External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G33120 364 / 2e-129
ATVAMP722, SAR1
ARABIDOPSIS THALIANA VESICLE-ASSOCIATED MEMBRANE PROTEIN 722, synaptobrevin-related protein 1 (.1.2.3)
AT1G04750 359 / 1e-127
ATVAMP7B, ATVAMP721, VAMP7B
VESICLE-ASSOCIATED MEMBRANE PROTEIN 7B, vesicle-associated membrane protein 721 (.1.2)
AT1G04760 358 / 2e-127
ATVAMP726
vesicle-associated membrane protein 726 (.1)
AT2G32670 354 / 2e-124
ATVAMP725
vesicle-associated membrane protein 725 (.1)
AT4G15780 296 / 8e-103
ATVAMP724
vesicle-associated membrane protein 724 (.1)
AT2G33110 286 / 5e-99
ATVAMP723
vesicle-associated membrane protein 723 (.1)
AT3G54300 277 / 5e-95
ATVAMP727
vesicle-associated membrane protein 727 (.1.2)
AT5G11150 162 / 3e-50
ATVAMP713
vesicle-associated membrane protein 713 (.1)
AT4G32150 157 / 4e-48
ATVAMP711, VAMP7C
vesicle-associated membrane protein 711 (.1)
AT5G22360 150 / 4e-45
ATVAMP714
vesicle-associated membrane protein 714 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G118300
404 / 2e-145
AT2G33120 359 / 8e-128
ARABIDOPSIS THALIANA VESICLE-ASSOCIATED MEMBRANE PROTEIN 722, synaptobrevin-related protein 1 (.1.2.3)
Potri.003G177700
359 / 8e-128
AT1G04750 389 / 1e-139
VESICLE-ASSOCIATED MEMBRANE PROTEIN 7B, vesicle-associated membrane protein 721 (.1.2)
Potri.002G240900
345 / 5e-122
AT2G33120 355 / 3e-126
ARABIDOPSIS THALIANA VESICLE-ASSOCIATED MEMBRANE PROTEIN 722, synaptobrevin-related protein 1 (.1.2.3)
Potri.001G050400
342 / 1e-120
AT2G33120 365 / 4e-130
ARABIDOPSIS THALIANA VESICLE-ASSOCIATED MEMBRANE PROTEIN 722, synaptobrevin-related protein 1 (.1.2.3)
Potri.008G209100
292 / 5e-101
AT4G15780 367 / 9e-131
vesicle-associated membrane protein 724 (.1)
Potri.010G239900
280 / 6e-96
AT3G54300 387 / 6e-138
vesicle-associated membrane protein 727 (.1.2)
Potri.008G019400
277 / 4e-95
AT3G54300 407 / 3e-146
vesicle-associated membrane protein 727 (.1.2)
Potri.018G025800
159 / 1e-48
AT4G32150 366 / 2e-130
vesicle-associated membrane protein 711 (.1)
Potri.009G018900
152 / 4e-46
AT5G22360 367 / 6e-131
vesicle-associated membrane protein 714 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036203
392 / 2e-140
AT1G04760 367 / 1e-130
vesicle-associated membrane protein 726 (.1)
Lus10040611
391 / 3e-140
AT1G04760 369 / 1e-131
