PHYC.1,phya (Potri.013G000300) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | PHYC.1,phya | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.013G000300.2 pacid=42811827 polypeptide=Potri.013G000300.2.p locus=Potri.013G000300 ID=Potri.013G000300.2.v4.1 annot-version=v4.1
ATGTCTTCTTCAAGGCCAAGCCACTCATCCAGCAATTCAGCAAGATCAAGACACAGTGCTAGGATTATTGCTCAGACCACTGTGGATGCCAAGCTCCATG
CTGATTTTGAGGAGTCAGGCAGCTCTTTTGATTACTCAAGCTCAGTGCGTGTTACTGACTCAGTTGGTGGAGATCAGCCACCCAGGTCTGACAAAGTAAC
TACAACTTATCTTCACCACATCCAGAAAGGCAAGCTGATCCAGCCTTTTGGATGCTTGCTAGCCTTAGATGAGAAAACTTTCAAAGTCGTTGCATACAGT
GAGAATGCCCCTGAACTGTTGACTATGGTCAGTCATGCAGTGCCAAGTGTTGGGGAACATCCTGTTCTTGGCATTGGAACTGACATCAGGACTATCTTCA
CTGCTCCTAGTGCATCTGCCTTGCAAAAAGCCATGGGATTCGGGGATGTTTCTCTTTTGAATCCTATATTGGTCCATTGCAAGACTTCTGGGAAGCCGTT
TTATGCCATTGTTCACCGGGTGACTGGCAGTTTGATCATTGACTTTGAACCTGTGAAGCCTTACGAGGTTCCCATGACTGCTGCTGGTGCTCTGCAATCA
TACAAGCTTGCAGCCAAAGCGATTACTCGGCTGCAGTCTTTGCCTAGTGGAAGCATGGAAAGGCTTTGTGATACAATGGTTCAGGAGGTTTTTGAGCTCA
CTGGATATGATAGGGCGATGGCCTATAAATTTCATGATGATGATCATGGAGAAGTGGTCTCTGAGGTTACAAAGCCAGGCATGGAGCCATATTTGGGTTT
GCACTATCCAGCCACTGATATCCCTCAGGCTTCACGCTTTCTATTTATGAAGAATAAAGTTCGAATGATTGTTGATTGTCATGCGAAACATGTCAAGGTG
CTTCAGGATGAGAAGCTTCCATTTGATCTGACATTGTGTGGTTCAACCCTGAGGGCCCCACACAGTTGTCATTTACAATACATGGAGAACATGAACTCCA
TCGCTTCTCTGGTTATGGCAGTTGTAGTCAATGATGGGGATGAAGATGGTGATACCCCTGATTCTGTGAACCCACAGAAAAGAAAGAGACTTTGGGGTTT
AGTAGTGTGTCATAATACAAGTCCGAGGTTTGTTCCTTTCCCTCTTAGGTATGCTTGTGAGTTTCTAGCTCAAGTATTTGCTATCCATGTCAACAAGGAA
TTGGAGTTAGAAAATCAAATTGTCGAGAAGAACATCCTCCGCACCCAGACACTCTTATGTGATATGCTCATGCGTGATGCACCATTGGGTATTGTGACAC
AGAGCCCGAACATTATGGATCTGGTAAAGTGTGATGGTGCTGTCCTGTTCTATAGGAATAAGATATGGAGGTTGGGGATAACTCCAAGTGATCTGCAGCT
