External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G55805 130 / 4e-40
BolA-like family protein (.1)
AT4G26500 117 / 6e-33
SUFE1, EMB1374, CPSUFE, ATSUFE
SULFUR E 1, MBRYO DEFECTIVE 1374, ARABIDOPSIS THALIANA SULFUR E, chloroplast sulfur E (.1)
AT5G09830 47 / 4e-08
BolA-like family protein (.1)
AT5G17560 46 / 5e-07
BolA-like family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G165400
117 / 9e-33
AT4G26500 404 / 1e-140
SULFUR E 1, MBRYO DEFECTIVE 1374, ARABIDOPSIS THALIANA SULFUR E, chloroplast sulfur E (.1)
Potri.001G468300
115 / 3e-32
AT4G26500 392 / 5e-136
SULFUR E 1, MBRYO DEFECTIVE 1374, ARABIDOPSIS THALIANA SULFUR E, chloroplast sulfur E (.1)
Potri.013G074100
47 / 2e-07
AT5G17560 156 / 6e-49
BolA-like family protein (.1)
Potri.001G309200
44 / 1e-06
AT5G09830 132 / 4e-42
BolA-like family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026101
171 / 8e-57
AT4G26500 124 / 1e-37
SULFUR E 1, MBRYO DEFECTIVE 1374, ARABIDOPSIS THALIANA SULFUR E, chloroplast sulfur E (.1)
Lus10002318
166 / 1e-54
AT1G55805 119 / 2e-35
BolA-like family protein (.1)
Lus10015605
121 / 5e-34
AT4G26500 387 / 2e-133
SULFUR E 1, MBRYO DEFECTIVE 1374, ARABIDOPSIS THALIANA SULFUR E, chloroplast sulfur E (.1)
Lus10032905
114 / 2e-31
AT4G26500 380 / 4e-131
SULFUR E 1, MBRYO DEFECTIVE 1374, ARABIDOPSIS THALIANA SULFUR E, chloroplast sulfur E (.1)
Lus10020347
49 / 7e-08
AT5G17560 158 / 2e-49
BolA-like family protein (.1)
Lus10009527
49 / 7e-08
AT5G17560 145 / 2e-44
BolA-like family protein (.1)
Lus10020861
46 / 1e-07
AT5G09830 138 / 4e-44
BolA-like family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01722
BolA
BolA-like protein
Representative CDS sequence
>Potri.012G036800.1 pacid=42783612 polypeptide=Potri.012G036800.1.p locus=Potri.012G036800 ID=Potri.012G036800.1.v4.1 annot-version=v4.1
ATGGGTTCGCGAGGAGTAGCAAACATGATACTGTCAAGAGCAAGCAGGATGAAAGCAAAATTACAATCATCACTAGAAGCAAGTCTTTTGGAAATCGAAG
ACGTATCACACCAGCACGCAGGCCATGCTGCCATGAAAGGCAATACTGCAGGGGAAACTCATTTCAATGTCAAGATTGTGTCTCCCAAATTTGACGGGCT
TAATCTAGTGAAGCGTCACCGCTTGGTTTATGATGCACTCTCGGAAGAGTTGCAGTCTGGGTTGCATGCTCTCTCCATAGTTGCCAAGACTCCACAAGAA
GAAGGAGCAGCTGCAAAAAAGTGA
AA sequence
>Potri.012G036800.1 pacid=42783612 polypeptide=Potri.012G036800.1.p locus=Potri.012G036800 ID=Potri.012G036800.1.v4.1 annot-version=v4.1
MGSRGVANMILSRASRMKAKLQSSLEASLLEIEDVSHQHAGHAAMKGNTAGETHFNVKIVSPKFDGLNLVKRHRLVYDALSEELQSGLHALSIVAKTPQE
EGAAAKK
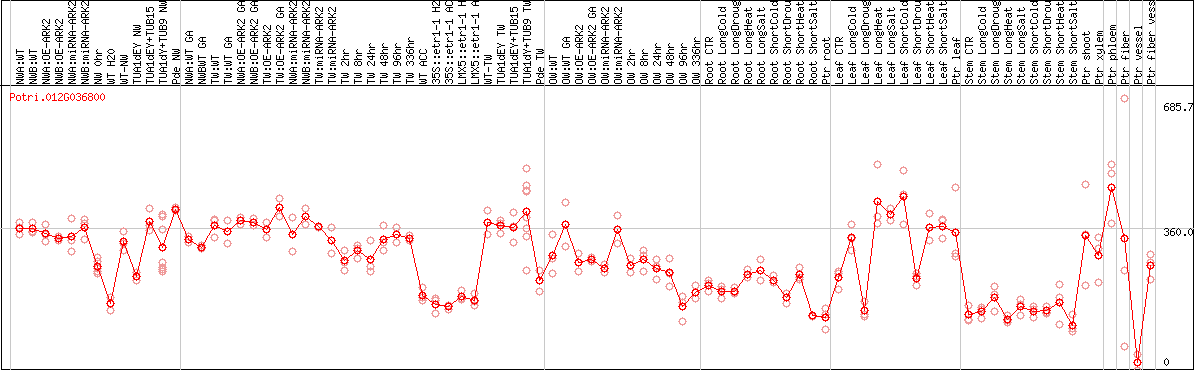
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G036800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.