Potri.012G103700 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.012G103700.1 pacid=42782543 polypeptide=Potri.012G103700.1.p locus=Potri.012G103700 ID=Potri.012G103700.1.v4.1 annot-version=v4.1
ATGGAAACTCCGAAAGAACACATAGAGCACATAAGAGAAACAACATTCTCAATAGGAAGAGAAAAGAACCCCTTGGCTCCCATGCTTGATCAGGCTGTCA
AGTATCTCTCCGCTGAGCTTTACGCCAAAGATGTTCACTTTCTCATGGAGCTCATTCAGAATGCTGAAGACAATGAATACTTGGAGGGGGTGGATCCTTC
ACTTGAGTTTGTTATAACATCTCGTGATATAACAAACACCGGTGCACCTGCTACTTTGCTCATGTTCAATAATGAGAAGGGCTTTTCTGCCAAAAATATT
GATTCAATTTGCAGCGTTGGAAATTCCACTAAGAAAGGGAATCGAAAGCGCGGTTATATTGGGGAAAAAGGGATTGGATTCAAGAGTGTGTTTCTTATAA
CTGCGCAGCCTTACATATTCAGCAATGGGTACCAGATACGATTCAATGAAAATCCCTGCCCACACTGCAATCTTGGATACATAGTTCCTGAATGGGTTCA
TGAGAGTCCATCTCTGTCTGACATCAAACAAATTTATGGTTCCACTTCTATGCTTCCGACTACCACGTTGATTTTACCTCTGAAGCCTGATAAGGTGACA
GCTGTGAAGCAGCAGCTGTCAAGTGTTCATCCCGAAGTCCTGCTGTTCCTTTCAAAGATAAAGCGCCTTTCTGTCAGGGAAGATAATGAGGATCCAAGTC
TCAACACTGTTAGTGCAATAGCTATTACCAAAGAGACTAATTTTGTGACAAGGAAAAACATTGATGCCGAGTCCTACACACTCCATCTGTCTGCAGAGGA
GAATGATGATGAATTTGCAAAAGGATGCAGCTACTACTTGTGGAAGCAGAAATTTCCTGTCAGGCAGGAAAACAGAGTCGACAGGAGAATGGAAGTGGAG
GATTGGGTGATCACTTTGGCTTTTCCTAATGGAGAGCGCCTTCTTAGAGGAATGAAGTACTCTCCTGGAATCTATGCATTTCTTCCAACCGAGATGGTGA
GCAACTTTCCCTTCATAATCCAAGCAGATTTTATTCTAGCATCATCAAGGGAAACAATACAATGGGACAACATATGGAACCAAGGAATCCTTGATTGTGT
TCCCTTTGCTTTTGTCAATGCATTAGTCTCATTAATCAAAACAGTAGATGATGCTCCAGTATCTAGTTTGCCTCCAATGTTCAAGTTCTTGCCAGTCCAC
AGCTCCCCCTTCGAAAAGTTGAATATAGTGAGGGAATCGATTAAATCAAAGCTGGCTGAAGAGGATATTGTTCCAAGTGAGTCATACACAGCGCAAAAGT
TCTTTCATAAACCGCGCCAAGTTTGCCGATTAATGCCTGCTTTCTGGAACATACTGAAGATGGCAAGGGAGCGAGGAGTGAGCTTGCACAATCTTTCATC
CCATGGTTGCTATGTTCTGAATTTTTCTTTTGATAAACCAGAGTATGATCACATACTGGATTTCTTGAGGGTGGAACCGGTAAGCAGTGAGTGGTATGTA
AAGTGCATCCAAGGTTCTCATATTGTAATGGGAGTCTCAGAGGAAACATACCTGGAGCTTCTTCACTTCCTTGCTGTTAATTGGCATTCCTTGTTTTACC
