External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G17260 91 / 4e-22
NAC
ANAC086
NAC domain containing protein 86 (.1)
AT3G03200 88 / 7e-21
NAC
ANAC045
NAC domain containing protein 45 (.1)
AT3G17730 84 / 4e-20
NAC
ANAC057
NAC domain containing protein 57 (.1)
AT5G09330 86 / 5e-20
NAC
VNI1, ANAC082
VND-interacting 1, NAC domain containing protein 82 (.1.2.3.4)
AT5G46590 84 / 6e-20
NAC
ANAC096
NAC domain containing protein 96 (.1)
AT2G33480 83 / 8e-20
NAC
ANAC041
NAC domain containing protein 41 (.1.2)
AT1G65910 85 / 1e-19
NAC
ANAC028
NAC domain containing protein 28 (.1)
AT1G34180 85 / 1e-19
NAC
ANAC016
NAC domain containing protein 16 (.1.2)
AT1G34190 83 / 4e-19
NAC
ANAC017
NAC domain containing protein 17 (.1)
AT5G13180 81 / 7e-19
NAC
VNDIP2, ANAC083, VNI2
VND-interacting 2, NAC domain containing protein 83 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020896
86 / 3e-20
AT5G09330 276 / 2e-88
VND-interacting 1, NAC domain containing protein 82 (.1.2.3.4)
Lus10026373
83 / 2e-19
AT3G17730 348 / 2e-121
NAC domain containing protein 57 (.1)
Lus10042284
83 / 2e-19
AT3G17730 348 / 1e-121
NAC domain containing protein 57 (.1)
Lus10028713
83 / 5e-19
AT1G34190 392 / 2e-130
NAC domain containing protein 17 (.1)
Lus10033493
80 / 3e-18
AT5G08790 315 / 4e-108
Arabidopsis NAC domain containing protein 81, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10020883
80 / 3e-18
AT5G08790 320 / 6e-110
Arabidopsis NAC domain containing protein 81, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10001809
78 / 4e-18
AT5G13180 271 / 6e-92
VND-interacting 2, NAC domain containing protein 83 (.1)
Lus10010959
79 / 1e-17
AT1G65910 389 / 9e-128
NAC domain containing protein 28 (.1)
Lus10020794
79 / 1e-17
AT5G10360 449 / 8e-153
Ribosomal protein small subunit 6b, embryo defective 3010, Ribosomal protein S6e (.1.2)
Lus10007377
79 / 2e-17
AT1G65910 431 / 7e-143
NAC domain containing protein 28 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Potri.012G024100.2 pacid=42783639 polypeptide=Potri.012G024100.2.p locus=Potri.012G024100 ID=Potri.012G024100.2.v4.1 annot-version=v4.1
ATGGCTAGCGCAGGACAAGCTTTGAACGTGCCAGTTCTCCTTGCTGGCTACAGATTTAGCCCTTACAATGATGATCTTATTGTTTATTATTTGAAGAGAA
AAATTCTGGGCCAACAACTTCCTGCTGATGTTATTACCACCACTGATGTATATGCTTCAAGCCCTGATAAACTCCCCTTAGATGATTTTAAGGGTGGGGT
GGCAAACGAGTGGTTTTTCTTTTCAAATAGAAGCAAGGATGATGACACTATTGCACTGGATGGTGGCTACTATGAAATCGATCCCGAGGGTGCTGGCCCG
ATCACATGGGAGGGAAAAGTTGTTGGTTATGTAAAGACTCTGAATTTCTATCAAGGAAGCTCACCTAATGGAACTGAAACTGAATGGATGGTAGAGGAGT
TCAGAGTGAATCCCGAGTTTCTTCCAGTTAACAACAACGACCGTAGCACTCAAGAAAAGGTAGTATAA
AA sequence
>Potri.012G024100.2 pacid=42783639 polypeptide=Potri.012G024100.2.p locus=Potri.012G024100 ID=Potri.012G024100.2.v4.1 annot-version=v4.1
MASAGQALNVPVLLAGYRFSPYNDDLIVYYLKRKILGQQLPADVITTTDVYASSPDKLPLDDFKGGVANEWFFFSNRSKDDDTIALDGGYYEIDPEGAGP
ITWEGKVVGYVKTLNFYQGSSPNGTETEWMVEEFRVNPEFLPVNNNDRSTQEKVV
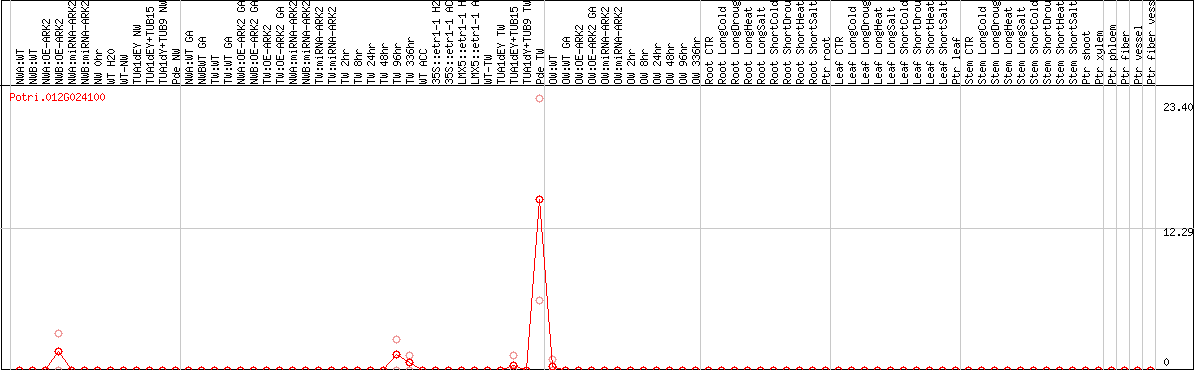
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G024100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.