TCAGGATATAGCTTTTTGGCTCTCCGAGTACCATATGGATTCTACAGGTTTGAGTACAGACAGCTTGTATGATGCAGGGTACCCAGGGGCTCTGGCTCTT
GGTGATGTAGTTTGTGGAATGGCAGCTGTGAGAATAACTTCCAAGGACATGCTTTTTTGGTTTCGATCCCAGACTGCTGCAGAAATTCGATGGGGTGGTG
CAAAGCATGAACCTGGGGAGAAGGATGATGGAAGGAGAATGCACCCTAGGTCATCTTTCAAGGCCTTCCTTGAAGTTGTCAAGACAAGGAGTTTGCCTTG
GAAGGACTATGAAATGGATGCAATCCATTCTCTGCAGCTTATCCTGAGGAACGCATTCAAAGACATTGAAACCATGGATGTGGATACCAAAACAATTCAT
GCAAGGCTAAGCGACCTCAAAATTGAAGGGATGCAAGAACTTGAAGCAGTTACTAGTGAGATGGTCCGTTTAATTGAAACAGCTACAGTGCCAATTTTGG
CAGTTGATGTTGATGGTCTAGTTAACGGGTGGAATACAAAAATTTCTGAATTGACTGGTCTTCTTGTTGATAAAGCAATCGGGAAGCATTTGCTAACACT
TGTGGAAGATTCTTCGGTTGACATAGTCAAAAGAATGTTATTCTTGGCATTGCAGGGCAAAGAAGAGCAAAACATACAATTTGAGATCAAGACACATGGT
TCCAAGTCTGAATGTGGCCCCATCTGCTTAGTTGTTAATGCCTGTGCAAGCAGGGACCTTCATGAAAATGTTGTGGGGGTGTGCTTTGTGGGTCAAGATA
TCACTGGCCAGAAGATGGTCATGGACAAGTTCACCCGGATTGAAGGGGATTACAAAGCAATTGTACAAAACCGAAACCCTTTGATTCCCCCAATATTTGG
GACGGATGAGTTTGGCTGGTGTTCTGAGTGGAATCCTGCCATGACAAATTTGACTGGTTGGAAACGAGAAGAAGTTCTAGATAAAATGTTATTGGGAGAG
GTTTTTGGGTTAAACATGGCATGCTGTCGTCTCAAGAACCAGGAGGCTTTTGTAAATCTTGGTGTTGTTCTTAATACTGCCATGACTGGCCAGGAATCCG
AGAAGGTTTCTTTTGGTTTCTTTGCTCGGACTGGAAAGTATGTGGAATGCCTGCTGTGTGTGAGTAAGAAATTGGACAGAGAGGGTGCAGTCACTGGGGT
ATTCTGCTTTCTACAGCTTGCAAGCCAAGAGCTGCAACAAGCACTTCATGTCCAGCGATTATCAGAGCAAACTGCTTTGAAAAGATTGAAAGCATTGGCT
TATTTAAAAAGGCAGATCTGGAATCCTCTCTCTGGAATTATATTTTCTGGAAAAATGATGGAGGGAACCGAGTTGGGAGCAGAACAAAAGGAGCTTCTGC
ATACCAGTGCCCAGTGCCAGTGCCAACTCAGCAAGATTCTTGATGACTCAGATCTAGATAGCATCATCGAAGGTTACTTGGATCTGGAAATGGTGGAGTT
TACCCTGCGTGAGGTACTAGTGGCAGCTACCAGTCAAGTCATGATGAAGAGCAATGAAAAGGGCATTCGAATAATCAACGATGCAGCTGAAGAGACGATG
GCTGAGACCCTGTATGGTGATAGCATCAGGCTTCAACAGGTGTTAGCTGATTTCTTGCAGATGTCAGTTAATTTTACACCATCTGGAGGCCTGCTTTCTG
