External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G17660 261 / 3e-88
AGD15
ARF-GAP domain 15 (.1)
AT5G54310 231 / 3e-73
NEV, AGD5
NEVERSHED, ARF-GAP domain 5 (.1)
AT4G05330 105 / 1e-26
AGD13
ARF-GAP domain 13 (.1)
AT4G21160 102 / 1e-25
ZAC, AGD12
ARF-GAP domain 12, Calcium-dependent ARF-type GTPase activating protein family (.1.2.3.4)
AT5G13300 100 / 9e-24
AGD3, VAN3, SFC
ASCULAR NETWORK DEFECTIVE 3, SCARFACE, ARF-GAP DOMAIN3, ARF GTPase-activating protein (.1)
AT3G53710 99 / 1e-23
AGD6
ARF-GAP domain 6 (.1.2)
AT4G17890 97 / 3e-23
UBP20, AGD8
ARF-GAP domain 8 (.1.2)
AT5G61980 98 / 4e-23
AGD1
ARF-GAP domain 1 (.1)
AT2G35210 95 / 1e-22
RPA, AGD10, MEE28
MATERNAL EFFECT EMBRYO ARREST 28, root and pollen arfgap (.1.2)
AT2G37550 96 / 2e-22
ASP1, AGD7
yeast pde1 suppressor 1, ARF-GAP domain 7 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G127000
232 / 1e-73
AT5G54310 466 / 4e-161
NEVERSHED, ARF-GAP domain 5 (.1)
Potri.001G406300
231 / 8e-73
AT5G54310 400 / 6e-135
NEVERSHED, ARF-GAP domain 5 (.1)
Potri.011G044100
226 / 1e-71
AT5G54310 347 / 3e-115
NEVERSHED, ARF-GAP domain 5 (.1)
Potri.004G035800
213 / 2e-66
AT5G54310 333 / 2e-109
NEVERSHED, ARF-GAP domain 5 (.1)
Potri.011G098500
104 / 3e-26
AT4G21160 506 / 0.0
ARF-GAP domain 12, Calcium-dependent ARF-type GTPase activating protein family (.1.2.3.4)
Potri.001G372000
103 / 5e-26
AT4G21160 506 / 0.0
ARF-GAP domain 12, Calcium-dependent ARF-type GTPase activating protein family (.1.2.3.4)
Potri.001G066800
104 / 2e-25
AT5G13300 1192 / 0.0
ASCULAR NETWORK DEFECTIVE 3, SCARFACE, ARF-GAP DOMAIN3, ARF GTPase-activating protein (.1)
Potri.003G163200
101 / 3e-24
AT5G13300 1146 / 0.0
ASCULAR NETWORK DEFECTIVE 3, SCARFACE, ARF-GAP DOMAIN3, ARF GTPase-activating protein (.1)
Potri.015G105500
100 / 4e-24
AT5G61980 1110 / 0.0
ARF-GAP domain 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10027249
224 / 1e-73
AT5G54310 213 / 2e-66
NEVERSHED, ARF-GAP domain 5 (.1)
Lus10018445
222 / 2e-70
AT5G54310 265 / 2e-83
NEVERSHED, ARF-GAP domain 5 (.1)
Lus10011238
221 / 1e-69
AT5G54310 261 / 5e-82
NEVERSHED, ARF-GAP domain 5 (.1)
Lus10038582
203 / 1e-62
AT5G54310 508 / 2e-178
