Potri.012G041300 [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info | |||||||||||||||||||||
|
Representative CDS sequence |
>Potri.012G041300.1 pacid=42784012 polypeptide=Potri.012G041300.1.p locus=Potri.012G041300 ID=Potri.012G041300.1.v4.1 annot-version=v4.1
ATGGCAATGATCGAGTTTTCACAAGAATGGAAGTCAGGATTTCCAATAGACACTGTGTCTAAAGCGCCACTCTTACTCTCAAAACAAACCTCAGAATCCC
TTATTGGTCCACTTGTCTTCAACCCAATCCCTGAATCATTGGCACACCTCTTTACCTCTCCTGCTCTCTCTCCTCCTCTTCTAAACCCACCACCCCATCT
TTCTCTCACAAGATTTATCTCCACTTCAACACTCGCTGACTCTCCTCTCCCTCTCTCTACTGCTTCCTCTATTGCCTTTTCTTTTGGCCCTCAAGATCTA
CACTTTTCTTCTCCTTTGTTAGCTTATAATAGACTCCAGTTCCTTAAATGCCCACATGATGATACTGTTGTTGTGTTTTTTTCCACTGGGACTAATCTTG
ACCGAGTTGGGTTTTTGTTACTTTCTGTTAAAGATAAGAGCTTGGTGGCTACTGGGGATCAAAAGGGTGGGATTTTTACTGCTAGTAAGAGTTTGGGGTC
AAAGATTGTGAGGGTCTTAGTGAACCCCATTGAGGATGATTCTTTTTTGAATGGTAATTATTCCTTTTCTGGCTCTTTTGGTTATTTGTTAGTTTATACT
ATGTATTCTGTGAATTGGTTCTGCGTTAAATATAGTGAAAGTATGAAAAGGCCTGTTTTGAGTTATTTAGGTTGTAAGAATTTTAAGAGTTGTGGAATTG
CGAGTGCTTGTTGGAGTCCTTATATAAAAGTTCAGAGTGTGGTTTTGTTGGAGAATGGTACATTATTTTTGTTTGATTTGGAAGCCGATTGTTCTGATAT
GTATTTTAGGGGTACTAAATTGAAAGTTTCGTGGGGTGATGAAGGTAAGTTAGGAGATGGTAAATGGTTAGGGTGTGAGTTTAGTTGGCATTGTAGGGTT
TTGATTGTTGCACGGTCAGATGCTGTTTTTATGATTGATTGGAAATGCGGTGGTTTTGATGTGACTTGTTTAGCAAGGATTGATATGTTTAGTGCATATG
CTCTTAGTGAAAAGGAACGGTTCCTAGCTATGTCCAGGGCAGTTTCGGATAGTCTTCATTTTGTTTTGGTTTCTGAAACTATGTTAGTTATTTGTGATGT
GCGCAAGCCTATGATACCATTGTTGCAATGGGCACATGGTCTTGATAAGCCATGTTTTATTGATGTGTTTAGATTGTCTGATTTGAGGTCCAACTCAAGG
GATGACACACATGACTGGGCGAATTCTTCAGGTTTTGGCATTATTTTGGGGTCATTTTGGAATTGTGAATTCAGCCTCTTTTGCTATGGACCATCCTTTC
CTCCTCGTAAAGGGTCCTTCGCCTTGGAAATTTCTAAATTTTCCAGTTGTTTATACGCTTGGGACCACCCTTCAGGTCTCATGTTATCAGGTGATGATTG
TCAACGTGGAGATTGCCTTGTAAGAGAACAGTTTTGGAAGGAAGCTCTTCCTGAATGGACTGATTGGCAGCAGAAAAAAGACATAGTGTTGGGCTTTGGT
GTTTTAAGCAACGATCTCTCTTCATTGTTGTTTGAGCCTGATGAATTTGGTGGTTTTGTTTTAATTAGATTGATGTCATCAGGAAAGCTTGAATCACAGA
GATATTGTGCCTCATGGGAATTGGTTAAAAATATAGAAGTGGCTCAAAGAGATCCAATGCTGCATTCAGAAGACAACTTACTTTACTTCATGGGTGATGA
GGAATACAAGGTCCCTAGAAAATTTAAGTATTTTGAACTGAACTATCTACATGCACATTTGAATGGCAACCTTTCTCAAGTTCTCGATTCAAACATGGCA
AAGCCTTGTGAGTGTCCTCATGAGAAGGAATTGTTCAGCCTAGAATTTCATGAAGTATTGTGCAAGAAGCTGAAAATTTGTGGGTTTGGTCAATTTAGAA
