Potri.012G084400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.012G084400.1 pacid=42783710 polypeptide=Potri.012G084400.1.p locus=Potri.012G084400 ID=Potri.012G084400.1.v4.1 annot-version=v4.1
ATGGCCTCAACTTCGAAAAAGACGACGCCTTCTCCTCCTTGTTGTGACTCTACTACTCCCGCTCCTCCTCCCATTCGCCGTGACGCCACCTCCTCCTCCT
CTTTGGTTCCGCAAGATTGGACGGTTCCCTCCGCTGATCGCCGCCCCGTTGTCGATTCTCTTCCAGAACCTTCCAAAACAGCCTCCGGTACTAAAACTGT
TATCCCAGTCATGTTAAGGGCTCAGACCAGCCATCCTTTGGATCCTTTGTCTGCTGCCGAAATCTCTGTTGCAGTGGCAACTGTTAGGGCTGCTGGAGCC
ACTCCTGAGCTTAGAGATAGCATGCGTTTCGTTGAGGTGGTTCTGTTTGAACCGGATAAGCATGTTGTTGCACTGGCAGATGCATATTTCTTTCCTCCTT
TCCAACCATCATTGCTTCCCAGATCAAAAGGTGGACCTATAATTCCTACAAAGTTGCCTCCAAGAAGGGCTAGACTTGTTGTTTACAACAAACGGTCAAA
TGAGACAAGCTTATGGATTGTGGAGCTCTCTGAAGTACATGCAGCGACTCGAGGTGGACATCATAGGGGAAAAGTGATTTCATCTCAAGTTGTTCCAGAT
GTTCAGCCTCCAATGGATGCTGTTGAATATGCTGAATGTGAAGCTGTTGTCAAAGACTTTCCTCCATTTAGAGAGGCAATGAAAAAAAGAGGTATTGAAG
ACATGGACCTTCTGATGGTTGATGCCTGGTGTGTTGGTTACCACAGTGATGCTGATGCTCCAAGCAGAAGACTTGCCAAACCACTTATATTTTGTAGAAC
TGAGAGTGACTGTCCAATGGAAAATGGTTATGCACGTCCAGTTGAGGGCATCCATGTACTTGTTGATATGCAAAACATGAGGGTCATTGAATTTGAAGAC
CGTAAACTTGTTCCCTTACCTCCAGCTGATCCATTGAGGAACTATACTCCTGGTGAAACAAGAGGTGGTGTTGATCGAAGTGATGTGAAACCTTTACAAA
TTATCCAGCCTGAAGGTCCAAGCTTTCGTGCTAATGGACACTATGTAGAATGGCAGAAGTGGAATTTTCGCATTGGATTCACTCCTAGGGAGGGCTTGGT
TATACATTCAGTTGCCTATGTTGATGGTAGTAGAGGGAGACGACCTATAGCCCACAGGTTAAGCTTTGTAGAGATGGTAGTTCCTTATGGAGATCCAAAT
GAGCCACATTACCGAAAGAATGCATTTGATGCAGGGGAAGATGGCCTTGGTAAAAATGCACATTCCCTAAAAAAGGGCTGTGATTGTTTGGGCTTTATTA
AATACTTTGATGCACACTTTACAAACTTCACTGGAGGTATTGAAACAATTGAAAATTGTGTATGTTTGCATGAAGAGGATCATGGAATTCTATGGAAGCA
TCAGGATTGGAGGACAGGTTTAGCAGAAGTGCGAAGGTCTAGGCGGCTGACGGTGTCCTTTATTTGCACTGTTGCTAATTATGAATATGGATTTTTCTGG
CACTTCTATCAGGATGGAAAAATCGAAGCTGAAGTTAAGCTTACGGGAATCCTTAGCTTAGGAGCATTACAACCTGGGGAGACTCGAAAATATGGTACTA
