External link
Symbol
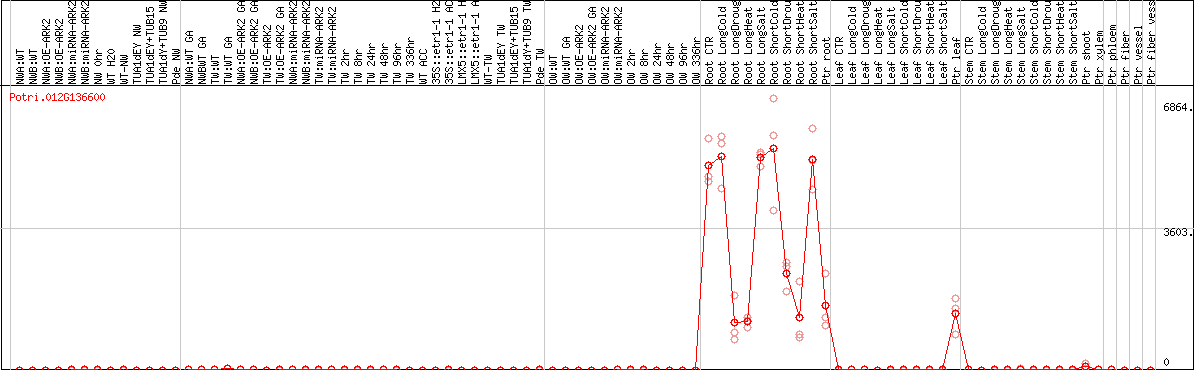
CYP80D1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G45560 360 / 4e-119
CYP76C1
"cytochrome P450, family 76, subfamily C, polypeptide 1", cytochrome P450, family 76, subfamily C, polypeptide 1 (.1.2)
AT1G33720 358 / 9e-119
CYP76C6
"cytochrome P450, family 76, subfamily C, polypeptide 6", cytochrome P450, family 76, subfamily C, polypeptide 6 (.1)
AT3G61040 357 / 2e-118
CYP76C7
"cytochrome P450, family 76, subfamily C, polypeptide 7", cytochrome P450, family 76, subfamily C, polypeptide 7 (.1.2)
AT2G45550 352 / 4e-116
CYP76C4
"cytochrome P450, family 76, subfamily C, polypeptide 4", cytochrome P450, family 76, subfamily C, polypeptide 4 (.1)
AT2G45570 350 / 3e-115
CYP76C2
"cytochrome P450, family 76, subfamily C, polypeptide 2", cytochrome P450, family 76, subfamily C, polypeptide 2 (.1)
AT5G07990 344 / 5e-113
CYP75B1, D501, TT7
TRANSPARENT TESTA 7, CYTOCHROME P450 75B1, Cytochrome P450 superfamily protein (.1)
AT3G52970 339 / 5e-111
CYP76G1
"cytochrome P450, family 76, subfamily G, polypeptide 1", cytochrome P450, family 76, subfamily G, polypeptide 1 (.1.2)
AT2G45580 315 / 6e-102
CYP76C3
"cytochrome P450, family 76, subfamily C, polypeptide 3", cytochrome P450, family 76, subfamily C, polypeptide 3 (.1)
AT4G12300 293 / 2e-93
CYP706A4
"cytochrome P450, family 706, subfamily A, polypeptide 4", cytochrome P450, family 706, subfamily A, polypeptide 4 (.1)
AT4G12320 281 / 9e-89
CYP706A6
"cytochrome P450, family 706, subfamily A, polypeptide 6", cytochrome P450, family 706, subfamily A, polypeptide 6 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G136800
968 / 0
AT2G45560 370 / 2e-123
"cytochrome P450, family 76, subfamily C, polypeptide 1", cytochrome P450, family 76, subfamily C, polypeptide 1 (.1.2)
Potri.015G138900
518 / 0
AT1G33720 359 / 6e-119
"cytochrome P450, family 76, subfamily C, polypeptide 6", cytochrome P450, family 76, subfamily C, polypeptide 6 (.1)
Potri.001G109300
513 / 3e-179
AT1G33720 363 / 1e-120
"cytochrome P450, family 76, subfamily C, polypeptide 6", cytochrome P450, family 76, subfamily C, polypeptide 6 (.1)
Potri.012G137200
493 / 1e-171
AT2G45550 374 / 1e-124
"cytochrome P450, family 76, subfamily C, polypeptide 4", cytochrome P450, family 76, subfamily C, polypeptide 4 (.1)
Potri.001G025200
463 / 8e-160
AT2G45550 322 / 2e-104
"cytochrome P450, family 76, subfamily C, polypeptide 4", cytochrome P450, family 76, subfamily C, polypeptide 4 (.1)
