Potri.012G074800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.012G074800.2 pacid=42784314 polypeptide=Potri.012G074800.2.p locus=Potri.012G074800 ID=Potri.012G074800.2.v4.1 annot-version=v4.1
ATGTTGACCAAGTTTGAGACCAAGAGTAATAGAGTGAAGGGGCTGAGTTTCCACAGTAAGAGACCATGGATCCTTGCTAGTCTCCACAGTGGTGTGATCC
AGCTATGGGACTATCGTATGGGCACTCTCATCGATCGCTTTGATGAGCATGATGGCCCTGTTCGTGGTGTTCATTTCCATAAATCTCAACCTCTTTTTGT
CTCTGGAGGTGATGATTACAAGATTAAAGTCTGGAACTACAAATTGCATAGGTGTCTGTTTACACTTCTTGGACATCTTGATTACATTCGTACCGTGCAG
TTTCATCATGAATACCCATGGATCGTGAGTGCCAGTGATGATCAGACCATCCGCATTTGGAACTGGCAGTCTCGTACTTGCATTTCTGTGTTGACAGGAC
ACAATCATTATGTGATGTGTGCCTCATTCCACCCTAAAGAAGACCTTGTTGTTTCTGCATCTCTGGATCAGACTGTTCGTGTTTGGGATATTGGTGCTTT
GAGAAAAAAGACGGTGTCCCCTGCTGATGACATCATGAGGCTTACACAGATGAACACTGATCTTTTTGGTGGTGTTGATGCTGTTGTTAAGTATGTTTTG
GAGGGCCATGACAGAGGAGTCAACTGGGCTGCATTCCACCCAACACTACCTTTGATTGTCTCAGGAGCAGATGATCGCCAAGTAAAGTTGTGGCGCATGA
ATGACACAAAGGCCTGGGAAGTGGACACGTTAAGAGGACACATGAACAATGTGTCATGTGTTATGTTCCATGCCAAGCAGGATATAATTGTATCAAATTC
AGAGGATAAAAGCATTCGTGTCTGGGATGTAACTAAACGAACTGGTGTTCAAACTTTCCGTCGAGAACATGACCGATTCTGGATCCTTGCATCTCATCCT
GAGATGAATCTTCTGGCTGCAGGTCATGACAGTGGCATGATTGTCTTTAAGTTAGAGAGAGAAAGGCCTGCTTTTGCTTTAAGTGGTGATTCTTTGTTCT
ATACTAAAGATCGCTTTTTACGGTTTTTTGAGTTTTCAACTCAAAGAGATACACAAGTAATTCCCATCCGACGACCTGGCACCACAAGCCTGAATCAAAG
TCCAAGAACTCTTTCGTACAGTCCTACAGAAAATGCTGTTCTTATCTGCTCGGATGTGGATGGAGGATCTTATGAGTTATATGTCATTCCTAAAGACAGC
ATTGCTAGGGGTGATGCTGTGCCAGAGGCAAAGAGAGGTGCTGGTGGATCTGCTGTCTTTGTGGCTCGGAATAGGTTTGCTGTGCTTGATAAGAGCAGCA
ATCAAGTCCTAGTTAAGAACCTCAAGAATGAAGTTGTCAAAAAGAGTGGTCTTCCTATCAGTTGTGATGCTATATTCTATGCTGGGACAGGTAACTTGCT
TTGCAGGGCAGAGGATAGGGTGGTCATATTTGATCTCCAGCAGAGGCTTGTTCTCGGGGAGTTGCAGACACCTTTTGTCAAGTATGTTGTTTGGTCCAAT
GACATGGAAAGTGTTGCCTTGCTAAGCAAACATGCCATCATCATTGCTAGCAAGAAGCTTGTCCACCAATGCACCCTCCATGAAACAATCCGTGTAAAGA
