External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G62165 218 / 3e-72
MADS
FYF, AGL42
FOREVER YOUNG FLOWER, AGAMOUS-like 42 (.1.2.3.4.5)
AT2G45660 200 / 4e-65
MADS
ATSOC1, SOC1, AGL20
SUPPRESSOR OF OVEREXPRESSION OF CO 1, AGAMOUS-like 20 (.1)
AT5G51870 191 / 2e-61
MADS
AGL71
AGAMOUS-like 71 (.1.2.3)
AT5G51860 182 / 2e-58
MADS
AGL72
AGAMOUS-like 72, K-box region and MADS-box transcription factor family protein (.1.2)
AT4G22950 182 / 6e-58
MADS
AGL19
AGAMOUS-like 19 (.1)
AT4G11880 173 / 3e-54
MADS
AGL14
AGAMOUS-like 14 (.1)
AT1G69120 152 / 1e-45
MADS
AGL7, AP1
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
AT5G60910 149 / 8e-45
MADS
FUL, AGL8
FRUITFULL, AGAMOUS-like 8 (.1.2)
AT4G09960 145 / 2e-43
MADS
AGL11, STK
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
AT1G24260 145 / 4e-43
MADS
AGL9, SEP3
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G074200
226 / 4e-75
AT2G45660 242 / 3e-81
SUPPRESSOR OF OVEREXPRESSION OF CO 1, AGAMOUS-like 20 (.1)
Potri.002G151700
222 / 1e-73
AT2G45660 212 / 8e-70
SUPPRESSOR OF OVEREXPRESSION OF CO 1, AGAMOUS-like 20 (.1)
Potri.001G112400
206 / 3e-67
AT4G22950 253 / 6e-86
AGAMOUS-like 19 (.1)
Potri.003G119700
204 / 2e-66
AT4G11880 226 / 4e-75
AGAMOUS-like 14 (.1)
Potri.004G115500
149 / 8e-45
AT3G02310 357 / 4e-126
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Potri.019G077200
147 / 2e-44
AT4G09960 274 / 5e-94
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
Potri.010G154100
148 / 4e-44
AT1G69120 324 / 7e-113
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Potri.013G104900
147 / 4e-44
AT4G09960 330 / 3e-116
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
Potri.017G099700
147 / 8e-44
AT3G02310 372 / 8e-132
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006715
185 / 1e-58
AT4G22950 223 / 1e-73
AGAMOUS-like 19 (.1)
Lus10014143
173 / 7e-54
AT4G22950 212 / 6e-69
AGAMOUS-like 19 (.1)
Lus10031665
155 / 3e-48
AT5G62165 179 / 8e-58
FOREVER YOUNG FLOWER, AGAMOUS-like 42 (.1.2.3.4.5)
Lus10026679
145 / 2e-43
AT1G69120 338 / 1e-118
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10005080
145 / 7e-43
AT3G02310 372 / 1e-131
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Lus10034663
145 / 7e-43
AT5G15800 267 / 4e-90
SEPALLATA1, AGAMOUS-like 2, K-box region and MADS-box transcription factor family protein (.1.2)
Lus10008264
145 / 1e-42
AT4G09960 328 / 3e-114
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
Lus10004637
143 / 1e-42
AT1G69120 318 / 2e-111
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10005302
143 / 6e-42
AT4G09960 330 / 2e-115
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
Lus10003125
139 / 2e-41
AT1G24260 290 / 9e-101
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00319
SRF-TF
SRF-type transcription factor (DNA-binding and dimerisation domain)
PF01486
K-box
K-box region
Representative CDS sequence
>Potri.012G133000.6 pacid=42782553 polypeptide=Potri.012G133000.6.p locus=Potri.012G133000 ID=Potri.012G133000.6.v4.1 annot-version=v4.1
ATGGCGAGAGGAAAGGTTCAGCTGAAAAGGATAGAGAATGCAACTAGCAGGCAAGTGACCTTCTCCAAGAGAAAAAATGGGTTATTGAAGAAAGCTTATG
AGCTATCAATTCTGTGCGATGCTGAAGTTGCAGTGATCATGTTTTCACAGAAAGGAACACTCTTTAAGTTTGCAAGCATTGATCAGATACAAAAGACGAT
TGATCGGTACCGTAAAAATGCAAAGCAATTGCACACTGACAGGATTGATGTGGAACAATCTAAGGAGCAATTAAGACAAGAATCAGCAAACATGGCCAAG
AAGATTGAGATGATCGAGATTTTGCAACGAAAGCTTTTAGGGCAAGATTTAGATTCATGTTCTCCCGAAGAGCTCCATGACATTGACAATCAGCTTGAGA
TCAGTTTAAGCAATATCAGGGCTAGAAAGACTCAGTTATTCAAGGAGCAGATAGAACAGCTGCAAGCAAAGGAAAGATTGTTGTTAATGGAGAATGCAAG
GTTAACTAAACAGTGTGATGCACAGCCATTGCAGCAATCAACTCAATCGAACCAAGTGGTGTCATACTTGACCTCATGTAGCAAGAGTTCAGATATCGTG
GAGACTGATCTGTACATTGGACTGCCACACATGCGCTGCTTGTAG
AA sequence
>Potri.012G133000.6 pacid=42782553 polypeptide=Potri.012G133000.6.p locus=Potri.012G133000 ID=Potri.012G133000.6.v4.1 annot-version=v4.1
MARGKVQLKRIENATSRQVTFSKRKNGLLKKAYELSILCDAEVAVIMFSQKGTLFKFASIDQIQKTIDRYRKNAKQLHTDRIDVEQSKEQLRQESANMAK
KIEMIEILQRKLLGQDLDSCSPEELHDIDNQLEISLSNIRARKTQLFKEQIEQLQAKERLLLMENARLTKQCDAQPLQQSTQSNQVVSYLTSCSKSSDIV
ETDLYIGLPHMRCL
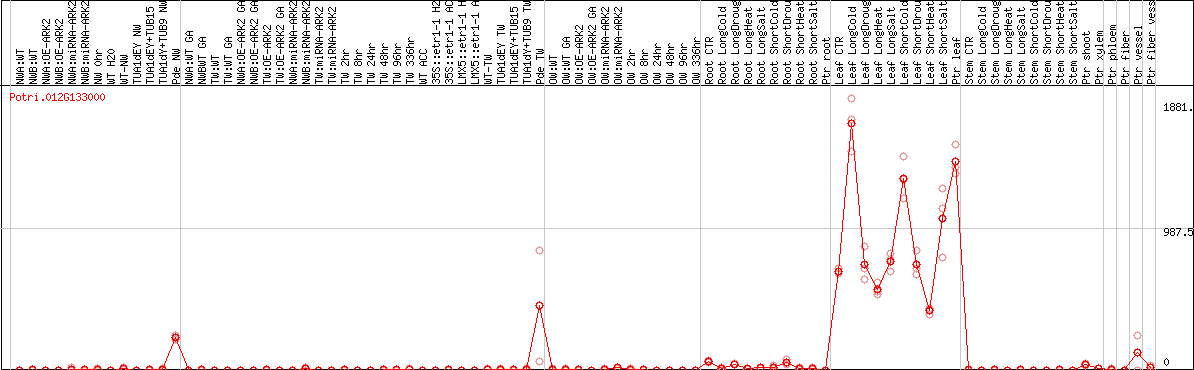
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G133000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.