Potri.002G150100
374 / 9e-125
AT2G45550 510 / 3e-178
"cytochrome P450, family 76, subfamily C, polypeptide 4", cytochrome P450, family 76, subfamily C, polypeptide 4 (.1)
Potri.003G118200
369 / 7e-123
AT2G45550 438 / 9e-150
"cytochrome P450, family 76, subfamily C, polypeptide 4", cytochrome P450, family 76, subfamily C, polypeptide 4 (.1)
Potri.002G150200
354 / 3e-117
AT2G45550 496 / 1e-172
"cytochrome P450, family 76, subfamily C, polypeptide 4", cytochrome P450, family 76, subfamily C, polypeptide 4 (.1)
Potri.002G150400
353 / 5e-117
AT2G45550 495 / 2e-172
"cytochrome P450, family 76, subfamily C, polypeptide 4", cytochrome P450, family 76, subfamily C, polypeptide 4 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005765
471 / 5e-163
AT2G45550 374 / 9e-125
"cytochrome P450, family 76, subfamily C, polypeptide 4", cytochrome P450, family 76, subfamily C, polypeptide 4 (.1)
Lus10006323
464 / 4e-160
AT1G33720 371 / 1e-123
"cytochrome P450, family 76, subfamily C, polypeptide 6", cytochrome P450, family 76, subfamily C, polypeptide 6 (.1)
Lus10027431
464 / 6e-160
AT2G45560 379 / 1e-126
"cytochrome P450, family 76, subfamily C, polypeptide 1", cytochrome P450, family 76, subfamily C, polypeptide 1 (.1.2)
Lus10027425
427 / 1e-145
AT3G61040 352 / 1e-116
"cytochrome P450, family 76, subfamily C, polypeptide 7", cytochrome P450, family 76, subfamily C, polypeptide 7 (.1.2)
Lus10027428
420 / 5e-143
AT2G45560 360 / 2e-119
"cytochrome P450, family 76, subfamily C, polypeptide 1", cytochrome P450, family 76, subfamily C, polypeptide 1 (.1.2)
Lus10005766
407 / 9e-138
AT3G61040 352 / 2e-116
"cytochrome P450, family 76, subfamily C, polypeptide 7", cytochrome P450, family 76, subfamily C, polypeptide 7 (.1.2)
Lus10005767
392 / 7e-132
AT2G45550 338 / 1e-110
"cytochrome P450, family 76, subfamily C, polypeptide 4", cytochrome P450, family 76, subfamily C, polypeptide 4 (.1)
Lus10027426
382 / 7e-128
AT2G45570 334 / 2e-109
"cytochrome P450, family 76, subfamily C, polypeptide 2", cytochrome P450, family 76, subfamily C, polypeptide 2 (.1)
Lus10040353
378 / 1e-126
AT2G45570 350 / 1e-115
"cytochrome P450, family 76, subfamily C, polypeptide 2", cytochrome P450, family 76, subfamily C, polypeptide 2 (.1)
Lus10027429
373 / 2e-124
AT2G45570 351 / 6e-116
"cytochrome P450, family 76, subfamily C, polypeptide 2", cytochrome P450, family 76, subfamily C, polypeptide 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00067
