Potri.012G022800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.012G022800.1 pacid=42782982 polypeptide=Potri.012G022800.1.p locus=Potri.012G022800 ID=Potri.012G022800.1.v4.1 annot-version=v4.1
ATGTTTCTGCTTTGCCAATGTTTTGTTCTAATCTCTCTTACCCTTTTCCAAGCTTCTTCACCAGCCTCATTACCTTCATCAGCAATCGACCTTCTCAAGT
TCCAAGATAGTCTTCCGTTACTTTCTCAAAAGCTCTTGCCATGGAACCAATCAAGCTCATCTTCCTCTCCTTGCCAATGGCCTGGAGTTTCTTGCTATCC
CAACAAAAGTTTTCAAGTCAAGGCCTTAAACCTGTCAGGATATGGTCTTTCTGGTGTCTTGAACAACTCCATTTCTTATCTCTGCCGCCATAAACATTTG
GTCTTACTTGATCTTAGTGGAAACCATTTCACCGGTGTTATTCCTCACTTGCTTGTAAACTGTGGTCAACTAAATACCATTCTTCTCAATGACAATGGCT
TGGAAGGTTCAATTCCTGCTGATGTTTTCAAATCAAAGAAGCTTGTACAGTTAGACTTCGGCTACAACTCTCTTTCTGGTAATATTCCTCCTGAGGTAAG
CTTTTGCACCAACCTTGAATACCTAGGGTTGTATAACAATTATTTGAGTGGAGCGGTTCCAAGTGAAATATTTTCTCTGCCAAAGTTGAATTTTATGTAT
CTTAACACAAATAACCTAACAGGGTTGTTGCCAAATTTCTTACCTTCCTGCGCAATCTCTGATCTTTTGATTCATGAGAATGCATTTTCTGGATCTCTTC
CTTCTACTCTCAGTAATTGCCAAAACCTTACCGTGTTTATCGCCTCTCAAAACAATTTTGAAGGAGTCATTGCACCAGAGATTTTCAAGGGTCTTTTGCA
ACTTGAAGTTCTTTATCTTGATGGAAACAAACTTGAAGGGGAAATTCCAGAAACTTTATGGGGTCTTGAAAATTTACAAGAGCTTGTTCTTTCAGGGAAC
AAGCTGAATGGCACAATATCTGAGAGGATTTCTCAATGCCCTCAGCTTATGACCATTGCTCTTTCTGGTAATAATCTTGTTGGTCATATTCCTAGATTAG
TTGGAACTCTCCAATATTTGACCAATCTAATTCTCTTTGACAATAAGCTTGATGGCTCATTGCCTGCTGAGCTTGGAAATTGCAGTTCACTTGTTGAATT
TAGGCTTCAGAATAACCTTATAGGAGGGAACATCCCTCCTGAAATATGTAATCTTGAGAATCTTGAGGTTCTTTTCTTGTCCAATAATTTTGTTGAAGGC
CATATTCCAAGGCAGATAGGGAGATTGAGCAACCTTAAAATATTAGCTCTGTATAGCAATAATTTGAGTGGTATAATCCCATCTGAAATCACCAATTTTA
CAAAACTCACCTATCTTTCATTTGCTCATAATGATCTTACTGGTGAGGTACCATTTGATCTTGGGAAAAATTCTCCAGATCTTGATAGACTCGACCTAAC
CTCGAACCACCTTTACGGACCAATTCCTCCTAATGTATGCAATGGCAATAATCTGAGAGTTCTGACCCTTGGGGACAACAGGTTTAATGGGATCTTTCCG
