Potri.012G067900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.012G067900.12 pacid=42784383 polypeptide=Potri.012G067900.12.p locus=Potri.012G067900 ID=Potri.012G067900.12.v4.1 annot-version=v4.1
ATGGCGCTGTTCAGACGTTTCTTTTACCGGAAGCCGCCGGATCGGCTTCTTGAGATCTCCGAGAGAGTCTACGTATTTGACTGTTGCTTCTCCACGGAAG
TGTTGGAAGAAGATGAGTACAAGGTTTACTTGGGTGGCATTGTAGCACAGCTACAAGACCACTTTCCTGATGCGTCCTTCATGGTTTTTAACTTTAGAGA
AGGGGAGAGGCGTAGCCAAATTTCAGACATACTTTCACAATATGACATGACAGTTATGGATTACCCTCGCCAATATGAGGGCTGTCCTATGCTGCCTCTG
GAGATGATCCACCACTTCCTTCGATCAAGTGAAAGCTGGCTGTCCTTGGAAGGGCAGCAGAATGTGCTGTTAATGCATTGTGAAAGAGGAGGTTGGCCTG
TACTTGCTTTCATGCTTGCAGGTCTTTTGCTATACCGGAAACAATATACTGGGGAGCACAAGACTCTTGAAATGGTCTACAAGCAAGCTCCGAGGGAACT
TCTCCATCTTTTGTCTCCTTTAAATCCACAGCCTTCCCAACTTAGATATCTTCAGTACATTTCTAGAAGAAATTTTGGTTCTGATTGGCCTCCATCAGAT
ACTCCTTTACAGTTGGATTGTCTGATGCTAAGATCCCTTCCTTTGTTTGAAGGGGGTAAAGGTTGCCGGCCAGTTGTACGTGTTTATGGCCAGGATCCTT
CAAAACCAGCCAATAGAACTTCCAAGCTTCTGTTTTCAACATCGAAGACTAAAAAACATGTTCGCCTCTACCGGCAGGAAGAGTGTATGCTAGTGAAAAT
AGATATACGGTGCCGGGTTCAAGGGGATGTTGTTCTTGAGTGCATCCATTTGGATGAAGATTTGGTACGTGAGGAGATGATGTTCAGAGTGATGTTTCAC
ACAGCATTTGTGCAGGCCAATATTTTGATGCTCGTCCGTGATGAGATTGACTTTCTCTGGGACGCCAAGGACCAATTTCCAAAGGACTTTAGAGCAGAGG
TACTTTTTGTGGATGCTGATGCTGTGGTGCCTAATGTCACCACAGTCGAGGCAAATGAAGATGGGAACGAGACAGAAAGTGCTTCACCTGAGGAATTTTT
TGAGGTGGAAGAAATTTTCAGCAATGTAGTAGACGGGCATGAAGCAAAGGGGTATGGTGCCTCTCATAAAGTCCATGACAACATGCCAGTTGATGTAGAC
GGTAAAGAAGTCTGGAAGGAGGACTCAGATCTTCACTCATTTGAAGACTGTGCTTCTGATGACGGGAATCACAAACAGGAGGGGAAGCTAGATTCTAGTG
TTGATGCAGTGAAGGACATTGCTGTGGATGATGTCAAGTACAAGGTAGATGAGAAGGTGGATTCTGATTTTTTAGCAGTGAAAGACATTACCGTGGATGA
TGGCGAAATCAAGGCAGATTCTGTGGTTTCTGCTACTGGTACGCTGATCAGGAAACAAACAACAGAAGTGATTGGAGATGTCGACGGGGAACTTAAAAAG
ATGGAAGATGAAGGTGATAGAGAAAATAGTGCTACAAAGAAGTTGGAATCCCAGGATCCACCAGTAGAGTTGAGTGCTGATGCTGGTAGACAAAAACTTG
AGCAATTAATGTTACCTTCACCAAGGAGGCAGCCTACATCAAATGCAAAACCTGCTGCAGATTCAATTATAACCGAACAAAAGACTAAACATAATGAACA
AGAAGGAGCTCATGGAAAACAAACAAAGCCAAATACCATTCCTCGGTGGGTTCCCCCTAATAGAGGCCCTTTTAGCAATTCCATGCACGTAGCACATCCG
