External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G14760 87 / 2e-22
KNATM
KNOX Arabidopsis thaliana meinox (.1.2.3)
AT4G08150 74 / 2e-16
HD
BP1, KNAT1
BREVIPEDICELLUS 1, BREVIPEDICELLUS, KNOTTED-like from Arabidopsis thaliana (.1)
AT1G70510 67 / 6e-14
HD
ATK1, KNAT2
ARABIDOPSIS THALIANA KN 1, KNOTTED-like from Arabidopsis thaliana 2 (.1)
AT1G23380 64 / 9e-13
HD
KNAT6S, KNAT6L, KNAT6
KNOTTED1-like homeobox gene 6 (.1.2)
AT1G62990 54 / 4e-09
HD
IXR11, KNAT7
KNOTTED-like homeobox of Arabidopsis thaliana 7 (.1)
AT1G62360 48 / 5e-07
HD
WAM1, WAM, SHL, BUM1, STM
WALDMEISTER 1, WALDMEISTER, SHOOT MERISTEMLESS, SHOOTLESS, BUMBERSHOOT 1, BUMBERSHOOT, KNOX/ELK homeobox transcription factor (.1)
AT5G25220 45 / 3e-06
HD
KNAT3
KNOTTED1-like homeobox gene 3 (.1.2)
AT5G11060 45 / 5e-06
HD
KNAT4
KNOTTED1-like homeobox gene 4 (.1)
AT4G32040 44 / 9e-06
HD
KNAT5
KNOTTED1-like homeobox gene 5 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G043500
65 / 4e-13
AT1G23380 371 / 3e-129
KNOTTED1-like homeobox gene 6 (.1.2)
Potri.008G188700
64 / 1e-12
AT1G23380 390 / 1e-136
KNOTTED1-like homeobox gene 6 (.1.2)
Potri.015G079100
61 / 9e-12
AT1G23380 309 / 3e-104
KNOTTED1-like homeobox gene 6 (.1.2)
Potri.005G014200
61 / 1e-11
AT1G23380 250 / 1e-81
KNOTTED1-like homeobox gene 6 (.1.2)
Potri.012G087100
59 / 5e-11
AT1G23380 323 / 1e-109
KNOTTED1-like homeobox gene 6 (.1.2)
Potri.013G008600
58 / 1e-10
AT1G23380 250 / 1e-81
KNOTTED1-like homeobox gene 6 (.1.2)
Potri.002G113300
57 / 3e-10
AT4G08150 421 / 7e-147
BREVIPEDICELLUS 1, BREVIPEDICELLUS, KNOTTED-like from Arabidopsis thaliana (.1)
Potri.004G004700
55 / 2e-09
AT1G62360 385 / 3e-133
WALDMEISTER 1, WALDMEISTER, SHOOT MERISTEMLESS, SHOOTLESS, BUMBERSHOOT 1, BUMBERSHOOT, KNOX/ELK homeobox transcription factor (.1)
Potri.011G011101
54 / 3e-09
AT1G62360 400 / 7e-139
WALDMEISTER 1, WALDMEISTER, SHOOT MERISTEMLESS, SHOOTLESS, BUMBERSHOOT 1, BUMBERSHOOT, KNOX/ELK homeobox transcription factor (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029125
69 / 2e-14
AT1G23380 417 / 3e-147
KNOTTED1-like homeobox gene 6 (.1.2)
