External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G23950 166 / 1e-50
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT1G07310 110 / 7e-28
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT4G01200 60 / 2e-10
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT2G13350 52 / 1e-07
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT1G04540 48 / 3e-06
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT3G04360 44 / 5e-05
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT3G16510 41 / 0.0005
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G249300
118 / 2e-31
AT1G07310 224 / 1e-71
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.008G064900
113 / 2e-29
AT1G07310 150 / 1e-42
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.009G043300
106 / 6e-27
AT1G07310 205 / 9e-64
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.008G172300
65 / 1e-11
AT2G33320 265 / 8e-82
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.013G048400
63 / 3e-11
AT2G33320 211 / 6e-63
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.010G065300
63 / 4e-11
AT2G13350 261 / 6e-83
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.019G021300
60 / 3e-10
AT3G04360 213 / 2e-65
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.019G065500
52 / 1e-07
AT2G40815 213 / 1e-66
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.002G166400
48 / 3e-06
AT4G01200 211 / 4e-68
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039259
214 / 2e-68
AT5G23950 160 / 3e-48
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10027502
201 / 2e-63
AT5G23950 164 / 8e-50
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10040692
115 / 6e-30
AT1G07310 189 / 2e-57
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10005867
112 / 6e-29
AT1G07310 206 / 3e-64
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10018213
73 / 4e-15
AT1G07310 94 / 1e-22
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10003570
64 / 3e-11
AT2G33320 222 / 7e-67
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10032780
63 / 5e-11
AT2G33320 227 / 1e-70
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10014466
58 / 2e-09
AT1G04540 245 / 7e-76
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10008731
45 / 3e-05
AT4G01200 222 / 3e-72
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10002155
44 / 7e-05
AT4G01200 224 / 7e-73
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0154
C2
PF00168
C2
C2 domain
Representative CDS sequence
>Potri.012G144600.2 pacid=42783791 polypeptide=Potri.012G144600.2.p locus=Potri.012G144600 ID=Potri.012G144600.2.v4.1 annot-version=v4.1
ATGGCTTCTCGATACGAGTTTGAAGTCACCATCTCTTCTGCCAAATACCTCAAAAACGTTAACTGGCGCCATGGCTCTTTGAAGCCATATGCTGTCGTGT
GGGTTGACTCCAACTACAAATGCTCCACTCAAGTCGACGACGAAGGCGACACCTCTCCTTTTTGGGACCAAACTCTGGTCATCCCCTTACCATCTGGTCG
CATCGAAGACCACACCCTCCACATCGACATCGTCCACGCCGGCTCCGAGGAGGGCACCAAGCCCCTCATCGGCTCCGCTAAGCTCAAGCTCATTGACGTT
CTCGACGACGTCGAAATCGGAGAACGAGCCACCCGCGCTCTCCAGCTCAAGCGTCCTTCCGGCAGACCACAGGGGAAACTCGACGTTAAGGTCACCATTA
GAGACCCTCGCTATCGTGCTCCTGATGCTTACCGTGAACCTCCTTACGGGGTTCCTCCTCCGTCATCTTCGAGAGACTACCCTTACGCGACACCCTATGG
TGCACCACCACCGCAGCCAAATCCGTACTATTCAACAGCAGCTCCGCCAACAGGGTATCCGTACAGCGCGCCCCCACCTCCGACATACGGGCAGCCGAGT
TACGGTTATGGGGGTTACGGGTATGGAGAGCAGCAACCTGTGGTTGTGGAGGAGAAGAAGAAAAGTAAGTTTGGAGGGATGGGGACAGGATTGGCCGTGG
GTGCGGTGGGTGGGGTCTTAGGTGGACTGGCATTGGCTGAGGCCATTGATGCTTTGGAAGACCATGTTGCTGATGACGTGGCAGAGAAAGTCGAGGATGA
TCTTGCCTTCGATATGATGCTTTCTGATCTCTAA
AA sequence
>Potri.012G144600.2 pacid=42783791 polypeptide=Potri.012G144600.2.p locus=Potri.012G144600 ID=Potri.012G144600.2.v4.1 annot-version=v4.1
MASRYEFEVTISSAKYLKNVNWRHGSLKPYAVVWVDSNYKCSTQVDDEGDTSPFWDQTLVIPLPSGRIEDHTLHIDIVHAGSEEGTKPLIGSAKLKLIDV
LDDVEIGERATRALQLKRPSGRPQGKLDVKVTIRDPRYRAPDAYREPPYGVPPPSSSRDYPYATPYGAPPPQPNPYYSTAAPPTGYPYSAPPPPTYGQPS
YGYGGYGYGEQQPVVVEEKKKSKFGGMGTGLAVGAVGGVLGGLALAEAIDALEDHVADDVAEKVEDDLAFDMMLSDL
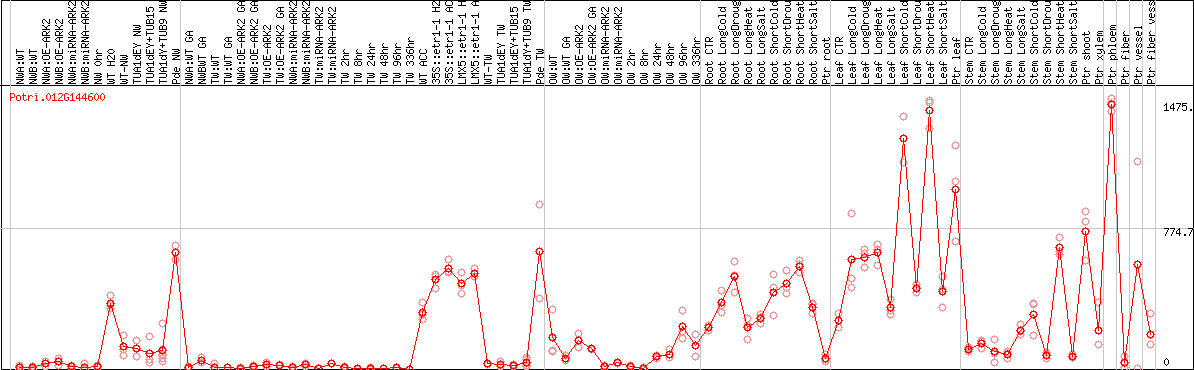
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.012G144600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.