CCATCAAGATATAATAGTGCACCACCGGCTCTTACATTTTGTGCTTCACCGGAGGACTCTAGTGCTGGTGGCCATGTAAAGATTTCTTCTGTTGCCACTG
GCCCTGGAGATATAATTTCAAATGATTTTCCTTCTCCAACTGAAGCTCCACCTTCTCTTGACCCGCAACAAATTGCCCTCCGTGGTCCCCCACCTCCACC
TCTGCCATATTCCAACAAATCTTCATTTTATGATTTCCAGGCCAGTTCAGGTGGTGAAGCACCACCACTTCATTCACAGATTGCTGATGCAGTTTCCTTT
CCCCCTCCGCCTCCCACATCATTTAGCAGGCAGAATATTCAGATGATTCCTCAACATTCATCTCCACCACCCCCTCCTCCTCTTCCCCAGCTGTCTAATA
GGCAGACTATTGGGATGGTTTTGCCTCCACCTCCTCCACCCCCTTGGAAGTCTGGGAATACTCCAGCTGTTTTCACTACAACTTATTCTCCACCCCCTCC
TCCTCCATCCCCTCTTCTCCCTTCAGGTGCCTCTACTACAAATCATGGCAGACTTGGAATTCCCAATCCACCTCCACCTCCACCTCCACCTCTTTCTCTT
GCACACACGTGTAGCACTCCTTTGGCTCAATCAATGCCCACTCATGGAGTTATACCTCCACCTCCTCCTCCACCCTCGAAGCCTGCACAAAGAGCTCCAC
CACCTTCGCAGCCTGCACATGGAGCTCCACCACCCCCACCTCCACCTCCTATGCGTGGTCCACCACTTCCTCCACTTGTGAGTCAGGCTCCTCCTCCACC
TCCTATGCGTGGTCCACCACCACCTCCACCTCCACCTCCACCTCCTATGCGTGGTCAACCACTACCTCCACTTGTGAGTCAGGCTCCTCCTCCACCACCG
CCTCCTCCAGGCCGTGGAGCCCCCCCTCCTCCCCCGCCTCCTCCAGGCCGTGGAGCCCCCCCTCCCCCGCCTCCTCCAGGCCGTGGACCACCTCCACCGC
CTCCACCAGGAGCTCGTGTTCCTGGACCTCCTGCATCACCAAGACCTCCAGGCAGTGCACCCCATCCACCTCCAGCCTTAGGTGTCAAAGGAGCTGCTGA
TGCAAGAGGTCTGCCTTCGGGAAGAGGGCGGGGTTTTTTACGTCCTTCAGGGATGGGGACATCTGCTACAGCACCCAGAAGATCTTCTTTAAAGCCTTTG
CATTGGAGCAAGGTAACAAGAGCAATACAAGGAAGCTTATGGGAGGAGCTGCAAAGGCATGGAGAGTCCCAAATTGCCCCAGAATTTGACGTGTCAGAGC
TAGAAAGCCTTTTCTCTGCAACTGTCCACAAACCTGCTGATTCGGGAGGTAAAGCTGGTGGAAGACACAAGTCTGTTGGATCAAAAACTGACAAAGTTCA
CCTGATTGATCTGCGGAGGGCGAATAACACTGAAATTATGCTTACAAAAGTTAAGATGCCGCTTTCTGACATGATGGCTGCAGTTCTAGCAATGGATGAG
TCAATATTAGATGTTGATCAAGTGGAAAATCTTATAAAGTTCTGTCCCACGAAAGAGGAGATGGAACTTCTCAAGGGCTATACAGGTGACAAGGAAAACC
TGGGGAAGTGTGAACAGTACTTCTTGGAGCTGATGAAAGTGCCACGTGTGGAGTCCAAGTTGAGAGTGTTTTCTTTCAAGATTCAATTCGGCTCTCAGAT
TTCAGAATTTAAAAAGAGCTTAAACACTGTAAACTCTGCATGTGATGAGGTCCGGAATTCCCTCAAATTGAAGGAAATTATGAAGAAAATTCTTTATTTG
GGGAATGCGCTGAATCAAGGAACTGCACGGGGTTCTGCAATTGGATTCAAGTTGGACAGTCTTTTAAAACTCACTGATACGCGTGCTTCTAACAACAAGA
TGACGCTCATGCATTATCTTTGCAAGGTGCTCGCTGCCAAGTCACAAGCACTTCTAGATTTTCACCGCGACCTAGTTAGCCTCGAAACTGCATCTAAGAT
ACAATTGAAGTCTTTAGCAGAAGAAATGCAGGCTATAATCAAGGGGTTAGAGAAGGTTAAGAAGGAGTTGGCAGCATCAGAAAATGATGGTCCTGTGTCC
