Potri.012G088500 [POPLAR]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Flax homologues |
|
|||||||||||||||
| PFAM info | ||||||||||||||||
|
Representative CDS sequence |
>Potri.012G088500.7 pacid=42783211 polypeptide=Potri.012G088500.7.p locus=Potri.012G088500 ID=Potri.012G088500.7.v4.1 annot-version=v4.1
ATGACATCAAGTATGTTGACCGCAGAGCGCAGATGGGCTTCTGCAAGAAAAGGTGGCATGAAAGTTTTGGGGAAAGTTCCAGTTCCAAAACCAATAAACC
TACCCAGCCAAAGGTTAGAAAATCATGGTCTGGACCCAAATGTTGAAATTGTGCCCAAGGGTACTCATAGCTGGGGCACCAGATCATCTTCTTCCACACC
TAATGCCTGGGGTTCTTCAACGCTTTCTCCAAACACTGATGGTGGTAGTGGTTCTCCAAGCCATCTCAGTGGCCGCCCTTCATCTGGTGGAAGTGGCACT
CGACCATCAACTGCAAGCAGTGATAGAACCCATGAACCCATCACTAATGCATGGGGTTCAAATTCTAGGCCATCTTCGGCATCAGGAGCATTGACATCAA
ATCAAACATCACCTGTGCCGCTGCGTCCTCGCAGTGCAGAAACAAGACCTGGTAGCTCACAGTTATCTCGGTTTGCTGAACCGTTGTCTGATAATTCAGT
GGCATGGGGTACAACTGGAACAGCAGAAAAGTTGGGTGTGACATCCTCTAAGAATGATGGGTTTTCTTTGACTTCTGGCGATTTTCCAACACTGGGTTCA
GAAAAAGAAATCTCTGGAAAAAACTTGGAGTCCCAAGAGCATGGTTCTTACAGTCGTCCTGGTTCATCATCAAGTGTAGTAGCTCCAGGGAATGAGAGTA
CTGGAAATTCTGCAGGTGATGCTTCTATAAAGACAAATGCAAAAATTGAATCTGCAAATTCTTGGAGGAGAGAGAATCCCATGTATGGTGAAGATGGGTT
GAGGCCTAACATGGAGAAGTGGCACCTTGATCCCCATTTGTATCCTAATTCTAATATTCGTCATCAGAATTATGATTCTTGGCGCGGTCCCCCTGTAAAT
AATCATCCTGGTGGTGTCTGGTACAGAGGGCCTCCAGGAGGTCCGCCTTTTGCACCCCCGATTGCTCCTGGTGGCTTTCCAATAGAGCCATTTCCTTATT
ATCGTCCACAGATTCCCCCTGCTACTCTTGCCAACCCACAGCAAGGTCCTCCACCAGGATCTGGACCAAGAGGACCACATCCAAAGAATGGAGATGTGTT
CAGACCCCATATGCATGATGCCTTTATTCGCCCAGGCATGCCATTTGGGCATGGCTTTTATCCTGGACCAGTCCCTTATGAGAACTACTATGGTCCTCCT
GTGGGCTACTGCAATTCCAATGATCGAGATATTCAGTTTATGGGAATGACAGTGGGTCCTGCTCCCTATAACAGATATTCCGGTCAAAATACTCCTGATC
CTGGAAATTCCCATGGAAGACCAGGTGGATATGGCCCTAGTGGTCACACGATGGTTTCTGAACAACTTGAGTCTGGTCATCAACAAGATACCCGAGGACC
