Potri.013G019000 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.013G019000.4 pacid=42812635 polypeptide=Potri.013G019000.4.p locus=Potri.013G019000 ID=Potri.013G019000.4.v4.1 annot-version=v4.1
ATGGATCACCTAAGATCATCCTCAGCGAATGGTGAGGAAAATGGTGGTGGGATTCCTGATGACTTGCGCTGCAAGAGGTCAGATGGGAAGCAGTGGAGAT
GTACTGCCATGTCTATGCCAGATAAAACTGTTTGTGAGAAGCATTATATCCAGGCAAAGAAGAGGGCAGCTAATTCTGCGTTAAGAGCCAGTCTGAAGAA
AGCCAAAAGGAAATCAATAGGTGAAAGCGATTTTTACTTGGAGAGTAAGAGTGATGATTTTGACATGCCACTCAGGAACATGAAAGTTGAGGAGGATCAG
CCACTTTCGGTCTCATCAAAGAGGTACAAGGAGAAAGTACCTAAAAGTCAGTCTCGCTATTCTCCAGAAACACTTATAAGGAGCTTGCGTGGTCAGAATT
CTCTAAAACTAAACGATGATTCCCAAAGAGATTTCGAGTTTGAAGAGAATTGGAGGTCATATAAGACGACACCTCGTTCAACAATGGAATCATCCAGGAG
TAGATCACAAAGGAGTTTTGATGCCAGTGCTATGACAGTGAGTGAAACTGTCACAGAATACTCAGATGCAAGCACCGATGCTTCTGAAGACACTGGTGGG
CAAACTTGTCATCAGTGCCGGAGGAATGACAGAAATAGTGTTACTTGGTGTCTCAAATGTGACAAGAGAGGTTTTTGTGATAGCTGTATCTCAGAATGGT
ACTCAGACATTCCATTAGAAGAAATTGAGAAGGTTTGTCCTGCATGTCGCGGGATCTGCAATTGCAGGGGTTGCTTACGAGGAGATAATATGGTAAAGGT
AAGAATACGGGAAATACCTGTTCTTGACAAGTTGCAATATCTCCACTGCCTATTATCTTCAGTGCTTCCTATAGTTAAGCAGATCCATCAAGAACAATGC
TTTGAAGTAGAGCTTGAACAAAGGCTGCGTGGAACTGATATAGATCTTGTCCGGGCCAAATTGAATGCAGATGAGCAGATGTGCTGCAATATATGTAGGA
TACCCATTATTGATTATCATCGGCATTGTGCAAATTGCTCGTATGATCTGTGCCTTCATTGCTGTCAAGATCTTCGAGGAGCATCTAAACATGGTGTTGA
AAATGAAGTGGATGATAATCAGATTGATGGGAGAAGTCAAGACAATGAGACTCCATTAGAACCAGTGAGAGAACCACAAGTAAGACTAAAGTTATCAGAT
AAGTACCAAGGCTGGAAGGCAAACAATGATGGCAGTATTCCATGCCCTCCGAAGGAGCACGGTGGTTGTAATTACTCATCATTGAACCTAAGCCGTATTT
TCAAGATGAATTGGGCAGCAAAACTTGTGAAAAATGTGGAGGAAATGGTCAGTGGCTGTAAGGTTTATGATGCTGGCACCCCACAGAAATCCGGGTTGAA
TGATTCCACACTCTGCCAGTATGCTCACAGAGAGGACAGTGATGATAATTTCTTATATTGCCCCTTATCTGAAGATGTAAAAGCTGATGGGATAAACAAG
TTTAGAAAACACTGGGTTCGAGGTGAACCTGTTATTGTGAAGCAGGTATTTGACAGCTCATCCATATCAAGCTGGGATCCAATGGCTATTTGGAGGGGGA
TCCGGGAAACATCAGATGAGAAAAAGAAAGGTGAAAACAGAATGGTGAAGGCCATAGATTGCTTACACTGGTCTGAGGTTGATATTGATCTTGATCAGTT
TATCCGAGGATACTCAGAGGGACGAATCAGGGAAAATGGTTCACCAGAGATGTTGAAGTTGAAGGATTGGCCTTCCCCTAGTGCTTCTGAAGAGTTCTTA
TTATACCAGAGACCTGAATCTATCAGTAAACTGCCTTTCCTTGAATTTATCCATTCAAGGGTGGGTGTTCTAAATGTTGCTGCAAAGTTGCCTCATTACT
CTTTGCAGAATGATGTAGGACCCAAGATTTGTATATCTTATGGGAGCCATGAGGATCTCGGTGTGGGTGATTCTGTAATCAAACTTCATTTCAAAACGCG