CCATTGCCCCAGGTCTTTATGCACCAGTTCATCAGCACTTTTTTGTTGCTCGTATGGACATGGCTGTTGATTGCAGACCAGGTGAAGCTTTCAATCAGGT
TGTGGAGGTAAATGTTGAAGTTGAGAAACCTGGGGAGAAAAATGTCCACAATAATGCATTTTATGCCAAAGAGACGCTGCTCCGATCAGAATTGGAAGCA
ATGCGTGCTTGCAATCCACAAACAGCTCGCCATTGGATTGTTAGAAACACTAGAACAGTAAACAGAACTGGACAATTAACAGGCTACAAGCTAGTACCTG
GTTCCAATTGTTTGCCATTGGCTGGTTCAGAGGCTAAGTTTTTGAGGCGAGCTGCATTTTTAAATCACAATCTTTGGGTTACACCTTATACACATGGTGA
GATGTTTCCTGGAGGAGAGTTCCCTAATCAAAATCCACGTGTTGATGAAGGACTGGCTACATGGGTTAAGCAGAATCGACCTTTAGAAGAAACTGATATA
GTTCTTTGGTATGTGTTTGGGATCACACATGTTCCTCGGTTGGAAGATTGGCCGGTCATGCCAGTGGAGCGCTTAGGCTTTATGCTTATGCCTCATGGGT
TTTTCAACTGCTCTCCTGCTGTGGATGTTCCTCCCAGCACATGCGAATTGGATGCCAAAGACAATGATGTTAAAGACAATGGTGTAACGAAGCCACTTCA
AAATGGGGTGCTAGCCAAGCTCTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.012G084400.1 pacid=42783710 polypeptide=Potri.012G084400.1.p locus=Potri.012G084400 ID=Potri.012G084400.1.v4.1 annot-version=v4.1
MASTSKKTTPSPPCCDSTTPAPPPIRRDATSSSSLVPQDWTVPSADRRPVVDSLPEPSKTASGTKTVIPVMLRAQTSHPLDPLSAAEISVAVATVRAAGA
TPELRDSMRFVEVVLFEPDKHVVALADAYFFPPFQPSLLPRSKGGPIIPTKLPPRRARLVVYNKRSNETSLWIVELSEVHAATRGGHHRGKVISSQVVPD
VQPPMDAVEYAECEAVVKDFPPFREAMKKRGIEDMDLLMVDAWCVGYHSDADAPSRRLAKPLIFCRTESDCPMENGYARPVEGIHVLVDMQNMRVIEFED
RKLVPLPPADPLRNYTPGETRGGVDRSDVKPLQIIQPEGPSFRANGHYVEWQKWNFRIGFTPREGLVIHSVAYVDGSRGRRPIAHRLSFVEMVVPYGDPN
EPHYRKNAFDAGEDGLGKNAHSLKKGCDCLGFIKYFDAHFTNFTGGIETIENCVCLHEEDHGILWKHQDWRTGLAEVRRSRRLTVSFICTVANYEYGFFW
HFYQDGKIEAEVKLTGILSLGALQPGETRKYGTTIAPGLYAPVHQHFFVARMDMAVDCRPGEAFNQVVEVNVEVEKPGEKNVHNNAFYAKETLLRSELEA
MRACNPQTARHWIVRNTRTVNRTGQLTGYKLVPGSNCLPLAGSEAKFLRRAAFLNHNLWVTPYTHGEMFPGGEFPNQNPRVDEGLATWVKQNRPLEETDI
VLWYVFGITHVPRLEDWPVMPVERLGFMLMPHGFFNCSPAVDVPPSTCELDAKDNDVKDNGVTKPLQNGVLAKL
|
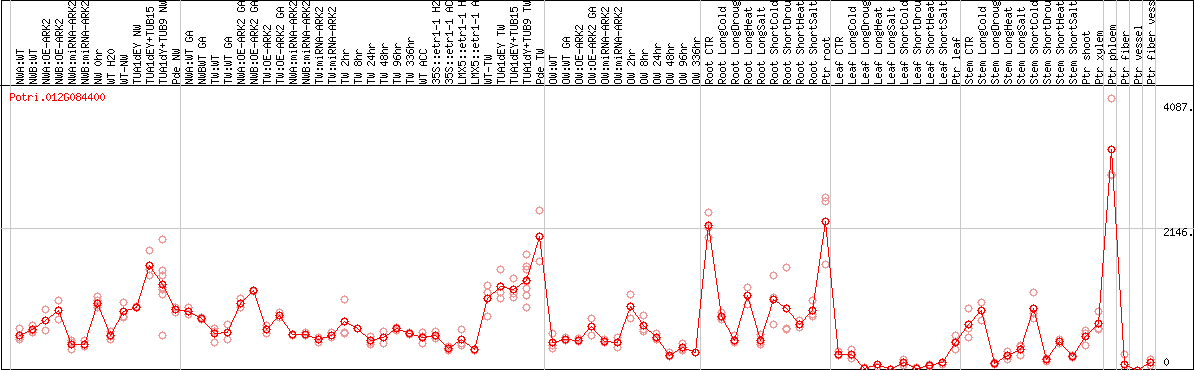
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G084400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
