Potri.012G124200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.012G124200.1 pacid=42782595 polypeptide=Potri.012G124200.1.p locus=Potri.012G124200 ID=Potri.012G124200.1.v4.1 annot-version=v4.1
ATGGCATCAATATCCAAATCATCCCAATTTTCCTTCTTCTCCACCATAACTTTCTTGGTTTACAACCTGCTTGCTTGTGCTACCTTCATAACTTCGATTC
CTGACAGTACTACTAGCGGTGCAGGATTCAAAGAAGCACAGGCCCTTCAGAAATGGAAAGCCAGTCTTGACAATGAAAGCCAATCCCTCCTGAGTTCATG
GAATGGAGACACCCCTTGCAAATGGGTTGGAGTTGATTGTTACCAAGCTGGTGGCATTGCCAATTTAAGTCTTCAGAATGCAGGTTTGAGAGGTACAATT
CATAGTCTTAATTTCTCGTCCTTCCCAAGCTTGATGAAGCTTAACCTCAGTAACAATTCACTCTATGGAACCATTCCTTCACAAATTAGTAACCTTTCCA
GGCTCACCATCCTTGACTTGTCTTACAATGATATTTCAGGAAACATTCCATCTGAAATAAGCTTCTTAAAAAGCCTCAGAATATTTTCACTATCCAATAA
TGATATGAATGGTTCCTTCCCTCCAGAAATAGGAATGATGAGTTCACTTTCCGAAATAAATTTAGAAAACAATCATCTCACGGGTTTTCTCCCTCATTCT
ATAGGAAATATGAGCCACTTATCCAAATTTCTTGTATCTGCTAATAAACTCTTTGGTCCCATACCAGAGGAAGTTGGAACGATGACATCTCTTGCAGTTC
TAGACCTGAACACCAACAGCCTAACCGGTGTTATTCCTAGGTCCATAGGAAACTTAACCAACTTGTTAAAACTTTGCCTTTATGAAAACAAACTCTCTGG
TTCCGTTCCTGAGGAAGTAGGGAACATGAGATCTCTCCTTTACTTTTACCTATGTGATAATAATCTCAGTGGCATGATACCATCTTCTATAGGAAACTTG
ACATCTCTCACTGTCCTAGACCTTGGACCTAATAATCTAACCGGTAAAGTCCCTGCTTCATTAGGAAATTTGAGAAACTTATCCCACCTTTACCTTCCAT
ATAACAACCTTTTTGGTTCCTTGCCTCCAGAAATAAATAACCTTACACATTTGGAACATTTGCAAATATATTCTAATAAATTCACTGGACATCTACCCCG
AGACATGTGCCTTGGGGGATCCCTTTTGTTTTTTGCTGCCTCGGGCAATTATTTTACTGGACCTATCCCAAAAAGTTTGAGAAATTGCACCAGCTTGTTG
AGATTCATGCTTAACAGAAACCAAATTAGTGGAAATATATCTGAAGATTTTGGCATATACCCACATTTATACTACATGGATTTGAGTGATAATGAGCTTT
ATGGTAAACTTTCATGGAAATGGGAGCAGTTCCATAATCTGACAACTCTAAAAATCTCGAGAAATAAAATCTCTGGTGAAATACCGGCTGAGCTAGGAAA
GGCATCTAATCTCAAAGCACTTGATCTCTCTTCAAACCACCTAGTTGGTCAAATTCCAATTGAAGTGGGAAAGTTGAAGTTGCTTGAACTTAAGCTCAGC
AATAACCGACTTTTAGGAGATATTTCTTCAGTGTTTGAAGTGTTGCCTGATGTTAAAAAACTTGACCTAGCAGCAAACAATTTAAGTGGACCAATTCCCA
GACAGATTGGCATGCACTCTCAATTATTATTCTTGAATCTGAGCAAGAATAGTTTCAAAGGGATTATTCCTGCAGAGATAGGGTATTTGCGCTTTCTTCA
AAGTCTTGATCTTAGTTGGAATTCTCTGATGGGAGACCTACCACAGGAGCTCGGAAACCTGCAACGGCTAGAGTCCTTGAACATTTCCCACAACATGCTG
TCTGGTTTCATTCCAACCACTTTTAGTTCTATGCGAGGCATGACAACCGTGGATGTTTCCAATAATAAGTTAGAGGGTCCGATTCCTGACATCAAAGCCT
TCCATGAGGCTCCATTTCAAGCAATTCACAACAATACCAACCTATGTGGCAATGCTACTGGTTTGGAGGTCTGTGAGACACTCCTAGGAAGCAGGACTTT
GCATAGAAAGGGAAAAAAAGTTGTCATTTTGATTACACTCCCTCTGTTGGGATTTTTATTTTTTCTTTTCACCTTGATTGGAGGTTTCTTCATTCTCCGG