GTGGAGCTTGGGATGATAATGGTGTTTTCATTTACACAACTTTAAATCATATCAAATACTGCCTGCCTAATGGAGATAGTGGTATAATTAGAACCCTTGA
TGTCCCAATATATATCACTAAGATTTCTGGAAATACTATCTTTTGCTTAGATCGGGATGGGAAGAATAAGCCTATTGTTATTGATGCTACAGAATACATC
TTTAAACTGTCTCTGCTGAAGAAGAGATATGATCATGTTATGAGCATGATAAGGAACTCACAGCTTTGTGGTCAGGCAATGATTGCTTATTTGCAACAGA
AGGGGTTCCCAGAAGTCGCTCTGCATTTTGTGAAAGATGAGAGAACTAGGTTTAATTTGGCTTTAGAGAGTGGAAATATTCAGATTGCTGTGGCTTCAGC
AAAGGAAATTGATGAGAAAGACCACTGGTATAGGTTGGGGGTTGAAGCTCTTCGCCAAGGGAATGCAGGTATAGTTGAATATGCCTACCAGAGGACAAAA
AATTTTGAGAGGTTATCTTTCCTTTATCTCATAACTGGTAATTTAGAAAAGCTGTCCAAGATGCTTAGAATTGCTGAAGTTAAGAATGATGTCATGGGTC
AGTTTCACAATGCCTTGTACCTGGGTGATGTCAGAGAACGTGTTAAGATCTTGGAGAATGCTGGCCATCTGCCACTTGCTTATGCCGCAGCTAAGGTCCA
TGGGCTGGAGGATGTTGTGGAACGTCTAGCAGCTGAGTTGGGAGATGATATTCCATCTTTTCCCAAGGGTAAAGAACCATCCCTTCTGATGCCCCCAGCC
CCTATTATGTGTGGTGGTGATTGGCCTCTTCTGAGAGTCATGAAAGGTATATTTGAAGGTGGGCTGGATAATATGGTCAGAGGTGGTGCTGATGAGGATG
AAGAAGAAGCTGCTGATGGTGATTGGGGCGAGGAACTGGACATGGTCGACGCGGTTGGTTTACAAAATGGGGATGTCACCGCAATTTTGGAGGATGGGGA
AGCAGCTGAAGAAAATGAAGAGGAGGAGGGAGGATGGGACCTTGAAGATCTGGAGCTACCTCCCGAGGCAGACACGCCAAGGGCTTCTGTGAGTGCCCGC
TCATCAGTTTTTGTTGCTCCAACTCCTGGTATGCCTGTAAGCCAGATTTGGATTCAGAGATCTTCACTTGCTGCTGAACACGCCGCGGCTGGCAATTTCG
ACACTGCCATGCGACTACTCAACAGGCAACTTGGAATTAAAAACTTTGTGCCTTTAAAACCCATGTTTCTTGATCTTCACTCAGGCAGCCATACCTATTT
ACGTGCATTTTCATCTACCCCAGTGATTTCACTGGCTGTTGAACGGGGATGGAACAAGTCTGCTAGCCCCAATGTTAGGGCCCCTCCAGCTCTGGTTTTC
GACTTCTCTCAATTGGAAGAGAAGCTTAAGGCTGGTTACAAAGCCACGACAGCTGGGAAATTTACCGAGGCACTTAAACTCTTTCTTAGCATTCTTCACA
CAATTCCTCTGATAGTTGTTGACTCGAGGAGGGAAGTTGATGAAGTCAAGGAATTGATTATTATAGTCAAAGAGTATGTTTTGGGTCTGCAAATGGAGCT
CAAGAGGAGGGAGATGAAAGACAATCCAGTACGCCAACAAGAGCTTGCTGCCTACTTCACACACTGCAACCTGCAAGCTCCTCACTTGAGGCTTGCGCTT
CAAAATGCAATGACTGTCTGCTTCAAGAATAAGAACCTTGCTACGGCAGCTAACTTTGCCAGACGGCTATTGGAGACAAACCCCCCGAATGAGAACCAGG
