External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G47140 251 / 1e-83
AtSDR5
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G29250 228 / 1e-74
AtSDR4
short-chain dehydrogenase reductase 4, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT2G47130 220 / 1e-71
AtSDR3
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G29260 216 / 3e-70
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT2G47120 216 / 6e-70
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G51680 192 / 5e-60
AtSDR2
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G26760 187 / 2e-58
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G26770 177 / 2e-54
NAD(P)-binding Rossmann-fold superfamily protein (.1)
AT3G42960 172 / 1e-52
ASD, ATA1
TAPETUM1, TAPETUM 1 (.1)
AT4G03140 174 / 2e-52
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G105600
501 / 0
AT2G47140 230 / 1e-75
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.015G104800
461 / 2e-166
AT2G47140 243 / 2e-80
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.015G065700
403 / 2e-143
AT2G47140 248 / 1e-82
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.015G065600
300 / 1e-102
AT2G47140 258 / 1e-86
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.014G115300
236 / 8e-78
AT2G47140 346 / 3e-121
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G207100
215 / 2e-69
AT4G03140 241 / 2e-78
NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.016G074000
208 / 1e-66
AT1G52340 246 / 5e-81
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.004G199900
208 / 1e-66
AT2G47140 256 / 1e-85
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Potri.006G105900
197 / 6e-62
AT3G51680 412 / 4e-146
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10042181
271 / 3e-91
AT2G47130 255 / 3e-85
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10009992
237 / 6e-78
AT2G47140 345 / 1e-120
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10038057
233 / 2e-76
AT2G47140 341 / 4e-119
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10021320
214 / 9e-69
AT1G52340 246 / 2e-81
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016997
211 / 1e-67
AT1G52340 249 / 2e-82
SHORT-CHAIN DEHYDROGENASE REDUCTASE 1, IMPAIRED SUCROSE INDUCTION 4, GLUCOSE INSENSITIVE 1, SHORT-CHAIN DEHYDROGENASE/REDUCTASE 1, ARABIDOPSIS THALIANA ABA DEFICIENT 2, ABA DEFICIENT 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10016178
204 / 6e-65
AT2G47140 210 / 2e-67
short-chain dehydrogenase reductase 5, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10026419
194 / 2e-60
AT3G51680 367 / 9e-128
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10014226
191 / 9e-60
AT3G51680 230 / 1e-74
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10029369
186 / 3e-58
AT3G51680 226 / 3e-73
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
Lus10021312
184 / 3e-57
AT3G51680 238 / 8e-78
short-chain dehydrogenase/reductase 2, NAD(P)-binding Rossmann-fold superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF08659
KR
KR domain
Representative CDS sequence
>Potri.012G105700.1 pacid=42783075 polypeptide=Potri.012G105700.1.p locus=Potri.012G105700 ID=Potri.012G105700.1.v4.1 annot-version=v4.1
ATGGCATCTACAAAACCTTGCAAGAACAAGGTTCAAGACAAGGTTGCTATCGTCACGGGAGGTGCCAGTGGCATAGGCGAGGCCACCGTGCTTGCCTTTG
CTGAAAATGGCGCACGAGCAGTCGTGATCGCAGATATTCAAGATGAAAAGGGCCAAAAGCTTGCAGAATCCATCGGAACTAACAGATCCACTTACATCCA
CTGTGATGTAGGTGATGAAAACCAAGTCAAATCCCTTGTGGAGTCCACTGTTCAACTTTATGGTCACCTTGACGTAATATTCTGCAATGCCGGCATTGCA
AGCTTTGGTAAGCAAAATGTTCTTGACTTCGACCTGGACTCGTGCGACAAACTCTTTGCAGTAAACGTGCGTGGCACAGCAGCATGCCTAAAGCACGCGG
CGCGTGCCATGGTGGACGGAGGTGTAAAGGGCAGCGTCATTTGCACATCAAGCGCCGCTGCGAATCTCGCAGGGGTCAGGTTCACAGATTATATCATGTC
CAAGAGCGGGGTTTTGGCACTGATGAAATGTGCAAGTTATCAGTTGGGTGAGCATGGTATAAGAGTCAACTGTGTGTCGCCAGGACCCGTGGCAACACCT
CTAGCATGCAAGACATTCGAAAAGGGAGTAGAGGAGGTGGAGAAGGCTTTTCAGTCGTCTTATTGCTTGAAAGGTGTGTTGAAAACGAAGCACGTAGCTG
ACGCAGTGCTGTTTCTCGCTTCTGACGATTCTGAGTTTGTGACTGGCCAGAATTTGATCGTCGATGGTGGGTTTAATTTTCAAGGTATTCCCAAATAA
AA sequence
>Potri.012G105700.1 pacid=42783075 polypeptide=Potri.012G105700.1.p locus=Potri.012G105700 ID=Potri.012G105700.1.v4.1 annot-version=v4.1
MASTKPCKNKVQDKVAIVTGGASGIGEATVLAFAENGARAVVIADIQDEKGQKLAESIGTNRSTYIHCDVGDENQVKSLVESTVQLYGHLDVIFCNAGIA
SFGKQNVLDFDLDSCDKLFAVNVRGTAACLKHAARAMVDGGVKGSVICTSSAAANLAGVRFTDYIMSKSGVLALMKCASYQLGEHGIRVNCVSPGPVATP
LACKTFEKGVEEVEKAFQSSYCLKGVLKTKHVADAVLFLASDDSEFVTGQNLIVDGGFNFQGIPK
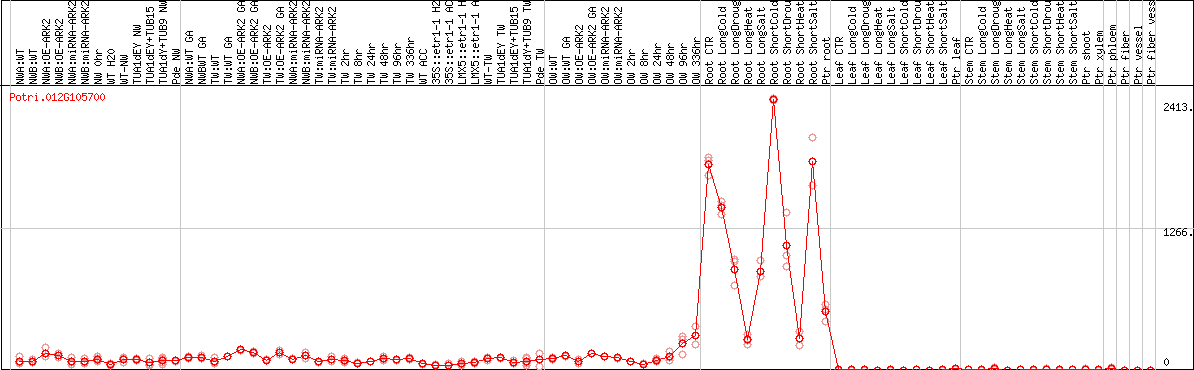
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G105700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.