External link
Symbol
Pt-LBD27.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G47870 180 / 5e-55
AS2
ASL29, SCP, LBD27
SIDECAR POLLEN, ASYMMETRIC LEAVES 2-like 29, LOB domain-containing protein 27 (.1)
AT3G13850 120 / 9e-33
AS2
LBD22
LOB domain-containing protein 22 (.1)
AT5G63090 100 / 1e-25
AS2
LOBB, LOB
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
AT3G27650 97 / 5e-25
AS2
LBD25
LOB domain-containing protein 25 (.1)
AT5G66870 100 / 9e-25
AS2
LBD36, ASL1
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
AT2G30130 98 / 1e-24
AS2
PCK1, LBD12, ASL5
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
AT1G65620 97 / 2e-24
AS2
AS2
ASYMMETRIC LEAVES 2, Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
AT1G72980 96 / 9e-24
AS2
LBD7
LOB domain-containing protein 7 (.1)
AT3G50510 95 / 1e-23
AS2
LBD28
LOB domain-containing protein 28 (.1)
AT1G31320 94 / 1e-23
AS2
LBD4
LOB domain-containing protein 4 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008629
296 / 2e-100
AT3G47870 175 / 2e-52
SIDECAR POLLEN, ASYMMETRIC LEAVES 2-like 29, LOB domain-containing protein 27 (.1)
Lus10036665
287 / 3e-97
AT3G47870 182 / 2e-55
SIDECAR POLLEN, ASYMMETRIC LEAVES 2-like 29, LOB domain-containing protein 27 (.1)
Lus10033111
283 / 2e-95
AT3G47870 194 / 5e-60
SIDECAR POLLEN, ASYMMETRIC LEAVES 2-like 29, LOB domain-containing protein 27 (.1)
Lus10042187
140 / 8e-41
AT3G47870 47 / 4e-06
SIDECAR POLLEN, ASYMMETRIC LEAVES 2-like 29, LOB domain-containing protein 27 (.1)
Lus10003789
124 / 4e-34
AT3G13850 162 / 7e-49
LOB domain-containing protein 22 (.1)
Lus10002513
121 / 4e-33
AT3G13850 161 / 1e-48
LOB domain-containing protein 22 (.1)
Lus10013510
108 / 4e-28
AT3G13850 143 / 1e-41
LOB domain-containing protein 22 (.1)
Lus10025840
102 / 8e-26
AT5G66870 184 / 4e-57
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
Lus10018993
100 / 1e-25
AT3G26660 134 / 2e-41
LOB domain-containing protein 24 (.1)
Lus10031285
99 / 3e-25
AT2G30130 226 / 3e-76
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03195
LOB
Lateral organ boundaries (LOB) domain
Representative CDS sequence
>Potri.012G072000.2 pacid=42783035 polypeptide=Potri.012G072000.2.p locus=Potri.012G072000 ID=Potri.012G072000.2.v4.1 annot-version=v4.1
ATGACTCTCAAGGGAGGCACCAGCCAAGCTTGTGCTTCGTGCAAGTATCAAAGGAGAAAGTGCTCCTCTGAGTGTCCTCTAGCACCTTACTTCCCTTCTG
AACAACCAAAGATGTTCCAAAATGCTCATAAGTTGTTTGGTGTACGCAAGATTCTCAGGATACTTGAGAATCTTGATTATCTCCAGAAGGAGGAAGCCAT
GCGTTCTATTATTTACCAATCCAATATTCGTGACAGGTTTCCTGTTCATGGCTGTTTGGGCATCATCTATCAACTGCATTTCCAGATTCGGCAGGCGGAA
GAAGAGCTCCGTGCTGTCTACGCACACCTTGAGATGTACAGGCAGCATCACCACCATCAGATTGCAGCTTTGACAGAGGATGTTCCCTCGCAATTAGAAT
TAGGAATGGCACTTCCTCAGCCTTGCTTGGCACAACAGTACTCCTACTCTATTAGCAGTAATATTGGTTACAGCTCTGGTTACTTGGATTCTAATGATAA
TGTTGGGAATCAGCTTTGGAGTCAACATCCTTATGCTACTAACAGTAACACCAATTACACGCCAATTATTCAGTCTCAGCTAGATGCTTCTCAGCCACTA
TCTACCCAACAAGAAATTGTTCAAGATTATGATGAGATGCATTCATTTTTTGATACTATTGACGATAGGCAATCATATATAGAAACTAAAGAACCCTATG
AGTCCAGCTCAGAAGAATCGTTGAAGGACACTACACAATCCATGGAGCATGTTGCAGAGAATGAATTGAAGAGTGCTGCTGCATGTTTCAGTCTTACCAG
TGTCAACTGA
AA sequence
>Potri.012G072000.2 pacid=42783035 polypeptide=Potri.012G072000.2.p locus=Potri.012G072000 ID=Potri.012G072000.2.v4.1 annot-version=v4.1
MTLKGGTSQACASCKYQRRKCSSECPLAPYFPSEQPKMFQNAHKLFGVRKILRILENLDYLQKEEAMRSIIYQSNIRDRFPVHGCLGIIYQLHFQIRQAE
EELRAVYAHLEMYRQHHHHQIAALTEDVPSQLELGMALPQPCLAQQYSYSISSNIGYSSGYLDSNDNVGNQLWSQHPYATNSNTNYTPIIQSQLDASQPL
STQQEIVQDYDEMHSFFDTIDDRQSYIETKEPYESSSEESLKDTTQSMEHVAENELKSAAACFSLTSVN
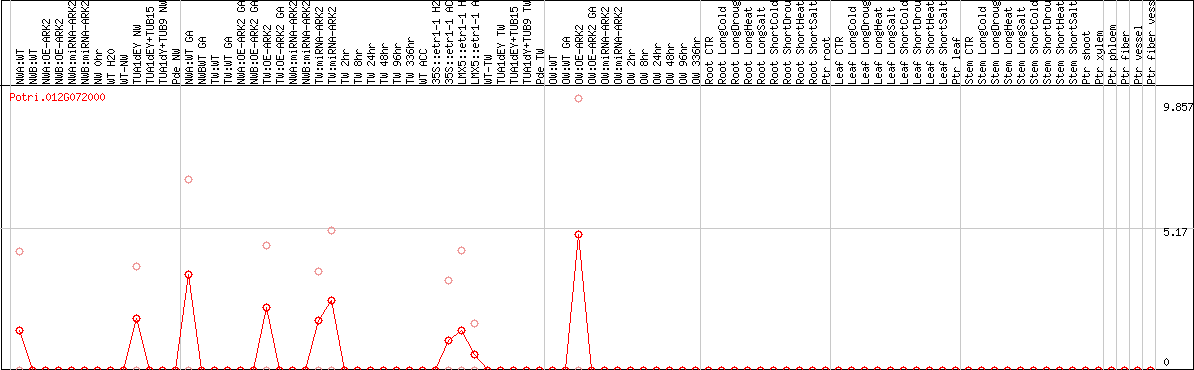
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G072000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.