ACACAGACATGGGGAGCATTCCACTCATTAAATATGTGGGTGTTGATGGGAGCGTGTCTTTGTGCACCGTTAATGAATCTGCTCAGTGGTATGGTAAAAC
TCTGTGCCTTTCACTTCTGTCATCTCATATCTCATGGTTGATTGATTGGAACAGGGAATTCCGATGCATGGCAAATCATTTTTTTATGCCAAGAAGTACA
CAAGAAGCTATCCGGTCATCATCTAGCAAAAATGAGGTGTTAGAATGGCTTGGAGATCCTGTTAAAGTTACTGCTTTAAGCGTAAATGATTATGCAGTTC
TCTGCGGAAATCAAGTCAGTAGTGACCGGAAGCTTGTCATTGCCTATGCTCATTTCTTATATCATTCATTCTCCAATAACTATCTATCAGGCAGGGAAGT
TGCCCCTTTATGTGATAAAATGCCACTTGTTGATAGCTATGGTCATGTGATTAAAGCAAGGAATGGGGTTCTTGTCCCGGCTCCTGAAAGCAAATGGGTG
CAATTGATTGGTTATAATCCTTGGAGGGGAGAAAGCTACGTTGAGTTAGGAGAAGACTACTTGCATCCTGGATATTTTGCTGGCACAAGCACAGAGGGAA
AGAAACTCTTGGAGTTCCTTAAAGCTTTTGTTAAAGCTTCTGATATTCCTCACATACCTCCCCCGATTGCTGGAATACCTACTGCGTCAACACCACTTAC
CAAACAAAATGCATTTCTGCTGTTGGACTGGATTCGTGAGCTGAAACGGAGTGGAATTTCCATTCCAGCAACGTTCATGAATTGCATAAAGGAGGGTAGC
TGGCTGAAAATTACTATGAATGGCTCTCCTGGTTACAAGCCACCATCACAGTCATTCCTACTTGGTTCAGTTAACAGAAGCTCAGACTGGGGAAACATTT
TGCAGAATGGATCCGTGCTTGTCGATATTCCGTTGATAGATCAAGGTTTTTATGGTTATAAAATTAACGAGTACAGAGAGGAGCTAATGACAGTCGGAGT
CATGTTCGAGTATGGAGAGGCATGTGAATTTATCGGGAATCGTCTCATGTCTTTGGCAGCTTCATCTACCTTGACTAAAAGCAATGTGATTTCCATACTG
AAATTCATCAGATTTTTGACGCTAAATCTTCTTCCTCCAGATAAATTCATTCTCAGAATTAAAGAAGGAAGGTGGCTAAAGACAGGTGGGGGATACAGGT
CTCCAGTTGGTTCTGTTCTATATGATCAGGAGTGGACAATTGCGAGGCAAATTAGTGACATCCCTTTTATTGATCAAGATTACTATGGCAAAGACATCCT
TGTTTTCAAATCAGAACTCCAGTTACTGGGCGTCGCAATTGGCTTCAGCGGAAGTTATCAATTGGTTGCTGATTACTTAAAATCGCCTTTATGGTTATCT
TATCTGACAATGGAGGCGTTTCTTCTGGTTTTAGATTGCATGCGTCATTCTAGTTCTGCTGGCAAGCTAGTTATCGCACTGAAAAGCACAAAATGCTTAA
ACACAACCTTGGGCTACAGATATCCTGATGACTGTTTCTTGTTTCATCCTGAATGGGGTTGCCTTCTAAATGTTTTTGGTGGTTTTCCATTGGTTGATAG
TAATTTCTATGGAAGCAACATCATCTCTTACAAAAAGGAGTTGAAGGACCTGGGAGTAAGAGTAGACTTCGAGGATGCTGTCGAAGTGTTTGTTGATACT
TTCAGGAAGCAGGCATCTTCCATGACAAAGGAAAGTGTATTTTCATTCATATCATGCTACAGAAAGCTGAAGGGAACTCCACACAAATTTCCTTCAGACC
TTAAGAAGTGCATCCGTGAGGAAAATTGGCTGCGGACCCGACTCGGTGATTATAAATCTCCAAGCAACTGTATCCTGTTTAGTCCTGAGTGGAAGTCAAT
