAATACTTAAGGCTGCTCAGGCTGGTTCTGCCTTGGTTCAAGATGAAGATGCTTTTGGTTCTGGAGCTGCTATCTCTGCTCATATTGCCTCTTCTT
ACATCATTAATCTTAGAGATTTAGACATGAAACATGTAAAAGATTTCATATTTGTGCATGATTATATTGAACCTGTGGTGGTTGTCCTTCACGAGCGAGA
GCTTACTTGGGCTGGGCGTGTTGTGTGGAAGCATCATACTTGCATGATTTCTGCTCTTAGTATCAGTACAACTTTGAAGCAACCTACCCTTATATGGTCT
ATTGGAAATCTCCCGCATGATGCGTATAAGCTGCTTGCAGTGCCATCTCCCATTGGCGGTGTTCTTGTGATTGGTGTGAACACTATCCATTATCACAGTG
AGTCAGCTTCTTGTGCATTGGCTCTGAACAGCTATGCTGCTTCTGTTGATAGCAGTCAAGAGCTGCCAAGAGCAACTTTCAGTGTGGAACTTGATGCTGC
CAATGCAACTTGGCTATTAAAAGATGTAGCCTTGCTGTCAACAAAAACTGGGGAACTCTTGTTACTGACCCTTGTTTATGATGGAAGGGTTGTGCAGAGA
CTTGATCTTTCCAAGTCTAAAGCTTCAGTACTTACTTCGGATATTACAACACTTGGAAACTCGTTCTTTTTTCTGGGAAGTCGTTTGGGAGACAGTTTGC
TTGTGCAGTTTACTTCTGGATTGGGATCTTCAATGTTGTCACCTGGTTTAAAGGAAGAGGTTGGAGATATTGAAGGTGATTTGCCTTCAGCAAAGCGATT
GAAGGTGTCCTCTTCTGATGCTTTACAAGATATGGTTAGCGGGGAGGAACTTTCCTTGTATAGTTCAGCGCCAAATAATGCAGAATCATCACAGAAAACT
TTCTCATTTACAGTGAGGGATTCGTTGATCAATGTTGGCCCTTTGAAGGACTTTGCATATGGTTTAAGGATTAATGCGGATGCAAATGCAACTGGAATCT
CCAAACAAAGCAACTATGAATTGGTGTGCTGTTCTGGACACGGTAAAAATGGTGCTCTCTGTGTCCTTCAGCAATCAATTCGCCCAGAAATGATTACAGA
GGTTGAACTTCCTGGATGTAAAGGAATTTGGACTGTTTACCACAAGAATGCACGTAGTCATAGTGTTGACTCATTGAAAATGGCTTCAGATGATGAATAT
CATGCATATTTGATCATAAGCATGGAGGCACGGACAATGGTGCTTGAAACTGCTGATCATTTGACGGAAGTCACTGAAAGTGTGGACTATTTTGTGCAAG
GAAGAACAATTGCTGCAGGGAACTTGTTTGGAAGACGTCGAGTTGTTCAGGTGTTTGAACGTGGTGCTCGAATCTTAGATGGTTCTTTTATGACTCAAGA
TTTAAGCTTTGGAGGTTCTAACTCGGAAACAGGTCGTTCTGAGAGTTCCACAGTGATGCATGTGTCTATTGTGGATCCATATGTTTTAGTTAGAATGGCT
GACGGGAGTATTCAGATTCTTGTTGGAGATCCTTCTGCTTGTACAGTTTCTGTAAACACTCCGTCTGCCTTTCAGAGCTCAACGAAATCAGTATCTGCCT
GCACACTGTATCATGATAAAGGGCCAGAGCCATGGCTCCGGAAGACAAGTACAGATGCATGGCTTTCTACTGGAATTAGCGAGGCTATTGATGGTGCTGA
CAGTGGAGCTCATGAGCAGGGTGATATATACTGTGTTGTTTGTTATGAGACTGGTGCCCTTGAAATATTTGATGTGCCAAATTTCAATTCTGTGTTCTTC
GTGGATAAATTTGTCTCTGGAAAAACTCATCTCCTTGATACCTGCACGGGGGAACCTGCTAAAGATATGATGAAGGGGGTGAAAGAAGAAGTTGCCGGTG