TGACATGGTGTATCTATTGGTACATACATGCGAAGCAAAGACAAAAGGTAGCCAGGAGAGTAGCTCAATTGACCCAGAAAAAAGCTTGGATGATGGAAGG
TTGCCTGATATATCACTCGATGGGCATGATATACAAGATGAAGTGAAGACAGCTGCAGATAAGGATGAGAAAATGGAGGATCAAGAGGTTGCTAATACCA
CTAGTATTGAAGAGATTGACAGAATTGAAGATCATGGGGCTGAAAGAACCACAGGTGTTCAAGAGGTTGAAAGAATGGAAACCACCAGAGTGGAAGAGGT
TGAGGGAATGGAGGATCAGCAGTTTAAGAAAGATAGTGAAGATATCCCTGTAGAGGTTTGTCCAGGAGTTAGTTGGGATGTCTTTCGTCGGCAGGATATT
CCAAAGTTAATTGATTATTTGAGAACATGCTATAAGGACCTTTGGAAGCCTGACAACATAGTGAATGATTTTGTCACAGATCCCCTTTATGACGGAACAG
TTTTCCTAAATGCGTTTCATAAGAGACAATTGAAGGAAGAGTTTGGAGTTGAACCATGGTCATTTGAACAACATTTGGGGCAAGCTGTATTCGTCCCAGC
TGGATGCCCTTTCCAAGCGAGGAATCTTCAGTCCAATGTTCAGTTGGGCCTTGATTTCTTATCTCCTGAAAGTCTGGGGGTGTCTGCTAGATTGGCAGAA
GAAATTCGCTGTCTTCCAAATGACCATGAAGCAAAACTTCAAGTTTTGGAGGTGGGTAAGATGTCACTCTATGCTGCTAGTTCAGCCATTAAAGAAGTCC
AGAAATTGGTGCTTGATCCTAAACTTGGTGCTGAAATTGGATTCGAGGATCGCAATTTAACTGCAGCAGTTGCTGAGAACTTGGAGAAAGGTGCTAAGCC
AAGACAGATAAGTTGTTCTTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.013G019000.4 pacid=42812635 polypeptide=Potri.013G019000.4.p locus=Potri.013G019000 ID=Potri.013G019000.4.v4.1 annot-version=v4.1
MDHLRSSSANGEENGGGIPDDLRCKRSDGKQWRCTAMSMPDKTVCEKHYIQAKKRAANSALRASLKKAKRKSIGESDFYLESKSDDFDMPLRNMKVEEDQ
PLSVSSKRYKEKVPKSQSRYSPETLIRSLRGQNSLKLNDDSQRDFEFEENWRSYKTTPRSTMESSRSRSQRSFDASAMTVSETVTEYSDASTDASEDTGG
QTCHQCRRNDRNSVTWCLKCDKRGFCDSCISEWYSDIPLEEIEKVCPACRGICNCRGCLRGDNMVKVRIREIPVLDKLQYLHCLLSSVLPIVKQIHQEQC
FEVELEQRLRGTDIDLVRAKLNADEQMCCNICRIPIIDYHRHCANCSYDLCLHCCQDLRGASKHGVENEVDDNQIDGRSQDNETPLEPVREPQVRLKLSD
KYQGWKANNDGSIPCPPKEHGGCNYSSLNLSRIFKMNWAAKLVKNVEEMVSGCKVYDAGTPQKSGLNDSTLCQYAHREDSDDNFLYCPLSEDVKADGINK
FRKHWVRGEPVIVKQVFDSSSISSWDPMAIWRGIRETSDEKKKGENRMVKAIDCLHWSEVDIDLDQFIRGYSEGRIRENGSPEMLKLKDWPSPSASEEFL
LYQRPESISKLPFLEFIHSRVGVLNVAAKLPHYSLQNDVGPKICISYGSHEDLGVGDSVIKLHFKTRDMVYLLVHTCEAKTKGSQESSSIDPEKSLDDGR
LPDISLDGHDIQDEVKTAADKDEKMEDQEVANTTSIEEIDRIEDHGAERTTGVQEVERMETTRVEEVEGMEDQQFKKDSEDIPVEVCPGVSWDVFRRQDI
PKLIDYLRTCYKDLWKPDNIVNDFVTDPLYDGTVFLNAFHKRQLKEEFGVEPWSFEQHLGQAVFVPAGCPFQARNLQSNVQLGLDFLSPESLGVSARLAE
EIRCLPNDHEAKLQVLEVGKMSLYAASSAIKEVQKLVLDPKLGAEIGFEDRNLTAAVAENLEKGAKPRQISCS
|
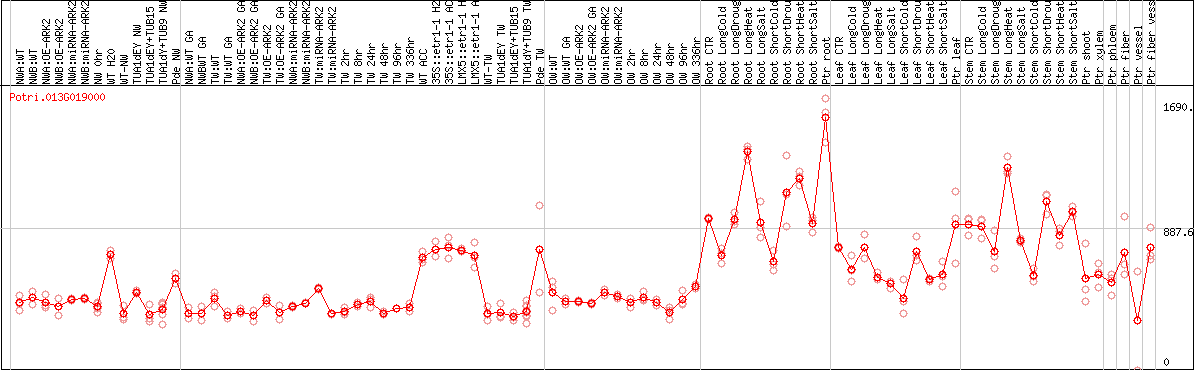
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G019000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