GTTGAGATTGGCAAGTGTTTGTCTCTAAGGAGGGTGATTCTTAGTAACAATCTTCTAGAGGGGAGTATACCAACCGATTTGGAGAGGAACTCTGGGATTT
CTTACTTGGAAGTTAGAGGAAACTTGATTGAAGGAAAAATACCTGCAGTGTTTGGTTCTTGGAGCAATCTATCGATGATTGATTTTTCAGGGAACAAATT
CTCTGGCTCTATACCTCCTGAGCTTGGAAAGCTTGCAAATCTTCAAGCCTTGAGACTTTCTTCTAATAATTTAACGGGAAGCATTCCCTCTGACCTCAGT
CATTGCAGAAAGTTCATTAAAATAGACTTGAGTAAGAATCAGTTGTCAGGAAAAATTCCTTCAGAGATTACTTCATTGGAAAAATTAGAAAGCCTTCTCC
TTCAAGAGAACAAACTTAGTGGAGCTATTCCTGACTCCTTCTCTCCTCTTCAAGGTCTCTTTGAATTGCAGCTTAGCAGTAACATGCTTGAAGGTCCTAT
TCCTTGCAGTTTGAGTAAGATCAACCATTTTAGTTCAGTTCTTAATCTGAGCTATAATAAGCTTTCTGGTAAAATCCCTGGGTGCCTTGGCAATCTAGAC
AAGTTACAGATTCTTGATCTCTCATGCAACAGCTTCTATGGTGAGATGCCGACAGAACTCAACAACATGATCTCCCTTTACTTTGTCAACATATCATTCA
ATCAACTCTCTGGCAAGCTTCCAACTTCATGGATTAGAATAATGGCTTCATATCCAGGGTCTTTTCTTGGAAATCCTGAACTTTGTTTGCCAGGTAATGA
TGCAAGAGACTGCAAGAATGTTAGAGAAGGCCATACAAGAAGGCTTGACCGGCATGCGTTGGCTGGTGTCATTATATGTGTTGTTATCTCTATGGCATTG
CTTTGTTCTGTGGTTTACATAATAGTGGTTCGAGTCCTTCAACATAAATATCACCGTGATCAATCACTTCTTCGTGAATGTCGATCGCATACTGAAGATT
TGCCAGAAGATTTACAGTTTGAAGATATAATGCGTGCTACAGAAGGCAGGAGTGAGGAATATGTAATTGGGAGAGGCAAGCATGGCACAGTTTACAGGAC
AGAATCCGCAAACTCCAGAAAGCATTGGGCTGTCAAGAAGGTGAGCTTGTCTGGGGATAACTTTAGCCTTGAAATGAGAACTTTGAGCGTAGTCAGGCAT
CGAAATATCGTGAGAATGGGGGGTTACTGCATTAAAGATGGTTACGGGTTCATTGTAACTGAATTCATGCCTGGAGGAACATTATTTGATGTCCTACACA
GACATGAACCACGCATGGCTTTGGATTGGGACACCCGATATCGCATTGCTCTTGGTGTTGCACAAGGCCTTTCTTACCTTCATCATGATTGCGTGCCACA
AATCATTCACAGAGATGTCAAATCAGATAACATTTTGATGGATTCTGAATTGGAACCCAAGGTTGGAGATTTTGGGATGTCAAAAATGCTTCTTGATTCT
GATTCAAGCTCGACGAGGTCTAGAATTGTAGGAACTCTAGGTTACATGGCACCAGAAAATGCATACTCAATCCGATTGACAGAGAAAGTCGATGTATACA
GCTACGGAGTCATTTTGTTAGAGATTGTTTGCAGAAAGTTTCCAGTAGACCCCAGCTTTGAAGAAGGTCTGGATATCGTTTCCTGGACGAGGAAGAAGCT