CTGGCAGAAAAGAAAGTACTCAAAACATGAAGGTTGTTGAATTGACCATGCTGAGATGGTCTGGACGTCACAGTCGCCCTTTTCTTTTTGGAATATTGAC
TGATGGGACAATTCTTTGTTACCACGCATACCTGTTTGAAGGTCCAGATGGTACTTCTAAACTAGAAGATTCTGTTTCTGCTCAAAATTCTGTTGGTGCT
AGCACCATCAGTGCTTCAAGGCTTAGAAATCTGCGATTCGTTCGGGTCCCTTTGGACACATACACTAGGGAGGAAACATCAAGTGAAACCTCATGCCAAA
GGATTACGACTTTCAAGAATATTAGTGGTTATCAAGGGTTTTTCCTTTCTGGATCAAGACCAGCTTGGTTTATGGTATTTAGAGAGCGACTGCGAGTTCA
TCCACAGCTATGTGATGGATCTATTGTTGCCTTCACTGTTCTTCACACTGTTAATTGTAATCATGGGCTCATTTATGTCACGTCACAGGGCAATTTGAAG
ATTTGTCACCTGTCATCTGTCTCGAGTTATGACAACTACTGGCCAGTGCAAAAGATTCCTTTGAAAGGAACTCCACATCAAGTGACTTATTTTGCAGAGA
GGAATTTATACCCTCTCATAGTTTCAGTTCCTGTTCAAAAGCCAGTAAATCAAGTTCTTTCATCACTGGTTGATCAAGAAGTTGGCCATCAGATTGAGAA
TCATAATTTGAGTTCTGAAGAAATACACCGAACTTACAGTGTAGATGAATTTGAGGTTCGGATTTTGGAACCATCTAATGGCCCTTGGCAAGTTAAGGCT
ACAATCCCAATGCAGACTTCTGAAAATGCATTGACTGTACGAATGGTTTCACTGTTTAATACAAGTACAAAGGAAAATGAAACACTTTTGGCTGTTGGGA
CTGCTTATGTGCAAGGGGAGGATGTTGCTGCTAGAGGACGTATTCTGCTGTTTTCTGTAGTCAAAAATCCAGAAAATTCTCAGATTTTGGTTTCAGAAGT
TTATTCTAAGGAATTGAAAGGTGCTATATCTGCTTTAGCCTCACTTCAAGGCCATCTGTTGATAGCTTCTGGTCCAAAAATTATTTTACACAAATGGACC
GGTACAGAGTTGACTGGTGTTGCATTTTCTGATGCTCCACCATTATATGTCGTGAGCTTAAATATTGTGAAGAATTTTATTCTCTTGGGTGACATTCACA
AAAGCATATACTTTCTTAGTTGGAAGGAGCAGGGGGCTCAGCTCAGCTTGCTGGCGAAGGATTTTGCCTCCCTTGATTGTTTCTCAACAGAGTTTTTGAT
TGATGGAAGTACTTTGAGCCTTGTGGTTTCAGATGAACAAAAGAATGTTCAGATATTCTATTATGCACCAAAGATGTCTGAGAGTTGGAAAGGACAGAAA
CTGCTTTCAAGGGCTGAATTTCATGTTGGTGCCCTTGTTACTAAGTTCATGAGGTTGCAAATGCTTTCCCCTTCACTTGATCGAAGTGGTGCCGCTCCAG
TTTCTGACAAGACCAATCGCTTTGCTTTACTGTTTGGTACTTTGGACGGCAGCATAGGTTGTATTGCACCTCTTGACGAGCTCACATTTCGTAGGCTGCA
GTCACTGCAGAAGAAACTTGTTGATGCTGTTCCCCATGTAGCTGGTTTAAACCCAAAATCATTCCGCCAGTTCCGCTCTGATGGAAAGGCTCACCGACCT
GGCCCTGAAAGCATAGTTGATTGTGAAATGCTTTCCTATTATGAGATGATCCCGTTGGAGGAACAGGTTGAAATTGCCCAACAGATTGGAACAACACGTG
