External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G73040 216 / 3e-72
Mannose-binding lectin superfamily protein (.1)
AT1G19715 110 / 3e-28
Mannose-binding lectin superfamily protein (.1.2.3)
AT3G16460 86 / 2e-19
JAL34
jacalin-related lectin 34, Mannose-binding lectin superfamily protein (.1.2)
AT1G52060 81 / 2e-18
Mannose-binding lectin superfamily protein (.1)
AT3G16420 79 / 1e-17
JAL30, PBP1
JACALIN-RELATED LECTIN 30, PYK10-binding protein 1 (.1.2.3)
AT2G25980 79 / 2e-17
Mannose-binding lectin superfamily protein (.1)
AT1G52070 79 / 2e-17
Mannose-binding lectin superfamily protein (.1)
AT3G16430 78 / 3e-17
JAL31
jacalin-related lectin 31 (.1.2)
AT3G16440 77 / 6e-17
ATMLP-300B, MEE36
maternal effect embryo arrest 36, myrosinase-binding protein-like protein-300B (.1)
AT3G16470 77 / 6e-17
JAL35, JR1
JASMONATE RESPONSIVE 1, jacalin-related lectin 35, Mannose-binding lectin superfamily protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G232400
135 / 5e-37
AT1G19715 609 / 0.0
Mannose-binding lectin superfamily protein (.1.2.3)
Potri.005G232500
131 / 8e-36
AT1G19715 486 / 2e-166
Mannose-binding lectin superfamily protein (.1.2.3)
Potri.002G030400
129 / 1e-34
AT1G19715 590 / 0.0
Mannose-binding lectin superfamily protein (.1.2.3)
Potri.017G010900
73 / 4e-15
AT3G16460 135 / 5e-34
jacalin-related lectin 34, Mannose-binding lectin superfamily protein (.1.2)
Potri.010G144900
47 / 7e-07
AT1G05760 92 / 5e-24
restricted tev movement 1, Mannose-binding lectin superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037605
195 / 1e-61
AT1G73040 198 / 1e-62
Mannose-binding lectin superfamily protein (.1)
Lus10006866
195 / 2e-58
AT1G73040 194 / 5e-58
Mannose-binding lectin superfamily protein (.1)
Lus10024290
134 / 1e-36
AT1G19715 607 / 0.0
Mannose-binding lectin superfamily protein (.1.2.3)
Lus10024291
130 / 4e-35
AT1G19715 438 / 1e-146
Mannose-binding lectin superfamily protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0568
Man_lectin
PF01419
Jacalin
Jacalin-like lectin domain
Representative CDS sequence
>Potri.012G140001.1 pacid=42782980 polypeptide=Potri.012G140001.1.p locus=Potri.012G140001 ID=Potri.012G140001.1.v4.1 annot-version=v4.1
ATGGAAAATGGGAAGGAACAATCAGCTGCGAGAAAGAAGAGCACCATATTAGTTGGACCATGGGGAGGGAATGGAGGGGATAGCTGGGATGATGGGATCT
ACCATGGAGTAAGAGAAATCACAATTGTTTATGATCAATGCATCGACTCGATCCAGGTTGTGTATGACAAGAATGGCAAGCCTATCACAGCAGAAAACCA
TGGAGGTGTTGGAGGCAGTAGAACAGCTGAGATTAAGTTGCAATATCCGGAGGAGTACTTAACCAGTGTGAGTGGCCATTACTGCCCGGTAGTCTATGGT
GGCAGTCCTGTGATTCGATCGTTAGCATTCAGCAGCAACAAAAGAACATTTGGACCATTTGGAGTTGAAGAAGGAACGCCATTTACACTTTCAATGGACG
GAGCATCGATTGTAGGCTTTAAGGGTAGAGGTGGATGGTATCTTGATGCCATTGGGTTTCGTTTATCTCGTATTCAATCCACCAAAGTTCTCAAAAAATT
TCAACAAAAGCTTCAAAGGCTCACCAGTACGGTTTCAAAGTCCTCTGCCTCCAAGGATGCTGAAAAAACCTATTGA
AA sequence
>Potri.012G140001.1 pacid=42782980 polypeptide=Potri.012G140001.1.p locus=Potri.012G140001 ID=Potri.012G140001.1.v4.1 annot-version=v4.1
MENGKEQSAARKKSTILVGPWGGNGGDSWDDGIYHGVREITIVYDQCIDSIQVVYDKNGKPITAENHGGVGGSRTAEIKLQYPEEYLTSVSGHYCPVVYG
GSPVIRSLAFSSNKRTFGPFGVEEGTPFTLSMDGASIVGFKGRGGWYLDAIGFRLSRIQSTKVLKKFQQKLQRLTSTVSKSSASKDAEKTY
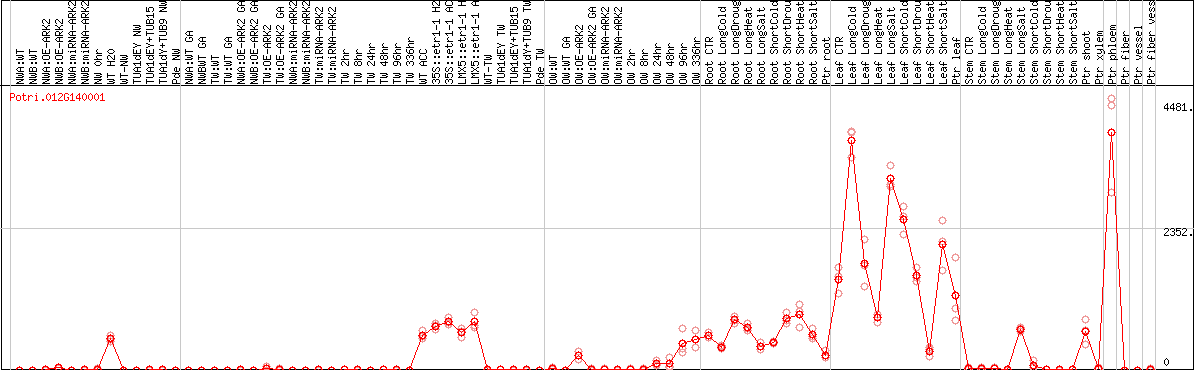
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G140001 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.