GCAAGAGAATGATGAATGTGTTTGCTTCCTTGATAGAGAAATCAGTTTCTGGGATAGAGATGAGCAACAAAAGGCTTTGAAACTGCTAGAATTGGCACTT
GAGTGCACTGAATCAGTGGCTGATAAGAGACCATCCATGAGAGATGTAGTAGGTTCTCTTATAAAATTGCATGATAAGCATGAAAGAAGAGTAAATAACA
GAAACAATTCGAACCAAAACATCTTATATCCTTCCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.012G022800.1 pacid=42782982 polypeptide=Potri.012G022800.1.p locus=Potri.012G022800 ID=Potri.012G022800.1.v4.1 annot-version=v4.1
MFLLCQCFVLISLTLFQASSPASLPSSAIDLLKFQDSLPLLSQKLLPWNQSSSSSSPCQWPGVSCYPNKSFQVKALNLSGYGLSGVLNNSISYLCRHKHL
VLLDLSGNHFTGVIPHLLVNCGQLNTILLNDNGLEGSIPADVFKSKKLVQLDFGYNSLSGNIPPEVSFCTNLEYLGLYNNYLSGAVPSEIFSLPKLNFMY
LNTNNLTGLLPNFLPSCAISDLLIHENAFSGSLPSTLSNCQNLTVFIASQNNFEGVIAPEIFKGLLQLEVLYLDGNKLEGEIPETLWGLENLQELVLSGN
KLNGTISERISQCPQLMTIALSGNNLVGHIPRLVGTLQYLTNLILFDNKLDGSLPAELGNCSSLVEFRLQNNLIGGNIPPEICNLENLEVLFLSNNFVEG
HIPRQIGRLSNLKILALYSNNLSGIIPSEITNFTKLTYLSFAHNDLTGEVPFDLGKNSPDLDRLDLTSNHLYGPIPPNVCNGNNLRVLTLGDNRFNGIFP
VEIGKCLSLRRVILSNNLLEGSIPTDLERNSGISYLEVRGNLIEGKIPAVFGSWSNLSMIDFSGNKFSGSIPPELGKLANLQALRLSSNNLTGSIPSDLS
HCRKFIKIDLSKNQLSGKIPSEITSLEKLESLLLQENKLSGAIPDSFSPLQGLFELQLSSNMLEGPIPCSLSKINHFSSVLNLSYNKLSGKIPGCLGNLD
KLQILDLSCNSFYGEMPTELNNMISLYFVNISFNQLSGKLPTSWIRIMASYPGSFLGNPELCLPGNDARDCKNVREGHTRRLDRHALAGVIICVVISMAL
LCSVVYIIVVRVLQHKYHRDQSLLRECRSHTEDLPEDLQFEDIMRATEGRSEEYVIGRGKHGTVYRTESANSRKHWAVKKVSLSGDNFSLEMRTLSVVRH
RNIVRMGGYCIKDGYGFIVTEFMPGGTLFDVLHRHEPRMALDWDTRYRIALGVAQGLSYLHHDCVPQIIHRDVKSDNILMDSELEPKVGDFGMSKMLLDS
DSSSTRSRIVGTLGYMAPENAYSIRLTEKVDVYSYGVILLEIVCRKFPVDPSFEEGLDIVSWTRKKLQENDECVCFLDREISFWDRDEQQKALKLLELAL
ECTESVADKRPSMRDVVGSLIKLHDKHERRVNNRNNSNQNILYPS
|
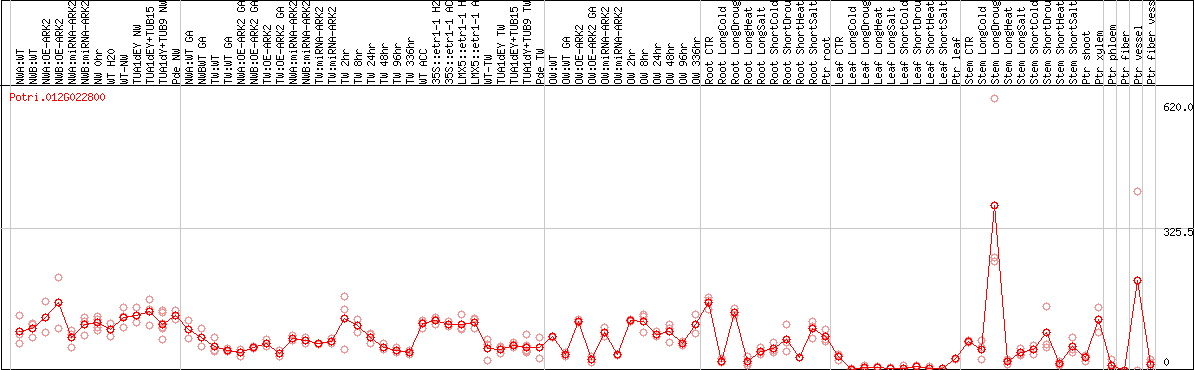
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G022800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