TTTCAGCTAGCTTAACCAAAGACCAGTTGGGGCAATCTGTTTATCTTGTGCATCTGGAACTCAGGATAAGGCATCCTGGTGCAGGGATTCCTGAAGCACT
GCTAGATCAAATGTTTGGGGAGGATACAGATGCGTCTGTGGAGGGGATCAGCTTGGTTATAAGCAGGAAGCTGGTGAAGCTGATGAACGGAGATGTTCGG
TACATGAGGGAAGCAGGGAAGTCGAGTTTTATCATATCAGTAGAACTTGCTGGTGGCCATAAATCGCAAAAAAGGGCATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.013G000300.2 pacid=42811827 polypeptide=Potri.013G000300.2.p locus=Potri.013G000300 ID=Potri.013G000300.2.v4.1 annot-version=v4.1
MSSSRPSHSSSNSARSRHSARIIAQTTVDAKLHADFEESGSSFDYSSSVRVTDSVGGDQPPRSDKVTTTYLHHIQKGKLIQPFGCLLALDEKTFKVVAYS
ENAPELLTMVSHAVPSVGEHPVLGIGTDIRTIFTAPSASALQKAMGFGDVSLLNPILVHCKTSGKPFYAIVHRVTGSLIIDFEPVKPYEVPMTAAGALQS
YKLAAKAITRLQSLPSGSMERLCDTMVQEVFELTGYDRAMAYKFHDDDHGEVVSEVTKPGMEPYLGLHYPATDIPQASRFLFMKNKVRMIVDCHAKHVKV
LQDEKLPFDLTLCGSTLRAPHSCHLQYMENMNSIASLVMAVVVNDGDEDGDTPDSVNPQKRKRLWGLVVCHNTSPRFVPFPLRYACEFLAQVFAIHVNKE
LELENQIVEKNILRTQTLLCDMLMRDAPLGIVTQSPNIMDLVKCDGAVLFYRNKIWRLGITPSDLQLQDIAFWLSEYHMDSTGLSTDSLYDAGYPGALAL
GDVVCGMAAVRITSKDMLFWFRSQTAAEIRWGGAKHEPGEKDDGRRMHPRSSFKAFLEVVKTRSLPWKDYEMDAIHSLQLILRNAFKDIETMDVDTKTIH
ARLSDLKIEGMQELEAVTSEMVRLIETATVPILAVDVDGLVNGWNTKISELTGLLVDKAIGKHLLTLVEDSSVDIVKRMLFLALQGKEEQNIQFEIKTHG
SKSECGPICLVVNACASRDLHENVVGVCFVGQDITGQKMVMDKFTRIEGDYKAIVQNRNPLIPPIFGTDEFGWCSEWNPAMTNLTGWKREEVLDKMLLGE
VFGLNMACCRLKNQEAFVNLGVVLNTAMTGQESEKVSFGFFARTGKYVECLLCVSKKLDREGAVTGVFCFLQLASQELQQALHVQRLSEQTALKRLKALA
YLKRQIWNPLSGIIFSGKMMEGTELGAEQKELLHTSAQCQCQLSKILDDSDLDSIIEGYLDLEMVEFTLREVLVAATSQVMMKSNEKGIRIINDAAEETM
AETLYGDSIRLQQVLADFLQMSVNFTPSGGLLSVSASLTKDQLGQSVYLVHLELRIRHPGAGIPEALLDQMFGEDTDASVEGISLVISRKLVKLMNGDVR
YMREAGKSSFIISVELAGGHKSQKRA
|
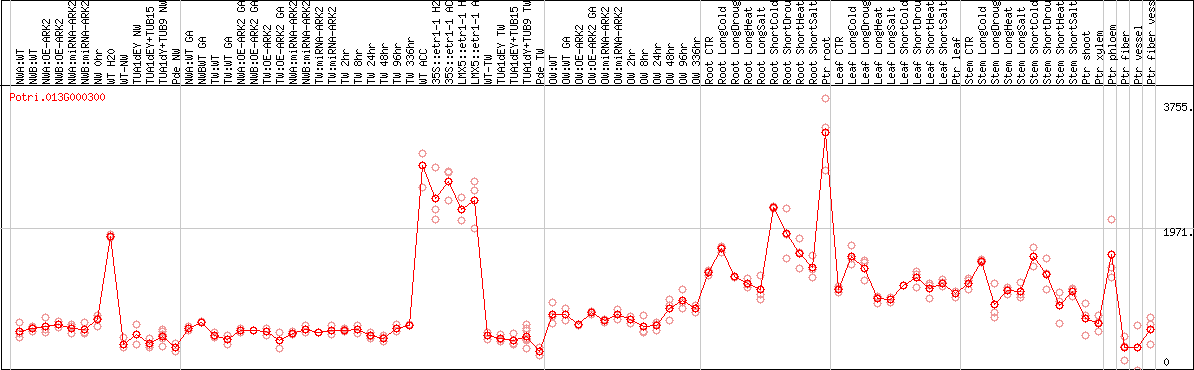
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G000300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