CTCAGATTTTGTCAAATTTAAACGACCTTACTCTGGGCACAAGTTTTCTGTGA
|
|||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.012G022700.2 pacid=42782616 polypeptide=Potri.012G022700.2.p locus=Potri.012G022700 ID=Potri.012G022700.2.v4.1 annot-version=v4.1
MSYAAYKMMHWPTTIDTCVSGFVTHSRSESAHLPQLHTDDLDSDWPSRRRHGGGIGPTPNLIVASGNVLELYVVRVQEEGARSSGELKRGGVMDGVAGAS
LELVCHYRLHGNVESMGVLSVEGGDDSRRRDSIILAFKDAKISVLEFDDSIHGLRTSSMHCFEGPDWRHLKRGRESFARGPLVKVDPQGRCGGVLVYDLQ
MIILKAAQAGSALVQDEDAFGSGAAISAHIASSYIINLRDLDMKHVKDFIFVHDYIEPVVVVLHERELTWAGRVVWKHHTCMISALSISTTLKQPTLIWS
IGNLPHDAYKLLAVPSPIGGVLVIGVNTIHYHSESASCALALNSYAASVDSSQELPRATFSVELDAANATWLLKDVALLSTKTGELLLLTLVYDGRVVQR
LDLSKSKASVLTSDITTLGNSFFFLGSRLGDSLLVQFTSGLGSSMLSPGLKEEVGDIEGDLPSAKRLKVSSSDALQDMVSGEELSLYSSAPNNAESSQKT
FSFTVRDSLINVGPLKDFAYGLRINADANATGISKQSNYELVCCSGHGKNGALCVLQQSIRPEMITEVELPGCKGIWTVYHKNARSHSVDSLKMASDDEY
HAYLIISMEARTMVLETADHLTEVTESVDYFVQGRTIAAGNLFGRRRVVQVFERGARILDGSFMTQDLSFGGSNSETGRSESSTVMHVSIVDPYVLVRMA
DGSIQILVGDPSACTVSVNTPSAFQSSTKSVSACTLYHDKGPEPWLRKTSTDAWLSTGISEAIDGADSGAHEQGDIYCVVCYETGALEIFDVPNFNSVFF
VDKFVSGKTHLLDTCTGEPAKDMMKGVKEEVAGAGRKESTQNMKVVELTMLRWSGRHSRPFLFGILTDGTILCYHAYLFEGPDGTSKLEDSVSAQNSVGA
STISASRLRNLRFVRVPLDTYTREETSSETSCQRITTFKNISGYQGFFLSGSRPAWFMVFRERLRVHPQLCDGSIVAFTVLHTVNCNHGLIYVTSQGNLK
ICHLSSVSSYDNYWPVQKIPLKGTPHQVTYFAERNLYPLIVSVPVQKPVNQVLSSLVDQEVGHQIENHNLSSEEIHRTYSVDEFEVRILEPSNGPWQVKA
TIPMQTSENALTVRMVSLFNTSTKENETLLAVGTAYVQGEDVAARGRILLFSVVKNPENSQILVSEVYSKELKGAISALASLQGHLLIASGPKIILHKWT
GTELTGVAFSDAPPLYVVSLNIVKNFILLGDIHKSIYFLSWKEQGAQLSLLAKDFASLDCFSTEFLIDGSTLSLVVSDEQKNVQIFYYAPKMSESWKGQK
LLSRAEFHVGALVTKFMRLQMLSPSLDRSGAAPVSDKTNRFALLFGTLDGSIGCIAPLDELTFRRLQSLQKKLVDAVPHVAGLNPKSFRQFRSDGKAHRP
GPESIVDCEMLSYYEMIPLEEQVEIAQQIGTTRAQILSNLNDLTLGTSFL
|
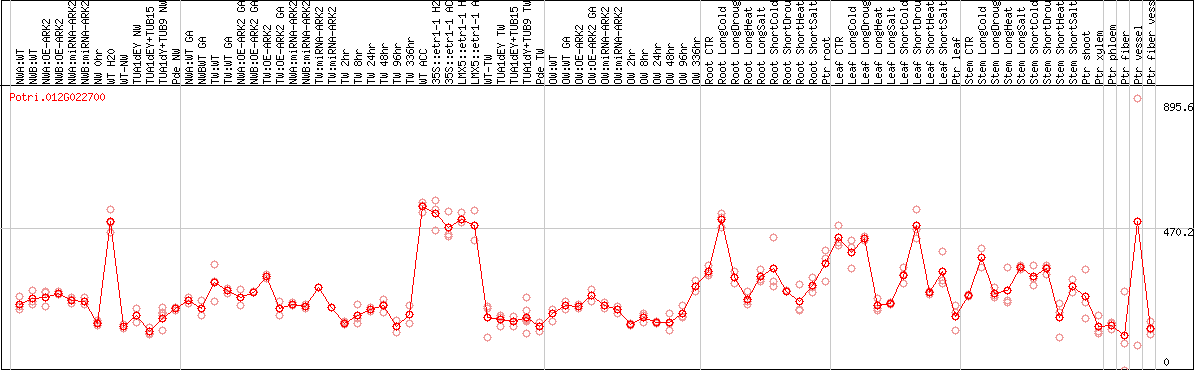
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G022700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
