Potri.012G044700 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.012G044700.1 pacid=42783241 polypeptide=Potri.012G044750.1.p locus=Potri.012G044700 ID=Potri.012G044700.1.v4.1 annot-version=v4.1
ATGGTGTTCAAGAACAAGCTATTTTTCTCATCTTCAAAGAAATCAGAGACTTCAAGTCCTGATGGATCCAACAACAGTCCAAGATCAATCGGGTCAAATT
CACCGATCCGGTCCGAGAAGAAAAAACCCGCCAAATCTAAAGATTCAACACCTACTACTCCAGCTAGCACTGCTTCTTCTAATAATTCTACTTGCAAGCA
AACCCAAGTTAAAGATGGGGTAAAAAAGAAAGACTCATTTTTTAAAGGCAAAGAAACTGTGAGCCAACCTCGAACTCCTACCAAACCAGGGATTTCAAAT
TCTGGGTCGGATTTAAAATCCAAAAAAGGTGGTGTCTTGGTTGATAATAAGGAGAAAGAAGCAGAAAAGTATTCTGTTTCGCCGATTTTAGCTTCATCTT
TGGGGTTGAATAGGATCAAAACAAGATCAGGGCCATTGCCACAAGAGACTTTCTTTAGTTTTAAAGGTGATAAAGGGAGTGGTGTATTGGGGTCTAGTAA
TTTGTCAAGGCCTAGTGCTAGTAGTGGGGATGGGGGGTCGAGTTCGAATTCAAGTTCTTTAGGGTCAGGAAAGAAGAAGGAAGGAATATTGGGGCAGAGT
AAGTTGAGGGTGTTTCAAGAGAGTGGTAACGGAGGTGATAATTCGGATAGTATGTCTACTGGAAGTGGAGGACAGTCTAGAGAAGTTAGTCCGAATTTGC
AGGCAAGAACTAGGTTGCAAAGTGGAGAGTCATCAAGCGAAGCAGGTCAACATAATTCATCACGGGGTCACTCTGGAGGCTTACGAAGTTCTGATGCTAT
CACACCTGAGACTTATGATTGTGAAAACCCAAAGGAATCTGAATCTCCCAGGTTTCAAGCAATACTCCGTTTGACAAGTGCCCCTAGAAAGAGGTTTCCT
GCTGACATAAAAAGTTTCTCACATGAATTAAATTCCAAAGGTGTACGGCCTTTCCCATTTTGGAAGCCTCGAGGGCTAAATAACTTGGAGGAAATCTTGG
TTGTCATACGTGCAAAATTTGACAAAGCAAAGGAAGAAGTGAATTCTGACCTGGCTATCTTTGCAGCGGATCTGGTTGGAATTCTTGAAAAAAATGCGGA
TAGTCATCCTGAGTGGCAGGAAACTATTGAAGACTTGTTGGTTTTGGCTCGGAGTTGCGCTATGACATCACCTGGAGAGTTTTGGCTTCAGTGTGAAGTT
ATAGTCCAAGAACTGGATGATAGACGCCAAGAACTTCCCCCAGGAATTTTGAAGCAACTTCATACCCGAATGCTTTTCATCCTAACAAGGTGTACCAGAT
TATTGCAATTCCACAAGGAAAGGGTGCTGGATGAGAATGAAAATGTTTTCGGGCTTCGTCAATCTAGGCTTTTGCATCCTGTAGATAAACGAATTCCTTC
TTTTGTTGGTAGGGATGGGAAAGTTTCTAGTGCTGCAAAGAAGGCAGCTTCAGGAAGGAAATCATATAGTCAAGAGCATAAGGCTGCATTAGTAAGAAAA
TCATACAGTCAAGAGCAGCGTGATTGGAGCAGAGAACAAGATATACTGCCAGGAAAACTGCTCTCTCTTGCTGATAACGCTCTTAAGAGTGACGAGTCTC
CCACAGGCAGGGATCGGATATCTTCTTGGAAGCCACTCCCATCTCCTCCTGGCAAAAGCACAAAGGAAGTTGTTCCTGTAGAGGAGCAGAATGACAGTAA