ATACAAAGTCCTTAAGCAGCACGATGGTTCAGAGGGAAAAGATGAAGAACACAAGTGGGATGCCATGATGACAACCAACACATCATATCCTGGGAAGGCA
GACCATCAAAGAAAGTCATCGTGGGAGAATGGCTGGAGAGCAGATGACAAGAAAAATGGAGAGAGAGATACAAGAAGATATGGAGAAGAATTTTCTTTTG
AGGCCACTGACAACCAAGGAGGTGCTAAAGTGAAACCTCTTGAACATGTGGGAAATTGGAAGGCTGCTGCTGATAGTTCAGTTAAGGAATTGGAACATTC
AGAACATGCAGCATCTGCTTTTCCAGAAGTTCCAGCAGCCCCAAAAGATCCCAGTCTGATTAGGAAAATAGGTTTAAATGCAAAGGCCCAGGCCTCTGAT
GGAAGGCAAGAAGTTAAATTTGTTTCCAGTAGAGAGGAGCAGAAGAATAGGTTACAAGTTGGTAATGCCAAGTCTAATCATTCTGCAAATGAAGCTGGTA
CTAGTTATGTATCTCAAAGAACTCATGTCAGTGGCATTGTTGACGCTGGTTTCCATGAAGACCGCATTTCTGCTGCAGATAAGAGCCTTGAGGTACAAAA
TGCTAATGAAACACCAAGCTCCAGGAGATCAACTCAAGGTATGCATGGAAGATCTGATCATCATGGAAAAGGGAGATTTATCACCCAAGAGCCTGATAGG
TGGCAGAGAAGATCCCAAGTTGTAGACTCGCCGTGTGTATTGTCTTCTCATTTTGAAAGTTCTAATGTTTATAGGCAAGACCATAGTTTTGCGGAGGCAA
CTGAAAAGTCTGGGTTATGTCATCAAGGGAAGGATGATGGAGTATCTGTGCCACCTCACCCTGATCCCGGTGATAGCCAGACACATCATGCTACTATTCA
ACGGATTAAACAGCGAGAGAAGGAAGAGGAGGAATGGGAAAGAGAGCAAAAGGCCAAGGCACTTGCAAAAGAGTTGAATAAGTGGACAAAAGCAGCAGAA
AGTTTAAGTGAGGTGCTGCCAGAGAAGCCAAAGGTGACCCATAAGGAGTCCATTGTGACACATGACCAATTGGAACCACTGCTACAGGATGTTAGTCATG
CTGATGCTGATCACCCTGATAATGCCCCACAGATTCATGACAGTAGAGCTTCTAAGCAGAAGCGTGTGAGCTACAGACAGAAGCAGAATGGTCCCTTGGG
AAAGACTTCCAATGATAAGTTATCATCTTCTACAACTGAAGCACCAAAAAATGTTACTGATATAGCAGCTAATGCTCCTGTTTCTCTTGAGGGTGTGAAC
AAACTTACCTCAAACTCAGAATCAACTTTACCCATCAACCTAACTGCCATGGCCGAGTCTTCAGTAAATCATAGAAGGAAGAACAAGAATGGAAAGAACA
AACACAAAATGGATGATGCATCAACATTGGCAGTTGTAACACCAACTCTATCAAAAGAATCAGCTGCTGCTTTAGACACTTCAGCTGGAAGTGGCAAGTC
GGCTTCTGAATCTCTTTTGGATCCAAGCTCATTTCAGCCTCAGACTGATTCTAGAGATGGAAATCAGTCCATGGATCAACGCACATCATCACCAAACGAA
GAAGCTCATGGAAGAGTAAATAACCAGTGGAAAGTGCAACATTTTCGCAGGATGCCAAGGAATCCACAAGCTAATAAATCAACTGAAAAATTCCCAAGTG