TTATCCAATTACTCGCCTTCCCTTTATTGATGACAGTGACAAATATTATGGGAATGACATTCATGAATATCAGAAAGAGCTGAAGAGTATGGGGGTCATT
GTTGAATTTAAAGCTGGTGTGAAGTTTGTCGCTGCTGGTCTTCGTTTTCCCCAAAATCCATGCCACATAGCTCGTGTGAATGTGCTTTCATTGCTGGAAT
GCATCCGTGCTTTATTGCAAGAAAAGGACTACTCCTTTCCTGAAATATTTCTGAAAAATATTTCTCAGGGATGGTTAAAGACCCATGCTGGTTTCAGGTC
TCCGGGCAACTGCTGTTTGTTCAATTCCCAGTGGAGTTCATACGTGAAGCCTACTGATGGACCATTCATTGATGAAGACTTTTATGGTTCTAACATCAAG
TTGTATGGAAAAGAGCTCAGTGCAATAGGAGTTCATTTAGAGGTAGAAAAGGCGTGTTCATTGCTTGCCAGTCATCTTGATTCCCATTCTGAGTTTTGCA
CTATAGTTCGAGTATATGATTTCCTGAGACAACATGAGTGGAAGCCAGATGGTGATGCAACCAGAAAGATTTGGATTCCAGATGGACTTGAGAATGGAAT
GTGGGTTAACCCAGAAGAATGTGTTCTGCATGACAAAGATGGTCTCTTTGGTCTTCAGCTGAATGTATTGGAGAAACATTATGAGCCAGAATTACTTCTT
TTCTTTTCCAGTTCATTCAAGGTTAGATCCAATCCTTCATTTGATGATTACTGTAAGCTTTGGAAGGTTTGGGAAAGTTTAGGACGCCCATTGACACATG
CTGAGTGCTGTGCATTTTGGAAGTGTGTTATGACGCATATGAGCTCAAAGACAGAGAGAACTCTCGCCGATGACTTGGTAAAGCTACCTGTTATTTTGGG
CTCTGGTGAAATAGTGTTGTTCAGGAAGGCTGACGTTTTTATTGCTGATGACCTTCTACTGAAGGACCTTTTCGAAAGGTTTTCATCACGCCCAATATTT
GTCTGGTGTCCCCAGCCAAACTTGCCTTCTTTACCTCGAACTAGGTTGCTTGATGTATACAGGAAAATTGGGGTTCGCACAATATCTGAGTCAGTGCAAA
AGGAGGAATTATCATTGGCAGATGGTGTGGAATTTAGTCAGATGAATCCGAGGAATGCCATGATTGGAAAAGAGCTGGTGAGACTGATTCTTGGCTTTCT
AGCAGATCCTTCTCTTGATATTGAAGCAACAAAAAGGCATGGAGCTGTCCAGTGCCTCTTGAACCTGAAGGTCCTGGAAACTATGGAGGCAATTGCTGTA
AGCTATAGTTTACCGCTTTCTGATGGGAAAATTCTGAAGGTGGAGAACGCACGCAGTATGATTCGCTGGGATAAGGAGAGTTCAAAGTTTCTCACACAGA
AGATGGATGAAGCTGGTGGCCAGAAAAATCTTATTGAATTTGCAACCATTTTTTCTGAAGTAATAGCTCGGGGGGTGCTATGGGATAAAGAAGATCAAAT
AAAAGCACTCTCCGAGCTGATCAGGTTGGCTTTCGTGCTGAATTTTGATGAACAAGCTGTTCAGTTCTTGATGAAATCCAATAATCTTCAAACTTTTCTG
GAGGATGAGGAGTTCCTTGCTGCTGCTTTCCCTTCTGTTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.012G103700.1 pacid=42782543 polypeptide=Potri.012G103700.1.p locus=Potri.012G103700 ID=Potri.012G103700.1.v4.1 annot-version=v4.1