CAAGGTCTGCCAGGCAAGTATTGGCAGCTTCAGAGAGGAATATGACAGATGCTGCTCAGCTGAACTATGATTTTAGAAATCCTTTTGTGGTATGTGGTGC
AACATATGTTCCAATTTATCGAGGACAAAAGGATGTCTCTTGCCCGTATTGTGGTTCCCGATTTGTACCAAGCCATGAAGGGCAGCTGTGTACTGTTTGT
GATCTTGCAGTAGTTGGGGCAGATGCTTCAGGGTTGCTCTGTTCTCCTTCCCAGATACGATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.012G074800.2 pacid=42784314 polypeptide=Potri.012G074800.2.p locus=Potri.012G074800 ID=Potri.012G074800.2.v4.1 annot-version=v4.1
MLTKFETKSNRVKGLSFHSKRPWILASLHSGVIQLWDYRMGTLIDRFDEHDGPVRGVHFHKSQPLFVSGGDDYKIKVWNYKLHRCLFTLLGHLDYIRTVQ
FHHEYPWIVSASDDQTIRIWNWQSRTCISVLTGHNHYVMCASFHPKEDLVVSASLDQTVRVWDIGALRKKTVSPADDIMRLTQMNTDLFGGVDAVVKYVL
EGHDRGVNWAAFHPTLPLIVSGADDRQVKLWRMNDTKAWEVDTLRGHMNNVSCVMFHAKQDIIVSNSEDKSIRVWDVTKRTGVQTFRREHDRFWILASHP
EMNLLAAGHDSGMIVFKLERERPAFALSGDSLFYTKDRFLRFFEFSTQRDTQVIPIRRPGTTSLNQSPRTLSYSPTENAVLICSDVDGGSYELYVIPKDS
IARGDAVPEAKRGAGGSAVFVARNRFAVLDKSSNQVLVKNLKNEVVKKSGLPISCDAIFYAGTGNLLCRAEDRVVIFDLQQRLVLGELQTPFVKYVVWSN
DMESVALLSKHAIIIASKKLVHQCTLHETIRVKSGAWDDNGVFIYTTLNHIKYCLPNGDSGIIRTLDVPIYITKISGNTIFCLDRDGKNKPIVIDATEYI
FKLSLLKKRYDHVMSMIRNSQLCGQAMIAYLQQKGFPEVALHFVKDERTRFNLALESGNIQIAVASAKEIDEKDHWYRLGVEALRQGNAGIVEYAYQRTK
NFERLSFLYLITGNLEKLSKMLRIAEVKNDVMGQFHNALYLGDVRERVKILENAGHLPLAYAAAKVHGLEDVVERLAAELGDDIPSFPKGKEPSLLMPPA
PIMCGGDWPLLRVMKGIFEGGLDNMVRGGADEDEEEAADGDWGEELDMVDAVGLQNGDVTAILEDGEAAEENEEEEGGWDLEDLELPPEADTPRASVSAR
SSVFVAPTPGMPVSQIWIQRSSLAAEHAAAGNFDTAMRLLNRQLGIKNFVPLKPMFLDLHSGSHTYLRAFSSTPVISLAVERGWNKSASPNVRAPPALVF
DFSQLEEKLKAGYKATTAGKFTEALKLFLSILHTIPLIVVDSRREVDEVKELIIIVKEYVLGLQMELKRREMKDNPVRQQELAAYFTHCNLQAPHLRLAL
QNAMTVCFKNKNLATAANFARRLLETNPPNENQARSARQVLAASERNMTDAAQLNYDFRNPFVVCGATYVPIYRGQKDVSCPYCGSRFVPSHEGQLCTVC
DLAVVGADASGLLCSPSQIR
|
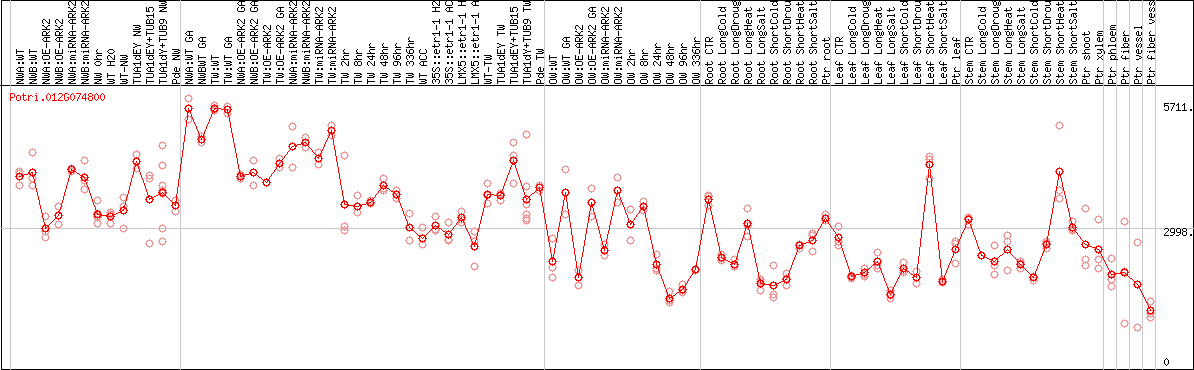
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G074800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