GCGATGCTGTTATCTGGGCTCCTGTTCGGTCACAAAGTAAAATTGAGGCTGCTGATGAAGCCACTCAGAAGAACGTGGCTGATGCCATTAGGGCACCCAT
GAAGAGTGATCAGCAAGTGCAGAATAATGCAAGAACTAAACGGGCAGAAATTGAAAGATACATTCCAAAACCTGTAGCCAAAGAAATGGCTCAGCAAGGA
AGTAGTCCCCAGTCAGTGGCTCCATTAATCAATCAGATTACACCAAATGAGACTGCCGGAAAACCCGAATCTGGTTCTCCAAGCGTTGAAAGTTCTCAAA
CCTCTTCCACAGGCATGGGAAAGGTAGGATCCACTTTGGAAGCTAAGAATGGCGATGGCAGGCAAAATAAGTCTGGAAAAATGCATGGATCATGGCGCCA
GCGGGGATCAGCAGAATCAACCACATCTTTTACAAGTAGGAATGTTCAAAAATCAATTGAGCATCAAGTCCAGAAACCTGATGTAAGTTCACCAAAAGAG
CAATTGAGTCATTCTGATGAATGGAATGAGCCTGATGGCTGGAACATTCTTGAAAATATTGATGTGCCAGTCACTACCCTTGCTATAAAAGATCAGGGTG
CAACAGCCAGAGGTAGGCGGCAATCATACAGGGGACAAAAAGGCACAGGATATAGTCATGAGCCTGATGAGAAGAGAATTAATACTGGGGACACCGAAAA
GGTTTATGTCCAAACTTCAGGCTCAGAGATGCATCAAGCTGACTTACCTGCAACTTCAAAAGAAAATCGGTCTGTTGGGGAAAGATCAGCATCTCATTGG
CAACCTAAATCTCAGCCATTTTCAGCAACTAATCAGCGAGGAAGCAGGACCAATGGTGGTCAGAACACAGGCTCTGAAGTTGGTAGGGGTAATAAAAAGG
ATTCCACTTCCCAAACTTTCATGCCACTGTTGTCTCAACCTGGCAGGGATATTGCTACAGTTAAGGCCCGGCCTCATCCTGATCGCTCTCTATCTGAAAA
GAGCATTTTGGAAGAAGTTCCACGCACTGCGCATCAAGAGGGTAAGAATGGAAGGAAGATACCTTCACATAAAGGACGACGTCCTAGTAGCCCAGTTGAA
CCATCTCCTTTGAATATGGATTTTCAACAAGAGCAACGTGTGTCTTCAGGGTTCCAAAAAAATGGAAACCAAAACAGCCGTTTCGGTGGAGAGCATGATT
CTCGTGGCGAGTGGAGTGGTTCTGGGAAAGATAACAAGCAACAAAACGTACCTGCAAACCGTGAAAGGCAGATACAGAATACACATTATGAGTGCCAGCC
AGTAGGGCCACAAAATACTTACAAAGCTAACAATTTTGAATCTTCAAAAGATGTTTCCCATAATTCTGTTGCAAGGTCCAGGGAGAGAGGTCAGGGTCGC
TCAAGGCATGGCGGGGGAAACTCTCATGGATGGCAAACTGGTAGTAGTGTTCGAGTAGATGCCAATTATGATTAG
|
|||||||||||||||
|
AA sequence
|
>Potri.012G088500.7 pacid=42783211 polypeptide=Potri.012G088500.7.p locus=Potri.012G088500 ID=Potri.012G088500.7.v4.1 annot-version=v4.1