CAGAGAATTAGAAGTAGACGAAAAATGTCAATGGAACGCGGAGATCTGTTCTCGATATGGGGCCATCAAGGTGAAATAAATCATGAGGACATCATTGAGG
CAACAGAGGGATTCAACCCCAGCCATTGTATTGGAGCAGGAGGGTTTGCAGCTGTCTACAAAGCTGCCCTGCCAACAGGGCTAGTGGTGGCTGTGAAGAA
ATTTCACCAATCACCAGATGATGAGATGATAGGTTTGAAAGCTTTTACAAGTGAGATGCATTCCTTATTGGGTATACGCCACCGAAACATCGTGAAGCTG
TATGGCTTCTGTTCACATAGAAAACACTCTTTTTTGGTTTATGAATTCCTTGAAAGGGGTAGTTTGAGGACTATTCTAGACAATGAGGAACAAGCAATGG
AGATGGATTGGATGAAAAGGATCAATCTTGTGAGAGGAGTGGCCAATGCTTTGTCCTACCTGCACCACAATTGCTCACCTCCGATCGTTCATCGAGATAT
TTCAAGCAACAATATTCTCTTGGATTCAGAATATGAGGCTCATGTGTCCGACTTTGGCACAGCTAGGCTATTATTGCCTGACTCATCTAATTGGACCTCC
CTTGCAGGTACCGCTGGGTACACAGCTCCAGAGTTGGCCTACACAATGGAAGTGAACGAAAAATGTGATGTCTATAGCTTTGGAGTGGTAGCTATGGAAA
TCATGATGGGAAGGCATCCAGGCGATTTCATTTCATCCTTGTTGTCATCAGCATCCTCATCAACAACAGCGGCAACTTCTCAGAATACGCTCTTCAAGGA
CATATTAGACCAGCGCCTCCCGCCTCCTGAACATCGGGTTGTAGCAGGTGTGGTCTATATTGCAGAACTAGCATTTGCTTGCTTGAATGCTGTTCCCAAA
TCTAGGCCATCGATGAAACAAGTTGCTTCGGATTTCTTAATTCGATGGCCCCCATTGTCATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.012G124200.1 pacid=42782595 polypeptide=Potri.012G124200.1.p locus=Potri.012G124200 ID=Potri.012G124200.1.v4.1 annot-version=v4.1
MASISKSSQFSFFSTITFLVYNLLACATFITSIPDSTTSGAGFKEAQALQKWKASLDNESQSLLSSWNGDTPCKWVGVDCYQAGGIANLSLQNAGLRGTI
HSLNFSSFPSLMKLNLSNNSLYGTIPSQISNLSRLTILDLSYNDISGNIPSEISFLKSLRIFSLSNNDMNGSFPPEIGMMSSLSEINLENNHLTGFLPHS
IGNMSHLSKFLVSANKLFGPIPEEVGTMTSLAVLDLNTNSLTGVIPRSIGNLTNLLKLCLYENKLSGSVPEEVGNMRSLLYFYLCDNNLSGMIPSSIGNL
TSLTVLDLGPNNLTGKVPASLGNLRNLSHLYLPYNNLFGSLPPEINNLTHLEHLQIYSNKFTGHLPRDMCLGGSLLFFAASGNYFTGPIPKSLRNCTSLL
RFMLNRNQISGNISEDFGIYPHLYYMDLSDNELYGKLSWKWEQFHNLTTLKISRNKISGEIPAELGKASNLKALDLSSNHLVGQIPIEVGKLKLLELKLS
NNRLLGDISSVFEVLPDVKKLDLAANNLSGPIPRQIGMHSQLLFLNLSKNSFKGIIPAEIGYLRFLQSLDLSWNSLMGDLPQELGNLQRLESLNISHNML
SGFIPTTFSSMRGMTTVDVSNNKLEGPIPDIKAFHEAPFQAIHNNTNLCGNATGLEVCETLLGSRTLHRKGKKVVILITLPLLGFLFFLFTLIGGFFILR
QRIRSRRKMSMERGDLFSIWGHQGEINHEDIIEATEGFNPSHCIGAGGFAAVYKAALPTGLVVAVKKFHQSPDDEMIGLKAFTSEMHSLLGIRHRNIVKL
YGFCSHRKHSFLVYEFLERGSLRTILDNEEQAMEMDWMKRINLVRGVANALSYLHHNCSPPIVHRDISSNNILLDSEYEAHVSDFGTARLLLPDSSNWTS
LAGTAGYTAPELAYTMEVNEKCDVYSFGVVAMEIMMGRHPGDFISSLLSSASSSTTAATSQNTLFKDILDQRLPPPEHRVVAGVVYIAELAFACLNAVPK
SRPSMKQVASDFLIRWPPLS
|
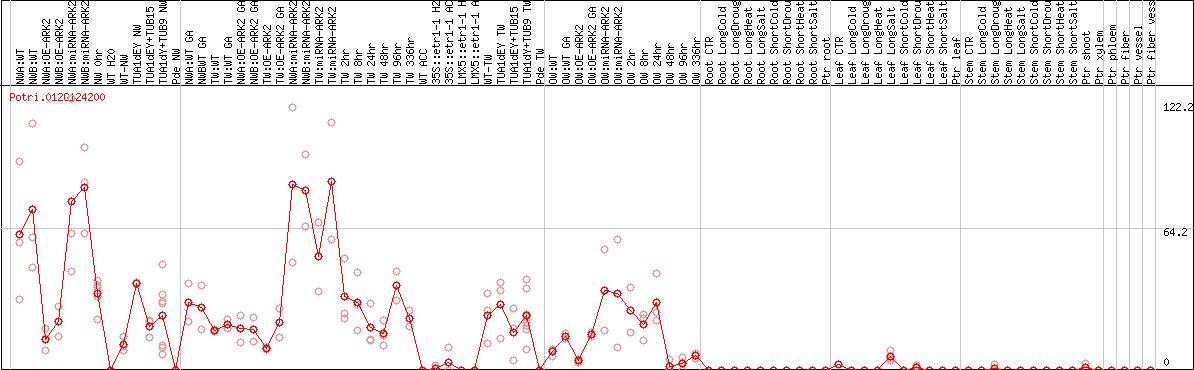
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G124200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