vesicle-associated membrane protein 726 (.1)
Lus10038342
390 / 6e-140
AT1G04760 366 / 3e-130
vesicle-associated membrane protein 726 (.1)
Lus10018296
389 / 2e-139
AT1G04760 366 / 2e-130
vesicle-associated membrane protein 726 (.1)
Lus10038166
363 / 3e-129
AT1G04760 383 / 3e-137
vesicle-associated membrane protein 726 (.1)
Lus10025933
360 / 5e-128
AT1G04760 381 / 2e-136
vesicle-associated membrane protein 726 (.1)
Lus10022804
359 / 2e-127
AT1G04760 377 / 1e-134
vesicle-associated membrane protein 726 (.1)
Lus10011870
353 / 3e-125
AT1G04760 376 / 3e-134
vesicle-associated membrane protein 726 (.1)
Lus10000754
343 / 5e-121
AT1G04760 372 / 8e-133
vesicle-associated membrane protein 726 (.1)
Lus10011737
296 / 8e-103
AT1G04760 324 / 4e-114
vesicle-associated membrane protein 726 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0445
SNARE-fusion
PF00957
Synaptobrevin
Synaptobrevin
CL0431
PF
PF13774
Longin
Regulated-SNARE-like domain
Representative CDS sequence
>Potri.012G119600.1 pacid=42782556 polypeptide=Potri.012G119600.1.p locus=Potri.012G119600 ID=Potri.012G119600.1.v4.1 annot-version=v4.1
ATGAACCAGAAATCACTGATTTACGCGTTTGTTTCACGAGGAACTGTGATTCTTGCTGAGTTCACTGAATTCAGCGGGAATTTCAATTCCATAGCGTTTC
AATGTCTCCAAAAACTTCCTGCTACTAACAACAAGTTTACTTACAACTGCGATGGTCATACTTTCAATTACCTCGCCGACAATGGGTTTACATATTGTGT
TGTTGCGGATGAATCGGCTGGAAGACAAGTGCCTATGGCTTTTCTGGAGCGTGTCAAGGATGATTTCGTGTCAAAGTATGGCGGTGGAAAGGCTGCCACA
GCCCAGGCCAATGGTCTTAACAAGGAATTTGGGCCAAAATTGAAGGAACATATGAAGTATTGTGCTGATCATCCTGAAGAGATAAGCAAGCTTGCGAAAG
TGAAAGCTCAGGTTTCAGAAGTTAAAGGTGTCATGATGGAGAACATTGAGAAGGTTCTGGATAGGGGAGAGAAAATAGAACTTCTTGTGGACAAGACCGA
GAACCTTCATTCACAGGCACAAGACTTTCGCAGTCAAGGGACACAAATCCGAAGAAAAATGTGGCTGCAGAACATGAAGGTTAAGCTGATTGTATTGGGA
ATACTGATTGCCTTGATCCTCATCATAGTCCTCTCTGTTTGCAAAGGCTTTAATTGTGGAAAATGA
AA sequence
>Potri.012G119600.1 pacid=42782556 polypeptide=Potri.012G119600.1.p locus=Potri.012G119600 ID=Potri.012G119600.1.v4.1 annot-version=v4.1
MNQKSLIYAFVSRGTVILAEFTEFSGNFNSIAFQCLQKLPATNNKFTYNCDGHTFNYLADNGFTYCVVADESAGRQVPMAFLERVKDDFVSKYGGGKAAT
AQANGLNKEFGPKLKEHMKYCADHPEEISKLAKVKAQVSEVKGVMMENIEKVLDRGEKIELLVDKTENLHSQAQDFRSQGTQIRRKMWLQNMKVKLIVLG
ILIALILIIVLSVCKGFNCGK
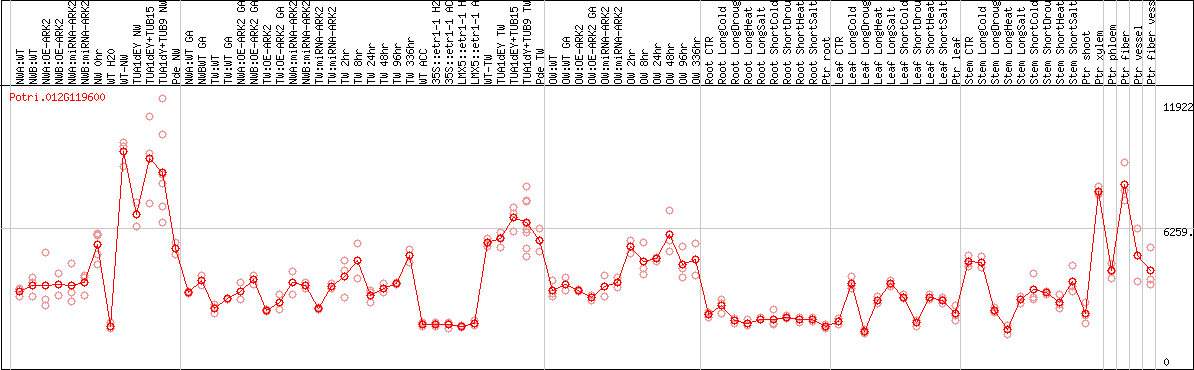
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G119600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.