Potri.012G088100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.012G088100.1 pacid=42783588 polypeptide=Potri.012G088100.1.p locus=Potri.012G088100 ID=Potri.012G088100.1.v4.1 annot-version=v4.1
ATGCCTGTAAATCCATGGCCCCTCTTTTCCTTCTTATTCTTCTCTTCTACCCTTGTGCTTCTCTTTCCTTTTACTGCTTTCGCCGTCAACCAACAAGGCG
AAACTCTTCTTTCATGGAAGAGAAGCTTGAATGGATCACCTGAGGGGTTGAATAATTGGGACTCGAGCAATGAGACTCCCTGTGGATGGTTCGGAATTAC
TTGCAACTTTAACAATGAAGTTGTGGCATTGGGTTTGAGGTATGTCAATTTATTTGGAACACTTCCCTCCAATTTTACTTTTTTGTCTTCCTTAAACAAG
CTTGTTCTGTCTGGAACCAATCTTACTGGCACAATCCCAAAAGAGATTGGCACTGCTCTTCCTCAATTGACTCACTTAGACTTGAGTGAGAATGCCTTGA
CCGGCGAGATTCCAAGTGAGCTTTGTAACTTTCCCAAGCTTGAACAACTCCTTCTCAACTCAAATCAACTTGAGGGCTCCATTCCAATCGAGATCGGAAA
CCTCACTAGCTTGAAATGGCTAATCCTCTACGACAATCAACTTAGCGGTTCCATTCCCAACACCGTAGGCAAGTTGAAGTATCTTGAAGTGATTAGAGCT
GGTGGCAACAAGAACCTTGAAGGTTCATTGCCTAAAGAAATTGGCAACTGCAGCAATTTGTTGATGTTAGGCCTAGCAGAAACAAGCATCTCAGGTTTTC
TCCCACCGAGCCTCGGCCTCCTGAAAAAACTCCAAACCGTTGCCATCTACACCACCCTCCTTTCCGGCCAAATCCCTCCAGAACTCGGTGACTGCACCGA
GCTCCAAGACATATACCTTTATGAAAACTCACTCACTGGCTCAATTCCGAAAACATTAGGCAAGCTCCGAAACCTTAGAAACCTTTTGCTCTGGCAGAAC
AACTTGGTTGGTATTATCCCTCCCGAGCTCGGAAACTGCAACCAAATGTTGGTGATTGACATATCAATGAATTCCTTAACCGGAAGCATTCCGCAATCAT
TCGGCAATCTAACTGAACTCCAAGAACTCCAGCTTAGCCTCAATCAAATCTCCGGAGAAATCCCAGCACAGCTTGGAAACTGCCAGAAGATCATTCACAT
CGAACTCGACAACAATCAAATCACTGGCTCAATCCCACCTGAAATAGGTAACTTATTTAATCTCACACTCTTTTACTTATGGCAGAACAAACTCGAAGGC
AACATACCGCCTTCCATTTCCAACTGTCAAAACTTAGAAGCCATAGATTTATCTCAAAATGGCTTAGTGGGGCCTATTCCAAAAGGAGTTTTTCAACTGA
AGAAACTCAACAAACTCTTGCTTCTTTCCAACAATCTCTCCGGTGAAATACCGCCCGAGATTGGAAACTGCTCGTCTTTGATTCGTTTTCGAGCTAATAA
CAACAAGGTTTCCGGAACTATTCCGGCCCACATTGGAAATTTGAAGAACCTGAACTTCTTGGATCTCGGATCGAACCGGATTACCGGAGTAATCCCGGAA
