External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G62610 228 / 4e-74
bHLH
bHLH079
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G59640 193 / 1e-60
bHLH
ZCW32, bHLH031, BPE, BPEUB, BPEP
BIG PETAL UB, BIG PETAL P (.1.2)
AT1G26260 129 / 4e-35
bHLH
CIB5, bHLH076
cryptochrome-interacting basic-helix-loop-helix 5 (.1.2.3)
AT3G07340 130 / 5e-35
bHLH
bHLH062
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G48560 130 / 1e-34
bHLH
bHLH078
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G23690 125 / 2e-33
bHLH
bHLH077
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G68920 125 / 6e-33
bHLH
bHLH049, ACE1
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
AT4G34530 119 / 1e-31
bHLH
bHLH063, CIB1
cryptochrome-interacting basic-helix-loop-helix 1 (.1)
AT4G36540 119 / 1e-31
bHLH
bHLH058, BEE2
BR enhanced expression 2 (.1.2)
AT5G50915 118 / 1e-31
bHLH
bHLH137
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G068100
406 / 1e-144
AT5G62610 263 / 7e-88
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.010G040000
206 / 2e-66
AT1G59640 273 / 1e-92
BIG PETAL UB, BIG PETAL P (.1.2)
Potri.008G190800
204 / 5e-65
AT1G59640 230 / 2e-75
BIG PETAL UB, BIG PETAL P (.1.2)
Potri.002G248500
134 / 1e-35
AT3G07340 349 / 5e-115
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.014G148900
131 / 1e-34
AT3G07340 254 / 2e-78
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.002G235400
127 / 4e-33
AT3G07340 257 / 2e-79
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.010G136100
126 / 5e-33
AT1G68920 387 / 3e-129
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Potri.008G113200
126 / 7e-33
AT1G68920 388 / 1e-129
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Potri.006G057200
122 / 1e-32
AT3G57800 280 / 2e-92
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10033120
302 / 1e-103
AT5G62610 265 / 9e-89
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10036659
290 / 2e-98
AT5G62610 256 / 3e-85
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10013284
222 / 3e-72
AT1G59640 303 / 3e-104
BIG PETAL UB, BIG PETAL P (.1.2)
Lus10030807
220 / 3e-71
AT1G59640 300 / 6e-103
BIG PETAL UB, BIG PETAL P (.1.2)
Lus10029110
139 / 9e-38
AT1G68920 320 / 5e-104
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Lus10013052
138 / 3e-37
AT1G68920 316 / 2e-102
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Lus10003172
127 / 1e-34
AT5G48560 205 / 4e-62
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10021968
125 / 4e-34
AT3G23690 163 / 5e-48
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10041261
124 / 7e-34
AT3G23690 172 / 3e-51
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10030581
130 / 8e-34
AT1G68930 910 / 0.0
pentatricopeptide (PPR) repeat-containing protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00010
HLH
Helix-loop-helix DNA-binding domain
Representative CDS sequence
>Potri.012G072700.1 pacid=42784105 polypeptide=Potri.012G072700.1.p locus=Potri.012G072700 ID=Potri.012G072700.1.v4.1 annot-version=v4.1
ATGGATCCTCCATTGATAAACGAGAAGTCCTTCTCTGCGGCCAACCCATCTTCCTACTCATTAACTGAGATTTGGCCTTTTCCTCCTCCTTCTTCAACTG
CTTTGGGCCTCCGGATGGCTAATTTGGCCGACCGAGATGGGTCCGTTGACGAATCCACTGTGACCGAGCAGAGGGGTGGAAATAGAAATGGAAATAGGAA
GGCTAGAGATTTGAGTTCCGAGGAGGATGACTCATCCATAATGGTTTCTACCACCACTAGTGCCCATGATTTGAATGATTTGAATGGTAAGCGGAGGAAA
ATTTCGGGGTCAAGGAATGAAAACAATGATTCAAGAGCTGAAATAGAAGCAAGTTCTGCTGCTAATAACAAGCCAGCTGAACCAAGCAGTAAACCATCTG
AGCCGCCTATGCAAGACTATATTCATGTCAGATCAAGAAGGGGCCAAGCGACTGATAGCCACAGTTTAGCAGAGAGAGCTCGGAGAGAGAGGATTGGTGA
GAGGATGAAGATACTCCAAGATTTGGTCCCTGGGTGTAATAAGGTCATTGGTAAAGCACTTGCCCTTGATGAGATAATTAATTACATCCAGTCACTCCAG
TGTCAGGTTGAGTTCCTCTCAATGAAGCTAGAAGCAGTAAATTCAAGGATGAGCACGAGCCCCGCGATTGAAGGCCTTCATCCAAAAGACCTTGGTGCAC
AGCCATTTGATGCTACTGGAATGATATTTGGCCCACAACCAACAAGAGACTATGTTCAAGGATCCCAACCGGAATGGCTACATATGCAGGTTGGTGGAAG
TTTTAAAAGAGCCACGTAA
AA sequence
>Potri.012G072700.1 pacid=42784105 polypeptide=Potri.012G072700.1.p locus=Potri.012G072700 ID=Potri.012G072700.1.v4.1 annot-version=v4.1
MDPPLINEKSFSAANPSSYSLTEIWPFPPPSSTALGLRMANLADRDGSVDESTVTEQRGGNRNGNRKARDLSSEEDDSSIMVSTTTSAHDLNDLNGKRRK
ISGSRNENNDSRAEIEASSAANNKPAEPSSKPSEPPMQDYIHVRSRRGQATDSHSLAERARRERIGERMKILQDLVPGCNKVIGKALALDEIINYIQSLQ
CQVEFLSMKLEAVNSRMSTSPAIEGLHPKDLGAQPFDATGMIFGPQPTRDYVQGSQPEWLHMQVGGSFKRAT
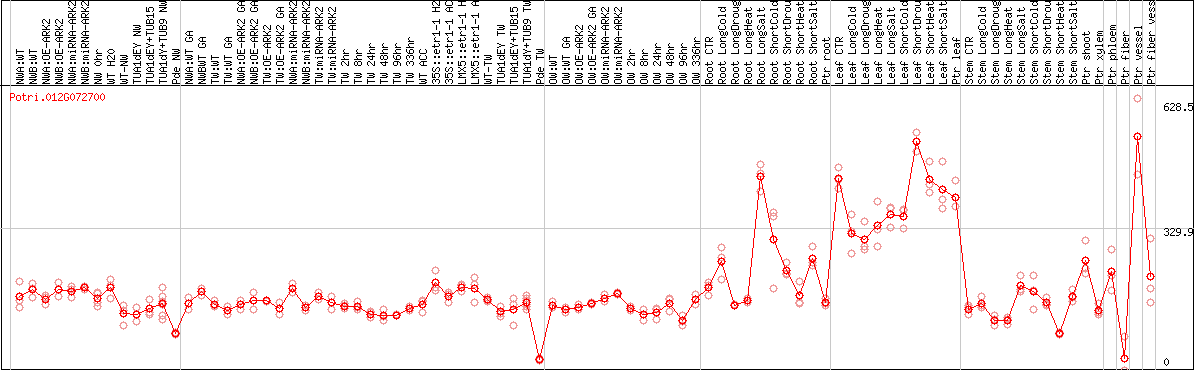
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G072700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.