Lus10013037
67 / 7e-14
AT1G23380 417 / 4e-147
KNOTTED1-like homeobox gene 6 (.1.2)
Lus10042103
56 / 1e-10
AT1G70510 125 / 1e-35
ARABIDOPSIS THALIANA KN 1, KNOTTED-like from Arabidopsis thaliana 2 (.1)
Lus10001238
52 / 3e-08
AT1G23380 250 / 1e-80
KNOTTED1-like homeobox gene 6 (.1.2)
Lus10001934
50 / 1e-07
AT5G25220 435 / 9e-152
KNOTTED1-like homeobox gene 3 (.1.2)
Lus10030003
45 / 8e-06
AT1G62360 315 / 5e-105
WALDMEISTER 1, WALDMEISTER, SHOOT MERISTEMLESS, SHOOTLESS, BUMBERSHOOT 1, BUMBERSHOOT, KNOX/ELK homeobox transcription factor (.1)
Lus10006291
45 / 8e-06
AT5G25220 489 / 3e-172
KNOTTED1-like homeobox gene 3 (.1.2)
Lus10001941
43 / 2e-05
AT5G25220 439 / 2e-153
KNOTTED1-like homeobox gene 3 (.1.2)
Lus10001195
40 / 0.0001
AT1G62990 189 / 2e-60
KNOTTED-like homeobox of Arabidopsis thaliana 7 (.1)
Lus10028244
39 / 0.0006
AT4G08150 370 / 1e-127
BREVIPEDICELLUS 1, BREVIPEDICELLUS, KNOTTED-like from Arabidopsis thaliana (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03790
KNOX1
KNOX1 domain
PF03791
KNOX2
KNOX2 domain
Representative CDS sequence
>Potri.012G043400.3 pacid=42782855 polypeptide=Potri.012G043400.3.p locus=Potri.012G043400 ID=Potri.012G043400.3.v4.1 annot-version=v4.1
ATGGAGAGTAGCAGCATGAACAATACTGTTGGGAAAGATGATGATCTCTTAGTTGACACTGCAGAAGCTGAGGCTCTCAAGAAGAGAATCTCTAGCCATC
CTTTGTATGGACTTTTGGTCCAAACTCACATCGACTGTTTGAAGGTTGGTACCGTTGGTGATGTTGACCGAATCCCAAGGGTGAGGCCGAATTTATCCTG
TCAATTTCCCAATCCAAGCTCCCTCAGTCAGCCAGAACTAGACAGTTTCATGGAAGCATACTGCTTTGCCCTGAGCAAGTTGAAAGAAGCAATGGAGGAA
CCCCAGCAAGAAACAGTAGCTTTCATCAACAGCATGCACTTACAGCTTAAAGAGCTCACAAGAACCCATTCCAAGAGTACTGCTGAACATTCCACTTCCA
CTACATCTTCCGGAGAAAGAATTGACTAG
AA sequence
>Potri.012G043400.3 pacid=42782855 polypeptide=Potri.012G043400.3.p locus=Potri.012G043400 ID=Potri.012G043400.3.v4.1 annot-version=v4.1
MESSSMNNTVGKDDDLLVDTAEAEALKKRISSHPLYGLLVQTHIDCLKVGTVGDVDRIPRVRPNLSCQFPNPSSLSQPELDSFMEAYCFALSKLKEAMEE
PQQETVAFINSMHLQLKELTRTHSKSTAEHSTSTTSSGERID
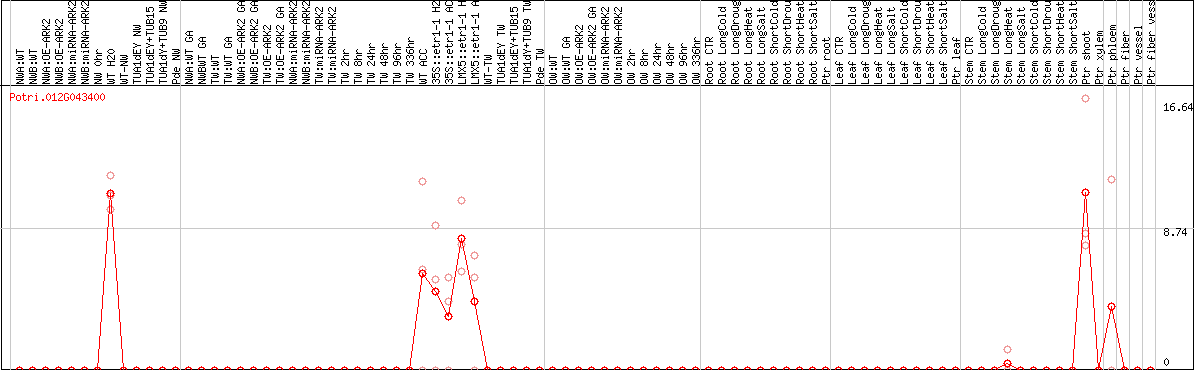
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G043400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.