GATTGAACCTTTGAAAACTTCAAATGACAGGAGAGGAGCCTCTGACGTGCATCTGGCTGCTGCAAAGGTTTCTGACCTTCCAATGGTGAAAGATGTGCAT
GAAAATTCTACGAAGCACCAACCTAAAATTTCTTGGGGAAACTGGGGCGATCAGCAGAATATTGCTGATGAGAGTTCAATAATTTGTCGCATTTGTGAGG
AGGAAGTTCCGACTTTACATGTAGAAGATCATTTAAGAATTTGTGCAATTGCAGATCGATGTGATCAGAAGGGTCTAAGTGTCAACGAGCGCTTGATCAG
AATTTCCGAAACACTTGAGAAGATGATAGTCCAGAAGGATATTCATCATGCAGTGGGAAGCCCAGACGTTGCTAAAGTATCAAATTCTAGTGTGACTGAA
GAATCTGATGTTCTCTCTCCAAAACTCAGTGATTGGTCTCATAGAGGGTCAGAGGACATGCTTGACTGCTTCCCTGAAGCTGACAATGCTGTTTTTATGG
ATGACCTGAAAGGGTTGCCTTCCATGTCATGTAAAACCCGATTTGGTCCCAAGTCTGATCAAGGAATGGCCACATCATCAGCTGGTAGCATGACTCCTAG
ATCTCCATTATTGACACCAAAGACCAGTCATATAGACTTGCTTTTAGCAGGGAAGAGTGCTTTTTCCGAGCATGATGATCTTCCTCAGCTGAATGAACTT
GCCGATATTGCAAGATGTGTAGCAACTACACCTCTAGAGGATGATCGCTCCACTCCCTATTTGCTAACTTGTCTTGGAGACTTGAGGGTTGTCATTGAGC
GGAGAAAGTTTGATGCGCTTACTGTGGAGACTTTTGGAACAAGAATTGAGAAGCTTATTCGGGAGAAATATTTGCAGCTTTGCGAGCTAGTGGAAGATGA
GAAGGTTGATATAGCAAGTACTGTAATTCATGAAGATACACCTTTGGAAGATGATGTTGTTCGAAGCTTGAGAACAAGTCCAATACATTCTTCTAAGGAT
CGTACCTCCATAGATGACTTCGAGATAATAAAACCAATAAGTCGTGGGGCATTTGGAAGGGTGTTTTTAGCTAAAAAGAGAGCAACAGGAGACCTTTTTG
CCATAAAGGTTCTTAAAAAGGCTGATATGATCCGCAAGAATGCTGTTGAAAGTATACTAGCAGAACGAGATATTTTAATTTCAGTCCGCAATCCCTTTGT
GGTTCGCTTCTTCTATTCATTTACTTGTCGTGAGAATTTGTATCTTGTGATGGAGTATTTGAATGGTGGCGATCTGTATTCATTGTTGAGAAATTTGGGC
TGCTTAGATGAAGATGTTGCTCGTGTATACATTGCAGAAGTTGTGCTCGCTTTGGAATATTTGCACTCACTGCGTGTAGTTCATCGTGATTTGAAGCCAG
ATAATTTGTTGATTGCACATGATGGACATATCAAGCTGACAGACTTTGGTCTTTCCAAGGTTGGTCTCATCAATAGCACCGATGATTTATCTGGTCCAGC
AGTTAGCGGGACTTCCATGCTTGTGGATGATGAACCTCAACTATCTACTTCCGAGCATCAGCGAGAAAGGCGGAAAAAACGTTCTGCTGTGGGCACACCT
GATTATCTTGCACCAGAGATTCTTCTGGGAACAGGACATGGAACAACTGCTGATTGGTGGTCTGTGGGTGTTATTTTATTTGAATTGATTGTTGGTATTC
CACCCTTTAATGCAGAGCATCCCCAGACGATATTTGATAATATTCTCAACTGTAAGATACCTTGGCCACGGGTGCCTGAGGAAATGAGTCCTGAAGCACA
GGATCTAATTGATAGATTATTAACTGAAGATCCTTACCAGAGACTTGGAGCTGGAGGAGCATCAGAGGTAAAGCAGCATGTATTCTTTAAAGATATCAAC
TGGGACACACTTGCCAGGCAAAAGGCTGCATTTGTGCCCAGTTCAGAGAGTGCACTTGATACAAGCTACTTCACTAGTCGATACTCTTGGAATACTTCAG
