External link
Symbol
Pt-PMSR1.2
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G61640 251 / 9e-86
PMSR1, ATMSRA1
ARABIDOPSIS THALIANA METHIONINE SULFOXIDE REDUCTASE A1, peptidemethionine sulfoxide reductase 1 (.1)
AT4G25130 245 / 2e-82
PMSR4
peptide met sulfoxide reductase 4 (.1)
AT5G07470 237 / 5e-80
PMSR3, ATMSRA3
ARABIDOPSIS THALIANA METHIONINE SULFOXIDE REDUCTASE 3, peptidemethionine sulfoxide reductase 3 (.1)
AT5G07460 233 / 2e-78
PMSR2, ATMSRA2
ARABIDOPSIS THALIANA METHIONINE SULFOXIDE REDUCTASE 2, peptidemethionine sulfoxide reductase 2 (.1)
AT2G18030 112 / 1e-30
Peptide methionine sulfoxide reductase family protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G087600
337 / 4e-120
AT5G61640 261 / 8e-90
ARABIDOPSIS THALIANA METHIONINE SULFOXIDE REDUCTASE A1, peptidemethionine sulfoxide reductase 1 (.1)
Potri.015G112268
245 / 2e-82
AT4G25130 351 / 4e-123
peptide met sulfoxide reductase 4 (.1)
Potri.012G115800
243 / 6e-82
AT4G25130 351 / 3e-123
peptide met sulfoxide reductase 4 (.1)
Potri.007G012900
115 / 6e-32
AT2G18030 361 / 2e-127
Peptide methionine sulfoxide reductase family protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10033039
303 / 1e-106
AT5G61640 256 / 6e-88
ARABIDOPSIS THALIANA METHIONINE SULFOXIDE REDUCTASE A1, peptidemethionine sulfoxide reductase 1 (.1)
Lus10043002
252 / 3e-85
AT4G25130 346 / 5e-121
peptide met sulfoxide reductase 4 (.1)
Lus10032504
251 / 1e-84
AT4G25130 348 / 6e-122
peptide met sulfoxide reductase 4 (.1)
Lus10013673
223 / 5e-75
AT5G07470 185 / 4e-60
ARABIDOPSIS THALIANA METHIONINE SULFOXIDE REDUCTASE 3, peptidemethionine sulfoxide reductase 3 (.1)
Lus10041922
114 / 3e-31
AT2G18030 370 / 9e-131
Peptide methionine sulfoxide reductase family protein (.1.2)
Lus10028467
112 / 1e-30
AT2G18030 364 / 1e-128
Peptide methionine sulfoxide reductase family protein (.1.2)
Lus10019755
96 / 6e-23
AT5G13800 589 / 0.0
Co-regulated with NYE1, pheophytinase (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01625
PMSR
Peptide methionine sulfoxide reductase
Representative CDS sequence
>Potri.012G091100.2 pacid=42782933 polypeptide=Potri.012G091100.2.p locus=Potri.012G091100 ID=Potri.012G091100.2.v4.1 annot-version=v4.1
ATGGCAACCCACACCACCAATCCTGCTCTGGATCCGGATCTAGACAAACCAGACAACCCCAATCACGAGTTTGCTCAATTCGGAGCTGGATGTTTCTGGG
GTGTCGAGCTTGCCTTTCAGAGATTACATGGTGTGGTCAAAACTGAAGTAGGCTACTCTCAGGGTAACGTCCCTGATCCGACTTACAAACTTGTATGCAC
CAAAACCACCAACCACGTTGAGGTTGTCCGGGTCCAATTTGACCCTGAAGTTTGCCCATATACCAACCTCCTCTCTCTTTTCTGGTCTCGCCATGATCCC
ACCACCCTCAATCGTCAGGGTGGCGATGTGGGTACACAGTATAGATCAGGGATATACTACTACAACGAGGCTCAAGCTAAGTTAGCCCAGGAATCAAAGG
AAGCTAAACAGTTGGGATTGAAGGATAACACGGTTGTGACGGAAATCCTGCCAGCAAAGAGATTTTATAGAGCTGAGGAGTACCATCAGAAGTATCTGGA
AAAAGGTGGGGGTCGAAGCGCTAAACAGTCTGCTGAAAAAGGTTGCAATGACCCTATTAGATGCTATGGTTAA
AA sequence
>Potri.012G091100.2 pacid=42782933 polypeptide=Potri.012G091100.2.p locus=Potri.012G091100 ID=Potri.012G091100.2.v4.1 annot-version=v4.1
MATHTTNPALDPDLDKPDNPNHEFAQFGAGCFWGVELAFQRLHGVVKTEVGYSQGNVPDPTYKLVCTKTTNHVEVVRVQFDPEVCPYTNLLSLFWSRHDP
TTLNRQGGDVGTQYRSGIYYYNEAQAKLAQESKEAKQLGLKDNTVVTEILPAKRFYRAEEYHQKYLEKGGGRSAKQSAEKGCNDPIRCYG
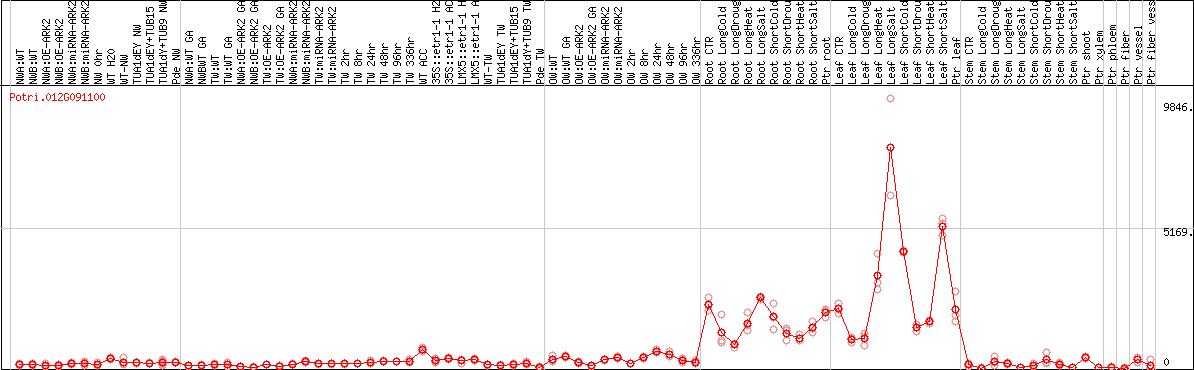
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G091100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.