NEVERSHED, ARF-GAP domain 5 (.1)
Lus10037882
108 / 3e-27
AT5G54310 344 / 4e-114
NEVERSHED, ARF-GAP domain 5 (.1)
Lus10001799
100 / 8e-24
AT5G13300 1201 / 0.0
ASCULAR NETWORK DEFECTIVE 3, SCARFACE, ARF-GAP DOMAIN3, ARF GTPase-activating protein (.1)
Lus10002572
99 / 1e-23
AT5G13300 1149 / 0.0
ASCULAR NETWORK DEFECTIVE 3, SCARFACE, ARF-GAP DOMAIN3, ARF GTPase-activating protein (.1)
Lus10000903
97 / 3e-23
AT4G17890 469 / 3e-165
ARF-GAP domain 8 (.1.2)
Lus10004562
97 / 5e-23
AT4G17890 512 / 0.0
ARF-GAP domain 8 (.1.2)
Lus10024165
96 / 1e-22
AT3G07940 491 / 1e-171
Calcium-dependent ARF-type GTPase activating protein family (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01412
ArfGap
Putative GTPase activating protein for Arf
Representative CDS sequence
>Potri.012G036900.1 pacid=42784201 polypeptide=Potri.012G036900.1.p locus=Potri.012G036900 ID=Potri.012G036900.1.v4.1 annot-version=v4.1
ATGAACAGCAAAGCCTCTGTCTCTAAGGAGCTTAATGCCAGGCACACCAAGATATTAGAGGGCCTTCTTAAGCTGCAGGAGAACAGAGAATGCGCTGATT
GTCATAGCAAGGCACCGCGATGGGCAAGTGTGAACCTTGGAATATTTATATGCATGCAATGTTCAGGAACTCATCGGGGTCTTGGGGTGCATATTTCACA
GGTACGGTCTACCACACTGGATACATGGTTGCCAGAGCAGGTTGCTTTCATGCAATCCGTGGGTAACAGGAGATCCAATAGTTTCTGGGAAGCAGAACTG
CCTCCAAATGTTGACAGAAGTGGGATTGACAGATTTATCCATGCTAAGTATGGGGAGAAAAGATGGGTTTCAAGAAATTCAAAGCAACCAACTGAAGTTT
TAAGTCGAATTAACTATACTAATGACATGTTGGTAGAAGGTGCAGCTAGTAGGGTTGTTCCGAGGCAAACACGGCCACAGTCTCTTGACGAAGAGAGTTT
TACCAGGATCACAGCACAACTTTCTCCTCCAATTACAAGACCTCGTTGGGCTTCGTTGGACATGAAGAGCGATCCAATTGCTTTTCCTACACCAAAGGGA
CTAACTGAGAGCATAAAGAGGACCGATGGTCCTACAGACCTCTACAGCTTGCTTTATGTCGATGATACGCAACAAAATACCTCTTCCATGGCAACTCCTT
CGAGCTGGGCAACTTTTGATTGA
AA sequence
>Potri.012G036900.1 pacid=42784201 polypeptide=Potri.012G036900.1.p locus=Potri.012G036900 ID=Potri.012G036900.1.v4.1 annot-version=v4.1
MNSKASVSKELNARHTKILEGLLKLQENRECADCHSKAPRWASVNLGIFICMQCSGTHRGLGVHISQVRSTTLDTWLPEQVAFMQSVGNRRSNSFWEAEL
PPNVDRSGIDRFIHAKYGEKRWVSRNSKQPTEVLSRINYTNDMLVEGAASRVVPRQTRPQSLDEESFTRITAQLSPPITRPRWASLDMKSDPIAFPTPKG
LTESIKRTDGPTDLYSLLYVDDTQQNTSSMATPSSWATFD
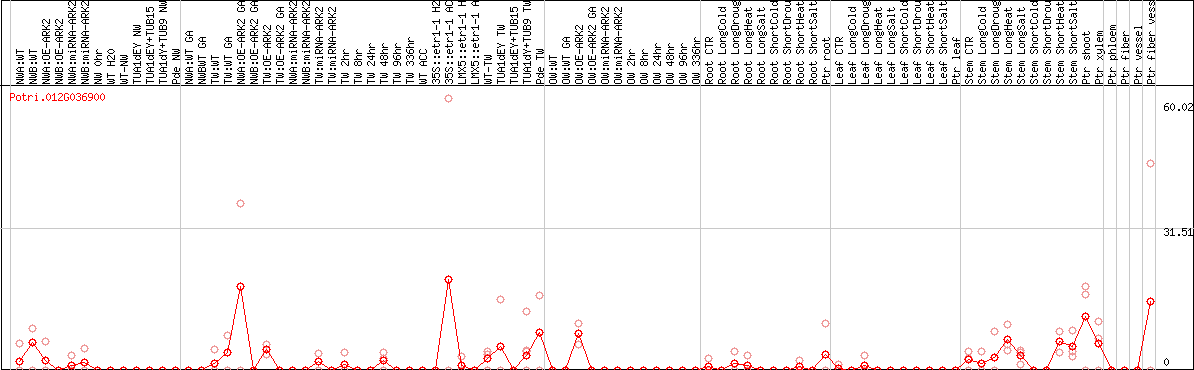
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G036900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.