GAAATATCGGGTTGTCAAAATCTCACATTTTTGGATTTGCATTCAAATGCTATTTCTGGTAACTTGCCACAGAGTTTTGACAAGCTCATATCACTCCAGT
TTATTGATTTCTCCAATAACTTGATTGAAGGCACACTGAGTCCAAGTCTTGGTTCACTGAGTTCACTCACCAAACTCACTCTTGCTAAGAACAGGCTGTC
AGGTTCCATTCCGAGTCAACTCGGTTCCTGCTCCAAGTTACAGTTATTGGACCTGAGTGGAAATCAGTTATCAGGGAACATTCCGTCAAGTGTTGGAAAG
ATCCCGTCACTGGAAATCGCTTTGAATCTAAGCTTGAATCAACTTAATGGGGAGATTCCGTCAGAGTTCACGGGGTTAAACAAGCTTGGCATTTTGGATA
TTTCGTATAATCATCTTACAGGCGATCTTCAACATCTCGCAGCTTTGCAGAATCTTGTCGTGCTTAATGTCTCGCACAATAATTTCTCGGGGCACGTGCC
TGACACGCCCTTCTTCTCGAAGCTTCCTCTCAGTGTACTTGCCGGGAATCCTGCTCTTTGCTTCTCCGGTAACCAGTGCGATTCTGGTGACAAGCACGTT
CAGCGAGGGACGGCAGCGCGTGTGGCAATGATTGTGCTGTTGTGTGCCGCATGTGCGCTCCTTTTGGCAGCGCTCTACATTATCCTAGCAAGCAAAAAAC
GTGGCAGTGGGGCCCAGGAGTGTGAAGGTGAGGATGACGTGGAGATGAGTCCGCCTTGGGAAGTGACTTTATATCAGAAACTAGACTTGTCAATTGCTGA
TGTGACACGATCGTTGACAGCTGGCAATGTGGTAGGACGAGGCCGGTCCGGTGTGGTTTACAAGGTTACCATTCCGTCTGGATTAATGGTAGCTGTTAAG
AGATTCAAGTCGGCAGAGAAGATTTCAGCTGCAGCATTTTCATCCGAGATTGCTACATTGGCGAGAATTCGGCACCGAAACATTGTCCGTTTACTCGGGT
GGGGAGCAAACCGGAAAACAAAGTTGCTGTTTTATGATTACATGGCTAATGGCACATTAGGGACATTGTTACATGAAGGGAATAATTTTGGATTGGTGGA
GTGGGAGACGAGGTTTAAGATCGCATTAGGGGTGGCAGAGGGGCTAGCGTATTTGCACCACGATTGCGTACCGCCGATTCTGCACCGAGATGTGAAAGCG
CATAATATCTTGCTTGGGGACCGGTTTGAGGCATATTTAGCCGATTTCGGGCTGGCTAGGCTGGTTGAAGACGAGCATGGATCATTCTCTGCAAACCCAC
AATTTGCAGGGTCCTATGGATACATTGCACCAGAGTACGCTTGCATGCTTAAAATCACTGAAAAGAGTGATGTATATAGCTATGGAGTGGTCCTTTTAGA
GACAATAACTGGAAAAAAGCCAGTCGATCCATCATTCCCGGATGGCCAACATGTTGTCCAATGGGTCAGAAACCACCTGAGGAGCAAGAAGGACCCAGTT
GAAATACTTGATCCTAAACTGCAAGGCCACCCAGACACACAAATACAGGAAATGCTTCAAGCTCTTGGAATCTCTCTTCTTTGCACAAGCAATCGCGCTG
