External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G51100 221 / 1e-72
FSD2
Fe superoxide dismutase 2 (.1)
AT4G25100 179 / 2e-57
ATFSD1, FSD1
ARABIDOPSIS FE SUPEROXIDE DISMUTASE 1, Fe superoxide dismutase 1 (.1.2.3.4.5)
AT5G23310 164 / 1e-50
FSD3
Fe superoxide dismutase 3 (.1)
AT3G10920 69 / 2e-14
MSD1, MEE33, ATMSD1
MATERNAL EFFECT EMBRYO ARREST 33, ARABIDOPSIS MANGANESE SUPEROXIDE DISMUTASE 1, manganese superoxide dismutase 1 (.1.2)
AT3G56350 65 / 6e-13
Iron/manganese superoxide dismutase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G110400
257 / 1e-86
AT5G51100 335 / 3e-115
Fe superoxide dismutase 2 (.1)
Potri.005G089600
149 / 1e-44
AT5G23310 340 / 1e-118
Fe superoxide dismutase 3 (.1)
Potri.013G092600
69 / 2e-14
AT3G10920 370 / 6e-132
MATERNAL EFFECT EMBRYO ARREST 33, ARABIDOPSIS MANGANESE SUPEROXIDE DISMUTASE 1, manganese superoxide dismutase 1 (.1.2)
Potri.019G057300
66 / 4e-13
AT3G10920 360 / 6e-128
MATERNAL EFFECT EMBRYO ARREST 33, ARABIDOPSIS MANGANESE SUPEROXIDE DISMUTASE 1, manganese superoxide dismutase 1 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026881
226 / 4e-74
AT5G51100 373 / 3e-130
Fe superoxide dismutase 2 (.1)
Lus10017378
169 / 7e-52
AT5G23310 368 / 5e-129
Fe superoxide dismutase 3 (.1)
Lus10010174
163 / 9e-50
AT5G23310 359 / 1e-125
Fe superoxide dismutase 3 (.1)
Lus10034222
69 / 3e-14
AT3G10920 370 / 2e-131
MATERNAL EFFECT EMBRYO ARREST 33, ARABIDOPSIS MANGANESE SUPEROXIDE DISMUTASE 1, manganese superoxide dismutase 1 (.1.2)
Lus10012883
69 / 8e-14
AT3G10920 356 / 1e-123
MATERNAL EFFECT EMBRYO ARREST 33, ARABIDOPSIS MANGANESE SUPEROXIDE DISMUTASE 1, manganese superoxide dismutase 1 (.1.2)
Lus10030534
57 / 6e-10
AT3G10920 241 / 8e-81
MATERNAL EFFECT EMBRYO ARREST 33, ARABIDOPSIS MANGANESE SUPEROXIDE DISMUTASE 1, manganese superoxide dismutase 1 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00081
Sod_Fe_N
Iron/manganese superoxide dismutases, alpha-hairpin domain
PF02777
Sod_Fe_C
Iron/manganese superoxide dismutases, C-terminal domain
Representative CDS sequence
>Potri.012G112301.2 pacid=42784002 polypeptide=Potri.012G112301.2.p locus=Potri.012G112301 ID=Potri.012G112301.2.v4.1 annot-version=v4.1
ATGTTTTTCTTACCTTCACTCTCTTTTTTCTTTCTTCATCCGACTCAGTGTTTTCTTGCATTTCAGGTCTTACAGCAACTGGTGTGCTGCGATGGACAGA
GGGTTGCCAGTATTACACGTTCGAGGCAATGTACCAGAAAGGCTGATGCTTTGACAGTTACAGCTAAATTTGAGCTGAAACCTCCTCCATATCCCATGGA
TGATTTAGAGCCGCATATGAGCAAGGACACATTTGAGTATCACCGGGGAAAGCATCACAGGGCTTATGTGGATAACTTAAACAAGCAAATTGACGGAACA
GAACGAGATGACATGTCCTTAGATGATGTTGTGCTCGTTACATACAACAAGGGTGGTCCACTTCCTGCTTTCAACAATGCTGCACAGGCATGGAACCATG
AATTCTTTTTGGAATCCATGAAACCGGGAGGTGGAGGAAAGGCATCAGGGGAACTTCTTCACTTGATTGAAAGAGATTTTGGTTCTTTTGATAGATTTGT
GCAAGAGTTCAAGTCGGCTGCAGCTACTCAGTTTGGTTCTGGATGGGCTTGGCTTGTTTCCACTCCATACAATTGA
AA sequence
>Potri.012G112301.2 pacid=42784002 polypeptide=Potri.012G112301.2.p locus=Potri.012G112301 ID=Potri.012G112301.2.v4.1 annot-version=v4.1
MFFLPSLSFFFLHPTQCFLAFQVLQQLVCCDGQRVASITRSRQCTRKADALTVTAKFELKPPPYPMDDLEPHMSKDTFEYHRGKHHRAYVDNLNKQIDGT
ERDDMSLDDVVLVTYNKGGPLPAFNNAAQAWNHEFFLESMKPGGGGKASGELLHLIERDFGSFDRFVQEFKSAAATQFGSGWAWLVSTPYN
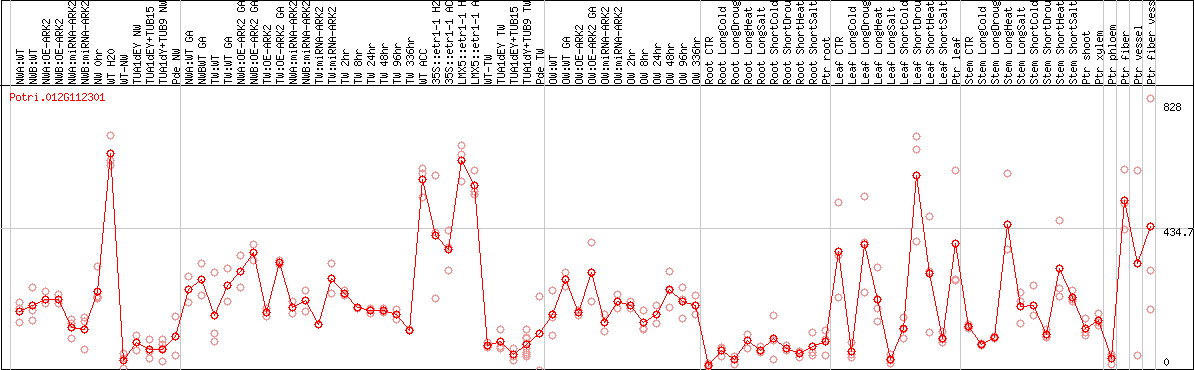
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G112301 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.