p450
Cytochrome P450
Representative CDS sequence
>Potri.012G136600.1 pacid=42782801 polypeptide=Potri.012G136600.1.p locus=Potri.012G136600 ID=Potri.012G136600.1.v4.1 annot-version=v4.1
ATGGTTTCAATAAGTGTTTTAGCAAACTCCTATCCATCTTTTCCCATGTTGTTTCTTCTAGCAATATTATTGTTGCTTTCCTTTGTCCTAAAACACAAAT
CTTCCAAGGTCCCGGCCATTCCACCAGGGCCAAAATCATGGCCTATTATAGGAAATGTACTTCAAATGGGAAACAAGCCCCATATTTCCTTGACTAAATT
AGCACAGGTTTATGGTCCACTCATGTCCCTCCGACTAGGGACTCAACTTGTTGTGGTAGGGTCTTCTCGGGAAGCAGCCTCCGAAATCCTCAAGACTCAT
GACCGAGAACTCTCCGGCCGGTGTGTGCCTCACGCATCCTTTGCTAAGGACCCTAAGCTGAATGAGGACTCCATAGCGTGGACGTTTGAGTGCACCGACC
GATGGAGGTTCTTTCGTTCCCTGATGAGGAATGAGCTATTCTCTTCTAAAGTCGTTGATGGACAGTCAAGAACAAGAGAAACCAAGGCCAGAGAAATGAT
AGATTTCTTGAAAAAGAAGGAAGGAGAGGGAGTCAAAATACGAGATATTGTGTTTGTTTATACATTTAATGCATTGGCGAACATTTACTTATCAAAAGAC
TTGGTAGATTATGATCAAATTGGAGAGTGTCAAAGAGTGTGTGGGCTTGTAAGGGAAATGATGGAGTTACATACAACTCTAAACATATCGGATTTGTATC
CTATACTAGGTAGTTTGGATCTTCAAGGTGTGAGTAGAAAGTGTAATGAGTGCGAGAGTAGAATTCAAGAATTATGGGGGAGTGTCATCAAAGAACGCAG
AGAAGGCCGCAACGATACAGGAGATGATGATGATAATTCCAGTAAACGTAAAGATTTTCTTGATGTTTTGGTTGATGGGGAATTTTCTGATGAGCAAATC
AGTTTATTCTTCGTGGAACTCTTAGCTGCAGTATCGGATAGTACTAGCTCAACAGTCGACTGGGCGATGGCAGAACTGATGAGGAATCCACAAGCAATGA
AACAACTTCGCGAAGAACTTGCTGGAGAAACTCCTGAAGACTTGATAACAGAATCAAGCCTAGCAAAATTCCCTTACTTGCATTTTTGTGTCAAGGAAAC
TTTGAGATTACACCCTCCAGCACCATTTCTAATACCTCACCGAGCAACAGAAGATTGTCAAGTTTTGGACTACACCATACCAAAGGACACCCAAGTTCTT
GTGAATGTATGGGCAATTGCAAGAGACCCTGCCAGTTGGGAAGACCCTTTGTGTTTTAAACCTGAAAGATTCCTGAATTCTGATCTGGATTACAAAGGAA
ACCATTTTGAGTTTTTGCCTTTTGGAAGTGGAAGGAGGATTTGTGCCGGGTTACCAATGGCTGTTAAGAAAGTTCAGCTGGCTTTGGCCAATTTAATCCA
TGGCTTTGACTGGTCTCTACCAAATAACATGCTTCCTGATGAACTAAACATGGATGAGAAGTATGGTATCACACTGATGAAGGAGCAACCTCTTAAGCTT
ATACCGAAATTGAGGAAATGA
AA sequence
>Potri.012G136600.1 pacid=42782801 polypeptide=Potri.012G136600.1.p locus=Potri.012G136600 ID=Potri.012G136600.1.v4.1 annot-version=v4.1
MVSISVLANSYPSFPMLFLLAILLLLSFVLKHKSSKVPAIPPGPKSWPIIGNVLQMGNKPHISLTKLAQVYGPLMSLRLGTQLVVVGSSREAASEILKTH
DRELSGRCVPHASFAKDPKLNEDSIAWTFECTDRWRFFRSLMRNELFSSKVVDGQSRTRETKAREMIDFLKKKEGEGVKIRDIVFVYTFNALANIYLSKD
LVDYDQIGECQRVCGLVREMMELHTTLNISDLYPILGSLDLQGVSRKCNECESRIQELWGSVIKERREGRNDTGDDDDNSSKRKDFLDVLVDGEFSDEQI
SLFFVELLAAVSDSTSSTVDWAMAELMRNPQAMKQLREELAGETPEDLITESSLAKFPYLHFCVKETLRLHPPAPFLIPHRATEDCQVLDYTIPKDTQVL
VNVWAIARDPASWEDPLCFKPERFLNSDLDYKGNHFEFLPFGSGRRICAGLPMAVKKVQLALANLIHGFDWSLPNNMLPDELNMDEKYGITLMKEQPLKL
IPKLRK