GAAGTTTTTCGAAAGACACTAAAAGAATTCATTAGTGTTGCTGAGACAGAGGTGGCATCCGTGACAAGTTTCTATGCTGTGGTGGGTAGAAATGCGGATG
CACTTGCACTGTACTTCGGTGAGGATCCTGCCCGTTGCCCATTTGAGCAAGTTACAGCTACTCTCTTAAATTTTGTGAGATTGTTTCGTAAAGCGCATGA
GGAAAATTTGAAACAGGCTGAATTGGAGAGGAAGAAAGCTGAGAAGGAGGCTGAAATGGAGAAGGCAAGGGGTATAAATCTTACAAAGAAGAACATGGAA
TAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.012G067900.12 pacid=42784383 polypeptide=Potri.012G067900.12.p locus=Potri.012G067900 ID=Potri.012G067900.12.v4.1 annot-version=v4.1
MALFRRFFYRKPPDRLLEISERVYVFDCCFSTEVLEEDEYKVYLGGIVAQLQDHFPDASFMVFNFREGERRSQISDILSQYDMTVMDYPRQYEGCPMLPL
EMIHHFLRSSESWLSLEGQQNVLLMHCERGGWPVLAFMLAGLLLYRKQYTGEHKTLEMVYKQAPRELLHLLSPLNPQPSQLRYLQYISRRNFGSDWPPSD
TPLQLDCLMLRSLPLFEGGKGCRPVVRVYGQDPSKPANRTSKLLFSTSKTKKHVRLYRQEECMLVKIDIRCRVQGDVVLECIHLDEDLVREEMMFRVMFH
TAFVQANILMLVRDEIDFLWDAKDQFPKDFRAEVLFVDADAVVPNVTTVEANEDGNETESASPEEFFEVEEIFSNVVDGHEAKGYGASHKVHDNMPVDVD
GKEVWKEDSDLHSFEDCASDDGNHKQEGKLDSSVDAVKDIAVDDVKYKVDEKVDSDFLAVKDITVDDGEIKADSVVSATGTLIRKQTTEVIGDVDGELKK
MEDEGDRENSATKKLESQDPPVELSADAGRQKLEQLMLPSPRRQPTSNAKPAADSIITEQKTKHNEQEGAHGKQTKPNTIPRWVPPNRGPFSNSMHVAHP
PSRYNSAPPALTFCASPEDSSAGGHVKISSVATGPGDIISNDFPSPTEAPPSLDPQQIALRGPPPPPLPYSNKSSFYDFQASSGGEAPPLHSQIADAVSF
PPPPPTSFSRQNIQMIPQHSSPPPPPPLPQLSNRQTIGMVLPPPPPPPWKSGNTPAVFTTTYSPPPPPPSPLLPSGASTTNHGRLGIPNPPPPPPPPLSL
AHTCSTPLAQSMPTHGVIPPPPPPPSKPAQRAPPPSQPAHGAPPPPPPPPMRGPPLPPLVSQAPPPPPMRGPPPPPPPPPPPMRGQPLPPLVSQAPPPPP
PPPGRGAPPPPPPPPGRGAPPPPPPPGRGPPPPPPPGARVPGPPASPRPPGSAPHPPPALGVKGAADARGLPSGRGRGFLRPSGMGTSATAPRRSSLKPL
HWSKVTRAIQGSLWEELQRHGESQIAPEFDVSELESLFSATVHKPADSGGKAGGRHKSVGSKTDKVHLIDLRRANNTEIMLTKVKMPLSDMMAAVLAMDE
SILDVDQVENLIKFCPTKEEMELLKGYTGDKENLGKCEQYFLELMKVPRVESKLRVFSFKIQFGSQISEFKKSLNTVNSACDEVRNSLKLKEIMKKILYL
GNALNQGTARGSAIGFKLDSLLKLTDTRASNNKMTLMHYLCKVLAAKSQALLDFHRDLVSLETASKIQLKSLAEEMQAIIKGLEKVKKELAASENDGPVS
EVFRKTLKEFISVAETEVASVTSFYAVVGRNADALALYFGEDPARCPFEQVTATLLNFVRLFRKAHEENLKQAELERKKAEKEAEMEKARGINLTKKNME
|
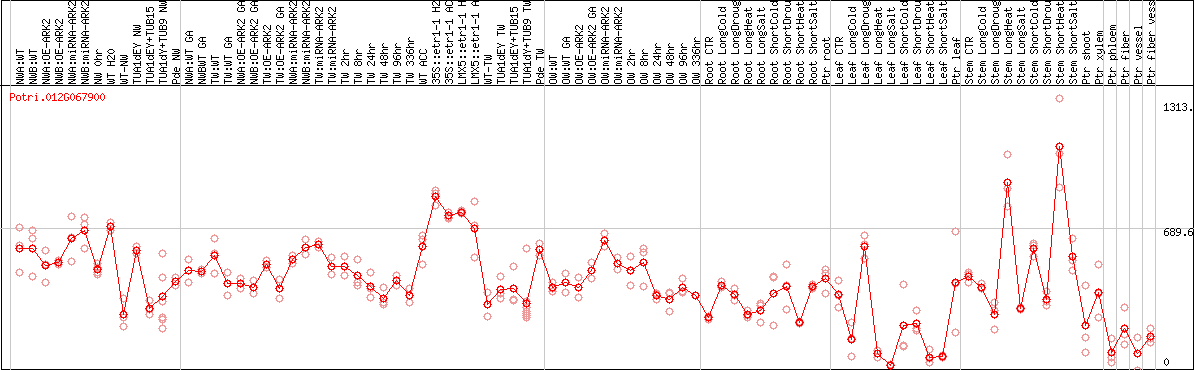
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G067900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