MTSSMLTAERRWASARKGGMKVLGKVPVPKPINLPSQRLENHGLDPNVEIVPKGTHSWGTRSSSSTPNAWGSSTLSPNTDGGSGSPSHLSGRPSSGGSGT
RPSTASSDRTHEPITNAWGSNSRPSSASGALTSNQTSPVPLRPRSAETRPGSSQLSRFAEPLSDNSVAWGTTGTAEKLGVTSSKNDGFSLTSGDFPTLGS
EKEISGKNLESQEHGSYSRPGSSSSVVAPGNESTGNSAGDASIKTNAKIESANSWRRENPMYGEDGLRPNMEKWHLDPHLYPNSNIRHQNYDSWRGPPVN
NHPGGVWYRGPPGGPPFAPPIAPGGFPIEPFPYYRPQIPPATLANPQQGPPPGSGPRGPHPKNGDVFRPHMHDAFIRPGMPFGHGFYPGPVPYENYYGPP
VGYCNSNDRDIQFMGMTVGPAPYNRYSGQNTPDPGNSHGRPGGYGPSGHTMVSEQLESGHQQDTRGPYKVLKQHDGSEGKDEEHKWDAMMTTNTSYPGKA
DHQRKSSWENGWRADDKKNGERDTRRYGEEFSFEATDNQGGAKVKPLEHVGNWKAAADSSVKELEHSEHAASAFPEVPAAPKDPSLIRKIGLNAKAQASD
GRQEVKFVSSREEQKNRLQVGNAKSNHSANEAGTSYVSQRTHVSGIVDAGFHEDRISAADKSLEVQNANETPSSRRSTQGMHGRSDHHGKGRFITQEPDR
WQRRSQVVDSPCVLSSHFESSNVYRQDHSFAEATEKSGLCHQGKDDGVSVPPHPDPGDSQTHHATIQRIKQREKEEEEWEREQKAKALAKELNKWTKAAE
SLSEVLPEKPKVTHKESIVTHDQLEPLLQDVSHADADHPDNAPQIHDSRASKQKRVSYRQKQNGPLGKTSNDKLSSSTTEAPKNVTDIAANAPVSLEGVN
KLTSNSESTLPINLTAMAESSVNHRRKNKNGKNKHKMDDASTLAVVTPTLSKESAAALDTSAGSGKSASESLLDPSSFQPQTDSRDGNQSMDQRTSSPNE
EAHGRVNNQWKVQHFRRMPRNPQANKSTEKFPSGDAVIWAPVRSQSKIEAADEATQKNVADAIRAPMKSDQQVQNNARTKRAEIERYIPKPVAKEMAQQG
SSPQSVAPLINQITPNETAGKPESGSPSVESSQTSSTGMGKVGSTLEAKNGDGRQNKSGKMHGSWRQRGSAESTTSFTSRNVQKSIEHQVQKPDVSSPKE
QLSHSDEWNEPDGWNILENIDVPVTTLAIKDQGATARGRRQSYRGQKGTGYSHEPDEKRINTGDTEKVYVQTSGSEMHQADLPATSKENRSVGERSASHW
QPKSQPFSATNQRGSRTNGGQNTGSEVGRGNKKDSTSQTFMPLLSQPGRDIATVKARPHPDRSLSEKSILEEVPRTAHQEGKNGRKIPSHKGRRPSSPVE
PSPLNMDFQQEQRVSSGFQKNGNQNSRFGGEHDSRGEWSGSGKDNKQQNVPANRERQIQNTHYECQPVGPQNTYKANNFESSKDVSHNSVARSRERGQGR
SRHGGGNSHGWQTGSSVRVDANYD
|
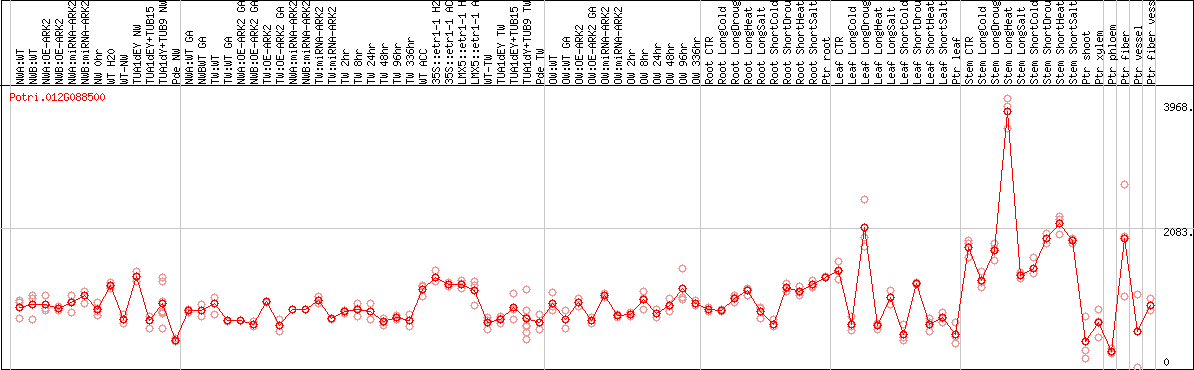
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G088500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