METPKEHIEHIRETTFSIGREKNPLAPMLDQAVKYLSAELYAKDVHFLMELIQNAEDNEYLEGVDPSLEFVITSRDITNTGAPATLLMFNNEKGFSAKNI
DSICSVGNSTKKGNRKRGYIGEKGIGFKSVFLITAQPYIFSNGYQIRFNENPCPHCNLGYIVPEWVHESPSLSDIKQIYGSTSMLPTTTLILPLKPDKVT
AVKQQLSSVHPEVLLFLSKIKRLSVREDNEDPSLNTVSAIAITKETNFVTRKNIDAESYTLHLSAEENDDEFAKGCSYYLWKQKFPVRQENRVDRRMEVE
DWVITLAFPNGERLLRGMKYSPGIYAFLPTEMVSNFPFIIQADFILASSRETIQWDNIWNQGILDCVPFAFVNALVSLIKTVDDAPVSSLPPMFKFLPVH
SSPFEKLNIVRESIKSKLAEEDIVPSESYTAQKFFHKPRQVCRLMPAFWNILKMARERGVSLHNLSSHGCYVLNFSFDKPEYDHILDFLRVEPVSSEWYV
KCIQGSHIVMGVSEETYLELLHFLAVNWHSLFYHTDMGSIPLIKYVGVDGSVSLCTVNESAQWYGKTLCLSLLSSHISWLIDWNREFRCMANHFFMPRST
QEAIRSSSSKNEVLEWLGDPVKVTALSVNDYAVLCGNQVSSDRKLVIAYAHFLYHSFSNNYLSGREVAPLCDKMPLVDSYGHVIKARNGVLVPAPESKWV
QLIGYNPWRGESYVELGEDYLHPGYFAGTSTEGKKLLEFLKAFVKASDIPHIPPPIAGIPTASTPLTKQNAFLLLDWIRELKRSGISIPATFMNCIKEGS
WLKITMNGSPGYKPPSQSFLLGSVNRSSDWGNILQNGSVLVDIPLIDQGFYGYKINEYREELMTVGVMFEYGEACEFIGNRLMSLAASSTLTKSNVISIL
KFIRFLTLNLLPPDKFILRIKEGRWLKTGGGYRSPVGSVLYDQEWTIARQISDIPFIDQDYYGKDILVFKSELQLLGVAIGFSGSYQLVADYLKSPLWLS
YLTMEAFLLVLDCMRHSSSAGKLVIALKSTKCLNTTLGYRYPDDCFLFHPEWGCLLNVFGGFPLVDSNFYGSNIISYKKELKDLGVRVDFEDAVEVFVDT
FRKQASSMTKESVFSFISCYRKLKGTPHKFPSDLKKCIREENWLRTRLGDYKSPSNCILFSPEWKSIYPITRLPFIDDSDKYYGNDIHEYQKELKSMGVI
VEFKAGVKFVAAGLRFPQNPCHIARVNVLSLLECIRALLQEKDYSFPEIFLKNISQGWLKTHAGFRSPGNCCLFNSQWSSYVKPTDGPFIDEDFYGSNIK
LYGKELSAIGVHLEVEKACSLLASHLDSHSEFCTIVRVYDFLRQHEWKPDGDATRKIWIPDGLENGMWVNPEECVLHDKDGLFGLQLNVLEKHYEPELLL
FFSSSFKVRSNPSFDDYCKLWKVWESLGRPLTHAECCAFWKCVMTHMSSKTERTLADDLVKLPVILGSGEIVLFRKADVFIADDLLLKDLFERFSSRPIF
VWCPQPNLPSLPRTRLLDVYRKIGVRTISESVQKEELSLADGVEFSQMNPRNAMIGKELVRLILGFLADPSLDIEATKRHGAVQCLLNLKVLETMEAIAV
SYSLPLSDGKILKVENARSMIRWDKESSKFLTQKMDEAGGQKNLIEFATIFSEVIARGVLWDKEDQIKALSELIRLAFVLNFDEQAVQFLMKSNNLQTFL
EDEEFLAAAFPSV
|
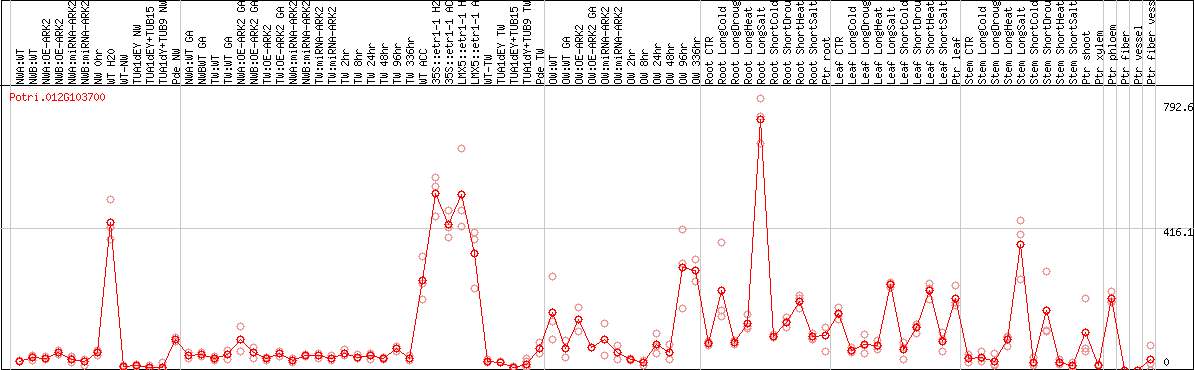
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G103700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