CATCCCCTGCCATTACAGTTACATTTAATGACATAAACTTGCCAACAAGCATCCATGAGGTAGCTTTGAGGAGAATGTGGGCTGAATTGCCAATGGAATT
CTTGCAACTCGCCTTTTCCAGCTATTCTGAATTGCATGAAGTACTTTTGGATCAGAAAAGGGTTGCTTTGGAATTTTCAGTTGTTCCAGAACTACCTCAA
TTACCACCATTTTTCTTGCGGAAGCCTTCAAACCACAGCAATAGGTGTTTACGTAAAGTGCAAAGTAGTGATGCCCTTGTGGGTCCAGCTCTTCCTCTCC
CTATTTTGTCTACACTTCATGAGCTTCGTAATGGTTGTCCAAATTCACAGGAAGAAACAGGCGGATTTTCATCAGAATCAGAACTTAGTGTTCGATGCAA
TGAAGTTATGCAGGTGGCCAAGGAAGTGGCTGTTTCAGATTCCACAACCAAGCTTCAGGATGATAATGCTATATCCCTTGATGATGACAGAGATGACTTT
TTGGATCATTCTGAAAAACCAAAGTCTTTCCTTCTATACCATCCAACTGCATGTCAGCTCTCTTTTCAAGTTCACAAAGAGGATAATTGTGTGTATCAGA
ATGACAAGTTTGCTTCCATGATTACAAAAGTGCATGAGAAACAGTCACCTCATCCTGAAAAGGTGGAAACTTTCAAACTAGAATTCTTTGATGATCTGTG
CCCAATCGACTTGAAATTTGATGCTCGTGAGGTGAAATTTAGTTCACAAGAGTCGAAAATTTCAAATTTATTGAAGAAGAATTTTTCAAAATGGCAAGAG
GAATTCACTCCTTATCGGGAATTTTGTTCCCGATTTACATCCCCTCGTGACAAGATGGCCTAA
|
||||||||||||||||||||
|
AA sequence
|
>Potri.012G041300.1 pacid=42784012 polypeptide=Potri.012G041300.1.p locus=Potri.012G041300 ID=Potri.012G041300.1.v4.1 annot-version=v4.1
MAMIEFSQEWKSGFPIDTVSKAPLLLSKQTSESLIGPLVFNPIPESLAHLFTSPALSPPLLNPPPHLSLTRFISTSTLADSPLPLSTASSIAFSFGPQDL
HFSSPLLAYNRLQFLKCPHDDTVVVFFSTGTNLDRVGFLLLSVKDKSLVATGDQKGGIFTASKSLGSKIVRVLVNPIEDDSFLNGNYSFSGSFGYLLVYT
MYSVNWFCVKYSESMKRPVLSYLGCKNFKSCGIASACWSPYIKVQSVVLLENGTLFLFDLEADCSDMYFRGTKLKVSWGDEGKLGDGKWLGCEFSWHCRV
LIVARSDAVFMIDWKCGGFDVTCLARIDMFSAYALSEKERFLAMSRAVSDSLHFVLVSETMLVICDVRKPMIPLLQWAHGLDKPCFIDVFRLSDLRSNSR
DDTHDWANSSGFGIILGSFWNCEFSLFCYGPSFPPRKGSFALEISKFSSCLYAWDHPSGLMLSGDDCQRGDCLVREQFWKEALPEWTDWQQKKDIVLGFG
VLSNDLSSLLFEPDEFGGFVLIRLMSSGKLESQRYCASWELVKNIEVAQRDPMLHSEDNLLYFMGDEEYKVPRKFKYFELNYLHAHLNGNLSQVLDSNMA
KPCECPHEKELFSLEFHEVLCKKLKICGFGQFRTSPAITVTFNDINLPTSIHEVALRRMWAELPMEFLQLAFSSYSELHEVLLDQKRVALEFSVVPELPQ
LPPFFLRKPSNHSNRCLRKVQSSDALVGPALPLPILSTLHELRNGCPNSQEETGGFSSESELSVRCNEVMQVAKEVAVSDSTTKLQDDNAISLDDDRDDF
LDHSEKPKSFLLYHPTACQLSFQVHKEDNCVYQNDKFASMITKVHEKQSPHPEKVETFKLEFFDDLCPIDLKFDAREVKFSSQESKISNLLKKNFSKWQE
EFTPYREFCSRFTSPRDKMA
|
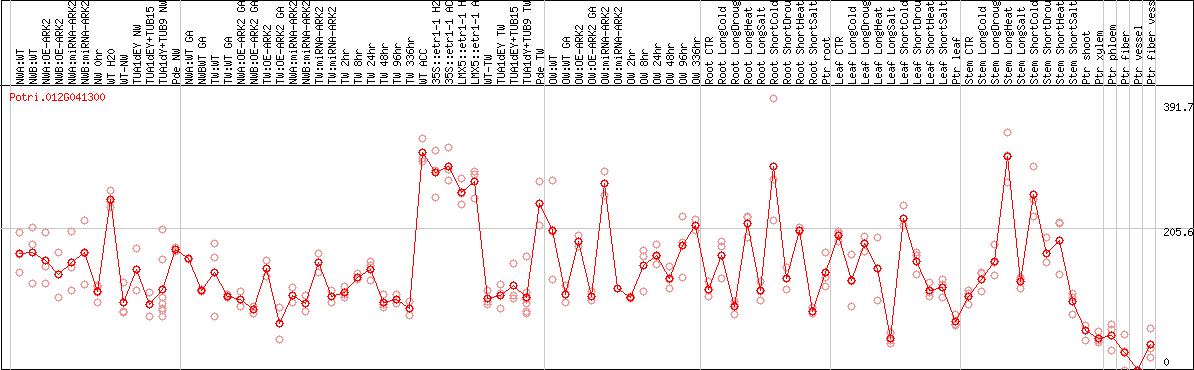
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G041300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
