External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G09070 157 / 7e-46
(AT)SRC2, (AT)SRC2, (AT)SRC2, (AT)SRC2, (AT)SRC2, (AT)SRC2, (AT)SRC2, (AT)SRC2, (AT)SRC2, (AT)SRC2,
soybean gene regulated by cold-2 (.1)
AT4G15755 113 / 1e-29
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT3G16510 113 / 5e-29
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT4G15740 97 / 1e-22
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT3G62780 84 / 9e-19
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT3G05440 45 / 5e-06
C2 domain-containing protein (.1)
AT3G04360 47 / 6e-06
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT5G11100 44 / 0.0001
SYT4, NTMCTYPE2.2, ATSYTD ,NTMC2T2.2 ,NTMC2TYPE2.2
synaptotagmin 4, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT4G01200 41 / 0.0004
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G026700
239 / 3e-78
AT1G09070 148 / 3e-42
soybean gene regulated by cold-2 (.1)
Potri.010G024200
157 / 2e-46
AT3G16510 135 / 5e-37
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.008G209800
134 / 2e-37
AT3G16510 137 / 8e-38
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.001G070400
75 / 4e-15
AT3G16510 82 / 4e-17
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.013G107175
72 / 3e-14
AT3G16510 77 / 1e-15
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.005G026650
66 / 3e-13
AT1G09070 53 / 9e-09
soybean gene regulated by cold-2 (.1)
Potri.003G160400
66 / 6e-12
AT3G16510 72 / 8e-14
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.018G025000
42 / 0.0004
AT5G11100 824 / 0.0
synaptotagmin 4, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.011G088400
40 / 0.001
AT5G55530 418 / 9e-145
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2), Calcium-dependent lipid-binding (CaLB domain) family protein (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029901
187 / 6e-58
AT1G09070 172 / 6e-52
soybean gene regulated by cold-2 (.1)
Lus10004500
184 / 1e-57
AT1G09070 179 / 2e-55
soybean gene regulated by cold-2 (.1)
Lus10006523
119 / 1e-31
AT3G16510 147 / 1e-41
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10005867
45 / 3e-05
AT1G07310 206 / 3e-64
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10040692
41 / 0.0007
AT1G07310 189 / 2e-57
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0154
C2
PF00168
C2
C2 domain
Representative CDS sequence
>Potri.013G018600.1 pacid=42811149 polypeptide=Potri.013G018600.1.p locus=Potri.013G018600 ID=Potri.013G018600.1.v4.1 annot-version=v4.1
ATGGAGTGCAGGCCATTAGAGATCACGGTTGCCTCAGGCAAAGATCTCAAAGATGTGAACGTTTTCGGCAAGATGGATCTCTATTGTGTTGTTTCCATCA
AGGGAGATCCACACAAGTCTAAGCAGAAACAGAAAACCCACGTTCATAAAGATTGTGGGCCTAACCCTCTTTGGAATTTCCCCATGAAATTCAACATCGA
CGAAGCCGCTGCTCAACAAAATCGCTTGCAAATCAAGTTCAAGCTTCTAGCTGAGCGAATGATGGGAGATAAAGAAGTTGGGGTGGTTTCTGTGCCTGTC
AAGGAATTACTGGATTCAAAAGATGGAAAAGGTGGTCTCATGAGCTATGCTGTCAAGACCCCATCTGGGAAGATGAAAGGAACGTTGAGTTTCTCCTTTA
ATTTTGGTGAGAAAGTCAGTGCTCCTGCGCCGGAGAAGGCGAAAAAGACGGGCGAGCATGTTGCTGCTGCTGCTTCTCCTGCAAAGGGATATCATGAGCC
CGTCACTGCCTACCCTGCAACGGGATATCAGGGTGCTCCTGGTTCAAGTTCTGCCTACCCAGCGCCACCGCCTCAAGCGGCTCCTTATCCATATCCTTAT
CCTTATCAGTATCCTCCACCACCCCAACATGGTTATGGTGGATACCCACCTGCACCCGGACATGGGTATCCCGGTTATCCACCACAACCCGGTTATGGTG
GGTATCCACCAGTGATGCAACAACAGAAGCCCAAGAAAAGTGGTAAAGGAAATATGGCTTTGGGATTGGGAGCTGGATTGCTTGGTGGATTGCTGGTTGG
TGATATGATTTCCGATGTTGGTGAGATGGCTGCCTATGACGATGGCTTTTCTGATGGACTTGATTTTTAG
AA sequence
>Potri.013G018600.1 pacid=42811149 polypeptide=Potri.013G018600.1.p locus=Potri.013G018600 ID=Potri.013G018600.1.v4.1 annot-version=v4.1
MECRPLEITVASGKDLKDVNVFGKMDLYCVVSIKGDPHKSKQKQKTHVHKDCGPNPLWNFPMKFNIDEAAAQQNRLQIKFKLLAERMMGDKEVGVVSVPV
KELLDSKDGKGGLMSYAVKTPSGKMKGTLSFSFNFGEKVSAPAPEKAKKTGEHVAAAASPAKGYHEPVTAYPATGYQGAPGSSSAYPAPPPQAAPYPYPY
PYQYPPPPQHGYGGYPPAPGHGYPGYPPQPGYGGYPPVMQQQKPKKSGKGNMALGLGAGLLGGLLVGDMISDVGEMAAYDDGFSDGLDF
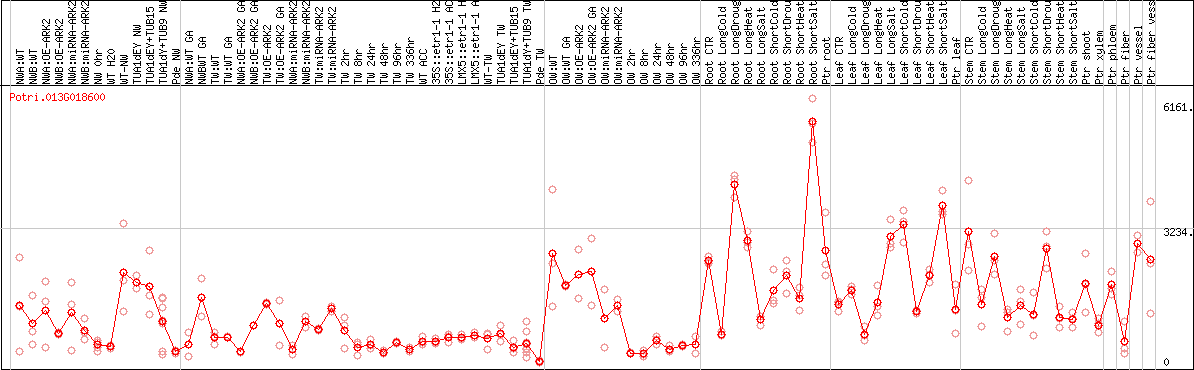
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G018600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.