External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G65260 273 / 2e-93
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT5G51120 266 / 1e-90
PABN1, ATPABN1
polyadenylate-binding protein 1 (.1.2.3)
AT5G10350 263 / 4e-90
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3)
AT3G12640 70 / 9e-14
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
AT2G16940 59 / 5e-10
Splicing factor, CC1-like (.1.2.3)
AT3G55460 55 / 6e-09
SCL30, At-SCL30
SC35-like splicing factor 30 (.1)
AT2G36660 54 / 2e-08
PAB7
poly(A) binding protein 7 (.1)
AT3G10400 52 / 6e-08
U11/U12-31K
U11/U12-31K, RNA recognition motif and CCHC-type zinc finger domains containing protein (.1)
AT2G35410 51 / 1e-07
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT1G74230 51 / 1e-07
GR-RBP5
glycine-rich RNA-binding protein 5 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G110800
390 / 1e-139
AT5G51120 266 / 1e-90
polyadenylate-binding protein 1 (.1.2.3)
Potri.005G073200
286 / 9e-99
AT5G10350 292 / 2e-101
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3)
Potri.007G095800
281 / 5e-97
AT5G10350 285 / 3e-98
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2), RNA-binding (RRM/RBD/RNP motifs) family protein (.3)
Potri.017G051600
257 / 2e-87
AT5G65260 288 / 4e-100
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.001G311600
234 / 2e-78
AT5G65260 238 / 2e-80
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.012G145050
82 / 2e-19
AT2G24350 107 / 4e-27
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.001G269200
71 / 3e-14
AT3G12640 432 / 3e-143
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.009G063800
71 / 7e-14
AT3G12640 418 / 6e-138
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.004G177000
61 / 2e-10
AT2G16940 538 / 0.0
Splicing factor, CC1-like (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10043008
327 / 5e-115
AT5G65260 278 / 4e-96
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10032509
336 / 2e-112
AT5G07440 771 / 0.0
glutamate dehydrogenase 2 (.1.2.3)
Lus10032510
317 / 9e-111
AT5G65260 278 / 2e-95
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10009323
248 / 4e-84
AT5G65260 268 / 2e-92
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10015840
244 / 1e-82
AT5G65260 261 / 9e-90
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10037701
79 / 1e-16
AT3G12640 412 / 2e-135
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10015696
78 / 2e-16
AT3G12640 417 / 1e-136
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10025075
61 / 9e-11
AT2G16940 565 / 0.0
Splicing factor, CC1-like (.1.2.3)
Lus10034466
61 / 1e-10
AT2G16940 569 / 0.0
Splicing factor, CC1-like (.1.2.3)
Lus10018277
57 / 3e-09
AT4G19610 770 / 0.0
nucleotide binding;nucleic acid binding;RNA binding (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Potri.012G115100.1 pacid=42783908 polypeptide=Potri.012G115100.1.p locus=Potri.012G115100 ID=Potri.012G115100.1.v4.1 annot-version=v4.1
ATGGATCACCTTCACGATCACGAAGAGGACGAGCACGAGCACGAGGTCTACGGAGGAGAAATCCCTGATGAAGGAGAGATGGATGCCGATGTTGACATGT
CGTCTAGAGCTGAAGAAGATGAGTACCAAGACCCTAACTCCAAGGATTTAGAGGATATGAAGAAAAGATTGAAAGAGATCGAAGAAGAAGCCGGTGCACT
ACGTGAAATGCAGGCTAAGGTTGAGAAAGAGATGGGAGCTGTTCAAGATTCTCCTGGTGCTTCTGCTACTCAAGCTGAAAAGGAGGAGGTGGATTCCCGA
TCCATTTATGTTGGTAATGTGGATTATTCTTGTACCCCTGAAGAAGTGCAGCAGCATTTTCAGTCCTGTGGAACTGTGAATAGAGTGACTATTTTGACTG
ACAAGTTTGGTCAACCAAAGGGATTTGCTTACGTTGAGTTTGTAGAGGTTGATGCTATTCAGAATGCTCTCTTATTAAATGAATCAGAACTTCATGGCCG
CCAGTTAAAGGTCTCTGCGAAGCGAACCAATGTTCCTGGAATGAAGCAATTTCGAGGAAGGCGCCCAAGCCCTTATGGGCTCAGGTCCCGGAGGCCATTC
ATGCCAGCTCCCTTCTATCCTGCATATGGTTATGGAAGGGTTCCGAGGTTCAGGAGACCTATGCGATACAGGCCATACTAG
AA sequence
>Potri.012G115100.1 pacid=42783908 polypeptide=Potri.012G115100.1.p locus=Potri.012G115100 ID=Potri.012G115100.1.v4.1 annot-version=v4.1
MDHLHDHEEDEHEHEVYGGEIPDEGEMDADVDMSSRAEEDEYQDPNSKDLEDMKKRLKEIEEEAGALREMQAKVEKEMGAVQDSPGASATQAEKEEVDSR
SIYVGNVDYSCTPEEVQQHFQSCGTVNRVTILTDKFGQPKGFAYVEFVEVDAIQNALLLNESELHGRQLKVSAKRTNVPGMKQFRGRRPSPYGLRSRRPF
MPAPFYPAYGYGRVPRFRRPMRYRPY
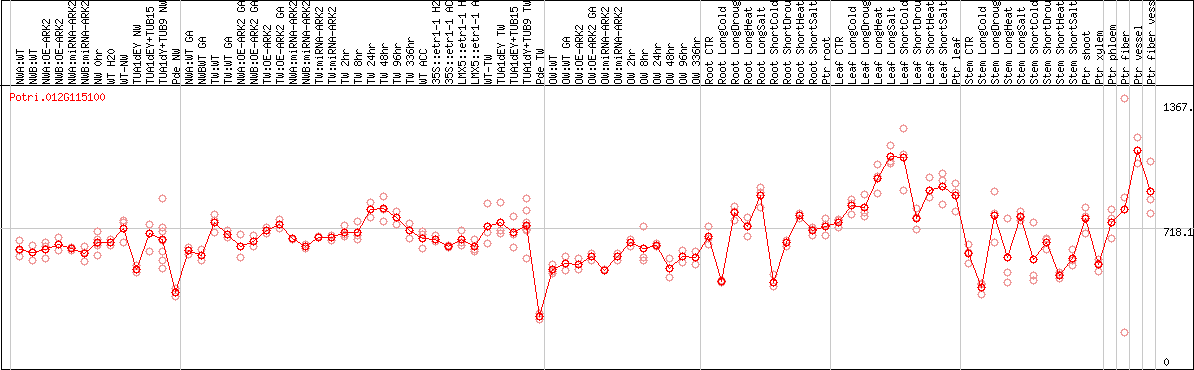
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G115100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.