External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G30700 101 / 1e-25
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G09190 99 / 8e-25
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G13410 97 / 2e-24
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G14820 97 / 3e-24
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G38010 96 / 6e-24
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT1G08070 96 / 1e-23
EMB3102, OTP82
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G42920 94 / 3e-23
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT5G40405 94 / 4e-23
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G15300 93 / 7e-23
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G08820 92 / 1e-22
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G034900
190 / 4e-59
AT4G21065 370 / 2e-122
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.011G052300
103 / 2e-26
AT2G33760 747 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.006G231200
103 / 3e-26
AT5G66520 504 / 1e-172
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.012G135600
100 / 2e-25
AT4G38010 672 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Potri.007G050200
100 / 2e-25
AT4G37380 817 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.008G121400
100 / 3e-25
AT1G13410 561 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.018G040100
100 / 3e-25
AT1G08070 974 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.012G109300
99 / 4e-25
AT5G50990 616 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G202600
98 / 2e-24
AT2G42920 660 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039974
101 / 1e-25
AT1G06150 612 / 0.0
EMBRYO DEFECTIVE 1444, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Lus10010385
99 / 1e-24
AT4G37380 758 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10024971
97 / 5e-24
AT4G37380 772 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10017446
96 / 7e-24
AT1G11290 1152 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10017243
96 / 1e-23
AT1G13410 539 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10029436
96 / 1e-23
AT3G08820 828 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10021069
94 / 4e-23
AT1G13410 533 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10021068
94 / 4e-23
AT1G13410 533 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10033026
94 / 5e-23
AT1G08070 921 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10035815
94 / 6e-23
AT4G30700 1026 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Representative CDS sequence
>Potri.012G044150.1 pacid=42782934 polypeptide=Potri.012G044150.1.p locus=Potri.012G044150 ID=Potri.012G044150.1.v4.1 annot-version=v4.1
ATGTTCAAGGATATTGAAACATCCCGCCAACAGATATTCCCAGCCTTAGTGGCATGGAATACTATGATTGGTAGTTATGTCTCTTGTGGTAAGTTCAAAG
AAGCACTTGGCATGTTCTCAAGGATGATGGAACTCGGCGTAGAGCCTGACGAGGCCACACTCGTTGAGACGTTCTCTGCAGGTTCTTCATTGGGTGCATT
AGATTTTGGGAGATGGGATCATTCATGTATTAGTAATACTGATCATGGGAGTATTATAGAGGTTAATTATTCACTCCTTAATATGTATGCAAAGTGTGGA
GCTCTTGAAGAAGCATATGAGACATTCGATGGGATGAGCAAGAAGAATACAGTAACGTGGAATACGATGATTTTGGGACTAGCAGTCATGGCTTTGCAAA
TGATGCATTGGTTCTCTTTTCCATGTTAG
AA sequence
>Potri.012G044150.1 pacid=42782934 polypeptide=Potri.012G044150.1.p locus=Potri.012G044150 ID=Potri.012G044150.1.v4.1 annot-version=v4.1
MFKDIETSRQQIFPALVAWNTMIGSYVSCGKFKEALGMFSRMMELGVEPDEATLVETFSAGSSLGALDFGRWDHSCISNTDHGSIIEVNYSLLNMYAKCG
ALEEAYETFDGMSKKNTVTWNTMILGLAVMALQMMHWFSFPC
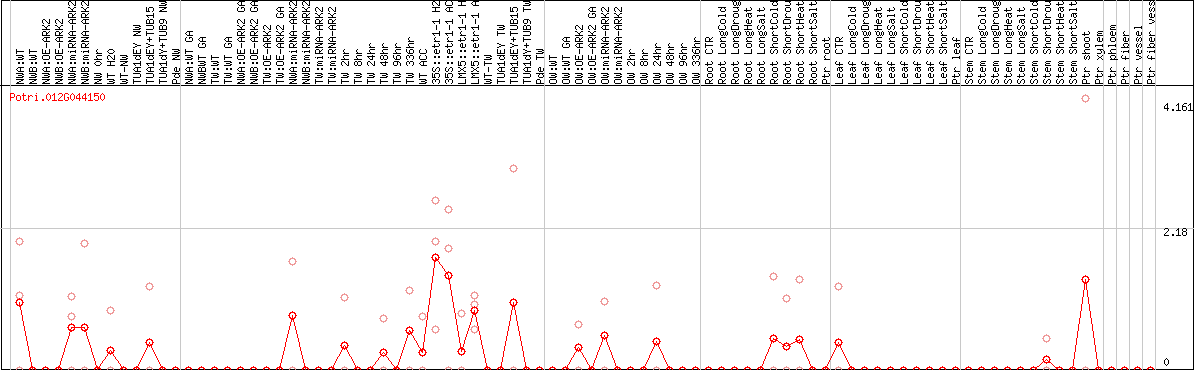
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G044150 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.