ATGATGCAATATACCCAGCCAGTGACTTTGAGGATTCGAGTGATGCAGATAGCTTGAGTGGCAGCAGTAGCTGCTTAAGCAATCGCCATGATGAAGTGGG
AGATGAATGTCAAGGTCTTGCTGAATTTGAGTCTGGTTCAGGTGTCAATTACTCCTTCAGTAATTTTTCATTCAAGAATCTCTCCCAGCTTGCATCAATC
AACTATGATATTCTCTCTAAGGGTTGGAAGGATGATCCACCAACAACTAATTTATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.012G044700.1 pacid=42783241 polypeptide=Potri.012G044750.1.p locus=Potri.012G044700 ID=Potri.012G044700.1.v4.1 annot-version=v4.1
MVFKNKLFFSSSKKSETSSPDGSNNSPRSIGSNSPIRSEKKKPAKSKDSTPTTPASTASSNNSTCKQTQVKDGVKKKDSFFKGKETVSQPRTPTKPGISN
SGSDLKSKKGGVLVDNKEKEAEKYSVSPILASSLGLNRIKTRSGPLPQETFFSFKGDKGSGVLGSSNLSRPSASSGDGGSSSNSSSLGSGKKKEGILGQS
KLRVFQESGNGGDNSDSMSTGSGGQSREVSPNLQARTRLQSGESSSEAGQHNSSRGHSGGLRSSDAITPETYDCENPKESESPRFQAILRLTSAPRKRFP
ADIKSFSHELNSKGVRPFPFWKPRGLNNLEEILVVIRAKFDKAKEEVNSDLAIFAADLVGILEKNADSHPEWQETIEDLLVLARSCAMTSPGEFWLQCEV
IVQELDDRRQELPPGILKQLHTRMLFILTRCTRLLQFHKERVLDENENVFGLRQSRLLHPVDKRIPSFVGRDGKVSSAAKKAASGRKSYSQEHKAALVRK
SYSQEQRDWSREQDILPGKLLSLADNALKSDESPTGRDRISSWKPLPSPPGKSTKEVVPVEEQNDSKIEPLKTSNDRRGASDVHLAAAKVSDLPMVKDVH
ENSTKHQPKISWGNWGDQQNIADESSIICRICEEEVPTLHVEDHLRICAIADRCDQKGLSVNERLIRISETLEKMIVQKDIHHAVGSPDVAKVSNSSVTE
ESDVLSPKLSDWSHRGSEDMLDCFPEADNAVFMDDLKGLPSMSCKTRFGPKSDQGMATSSAGSMTPRSPLLTPKTSHIDLLLAGKSAFSEHDDLPQLNEL
ADIARCVATTPLEDDRSTPYLLTCLGDLRVVIERRKFDALTVETFGTRIEKLIREKYLQLCELVEDEKVDIASTVIHEDTPLEDDVVRSLRTSPIHSSKD
RTSIDDFEIIKPISRGAFGRVFLAKKRATGDLFAIKVLKKADMIRKNAVESILAERDILISVRNPFVVRFFYSFTCRENLYLVMEYLNGGDLYSLLRNLG
CLDEDVARVYIAEVVLALEYLHSLRVVHRDLKPDNLLIAHDGHIKLTDFGLSKVGLINSTDDLSGPAVSGTSMLVDDEPQLSTSEHQRERRKKRSAVGTP
DYLAPEILLGTGHGTTADWWSVGVILFELIVGIPPFNAEHPQTIFDNILNCKIPWPRVPEEMSPEAQDLIDRLLTEDPYQRLGAGGASEVKQHVFFKDIN
WDTLARQKAAFVPSSESALDTSYFTSRYSWNTSDDAIYPASDFEDSSDADSLSGSSSCLSNRHDEVGDECQGLAEFESGSGVNYSFSNFSFKNLSQLASI
NYDILSKGWKDDPPTTNL
|
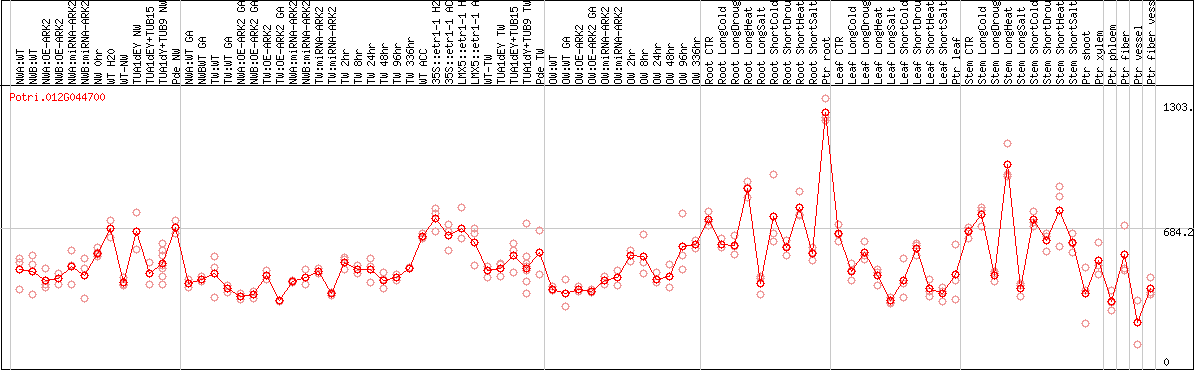
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G044700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