AAGATCGCCCGACAATGAAAGATGTGGCAGTATTATTGAAGGAAATTCGACAGGAACTGATCACAGGCGGTGAGGCCCAGAAGCCGACCAACAAATCATC
GAAAACGATGGAAAGCAATCCATCATATTCATCATCCTCGGTGACCCCAGCTCAATTGTTGATGCTTCAACAAGGGTCTTCTCGTTGCTCACTTGCTTAC
TCATCTTCATCATCAACTAGTATTATGTCAAGTGCTAATCAATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.012G088100.1 pacid=42783588 polypeptide=Potri.012G088100.1.p locus=Potri.012G088100 ID=Potri.012G088100.1.v4.1 annot-version=v4.1
MPVNPWPLFSFLFFSSTLVLLFPFTAFAVNQQGETLLSWKRSLNGSPEGLNNWDSSNETPCGWFGITCNFNNEVVALGLRYVNLFGTLPSNFTFLSSLNK
LVLSGTNLTGTIPKEIGTALPQLTHLDLSENALTGEIPSELCNFPKLEQLLLNSNQLEGSIPIEIGNLTSLKWLILYDNQLSGSIPNTVGKLKYLEVIRA
GGNKNLEGSLPKEIGNCSNLLMLGLAETSISGFLPPSLGLLKKLQTVAIYTTLLSGQIPPELGDCTELQDIYLYENSLTGSIPKTLGKLRNLRNLLLWQN
NLVGIIPPELGNCNQMLVIDISMNSLTGSIPQSFGNLTELQELQLSLNQISGEIPAQLGNCQKIIHIELDNNQITGSIPPEIGNLFNLTLFYLWQNKLEG
NIPPSISNCQNLEAIDLSQNGLVGPIPKGVFQLKKLNKLLLLSNNLSGEIPPEIGNCSSLIRFRANNNKVSGTIPAHIGNLKNLNFLDLGSNRITGVIPE
EISGCQNLTFLDLHSNAISGNLPQSFDKLISLQFIDFSNNLIEGTLSPSLGSLSSLTKLTLAKNRLSGSIPSQLGSCSKLQLLDLSGNQLSGNIPSSVGK
IPSLEIALNLSLNQLNGEIPSEFTGLNKLGILDISYNHLTGDLQHLAALQNLVVLNVSHNNFSGHVPDTPFFSKLPLSVLAGNPALCFSGNQCDSGDKHV
QRGTAARVAMIVLLCAACALLLAALYIILASKKRGSGAQECEGEDDVEMSPPWEVTLYQKLDLSIADVTRSLTAGNVVGRGRSGVVYKVTIPSGLMVAVK
RFKSAEKISAAAFSSEIATLARIRHRNIVRLLGWGANRKTKLLFYDYMANGTLGTLLHEGNNFGLVEWETRFKIALGVAEGLAYLHHDCVPPILHRDVKA
HNILLGDRFEAYLADFGLARLVEDEHGSFSANPQFAGSYGYIAPEYACMLKITEKSDVYSYGVVLLETITGKKPVDPSFPDGQHVVQWVRNHLRSKKDPV
EILDPKLQGHPDTQIQEMLQALGISLLCTSNRAEDRPTMKDVAVLLKEIRQELITGGEAQKPTNKSSKTMESNPSYSSSSVTPAQLLMLQQGSSRCSLAY
SSSSSTSIMSSANQ
|
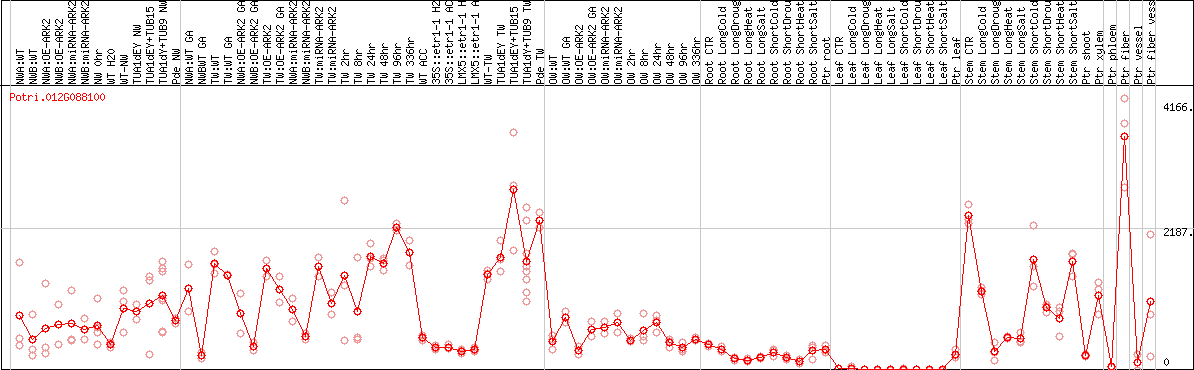
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G088100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
